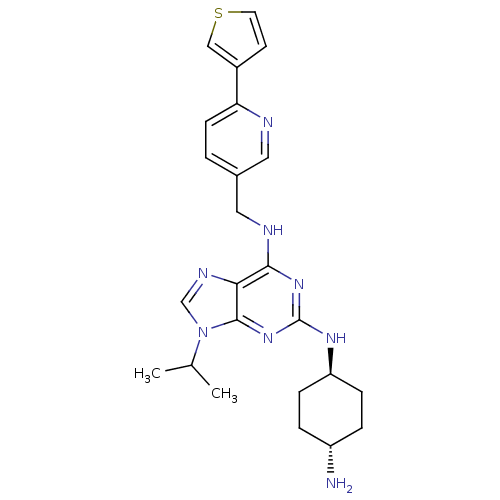
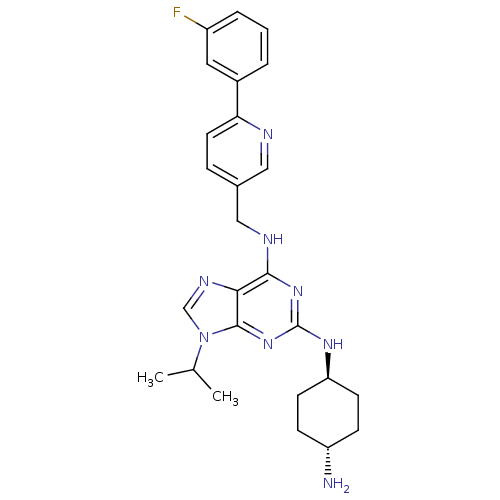
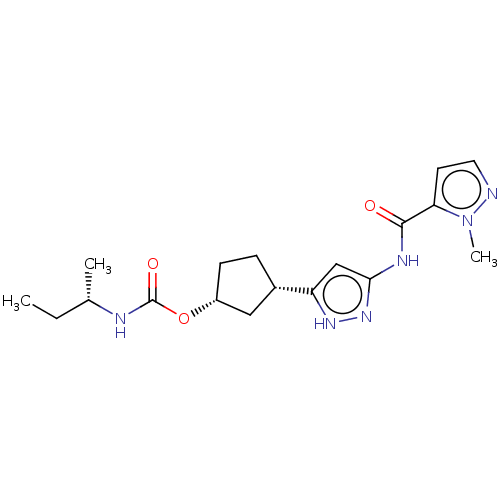
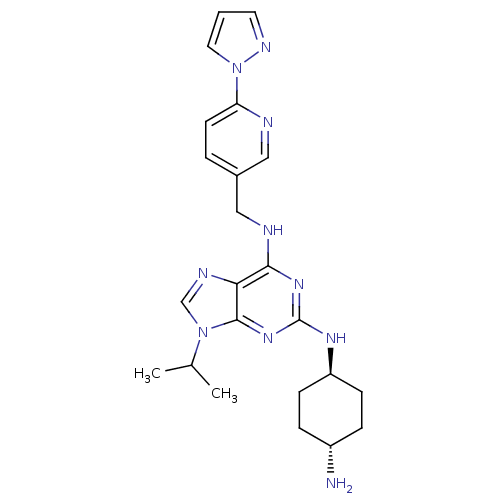
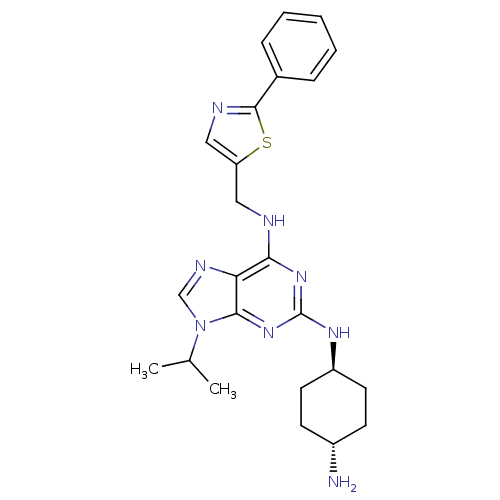
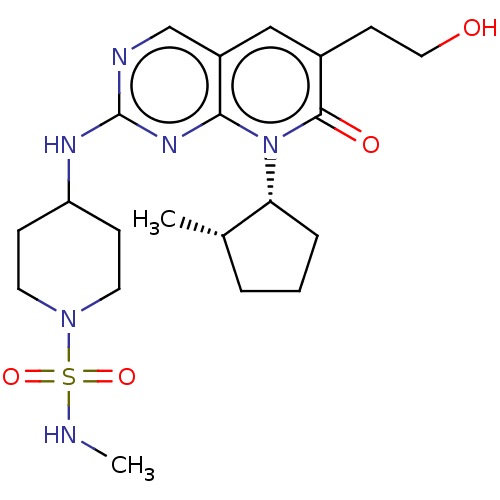
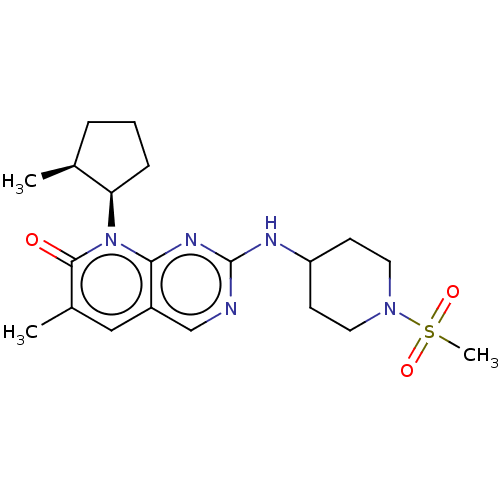
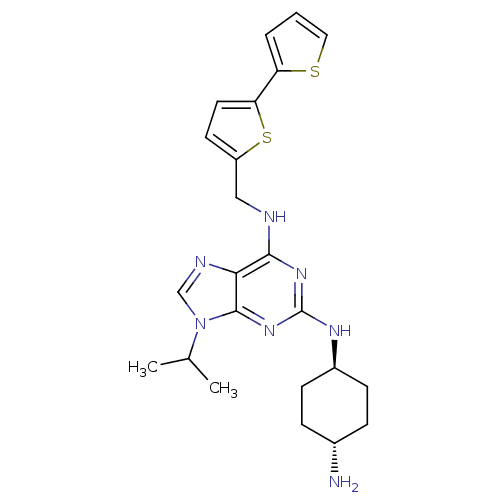
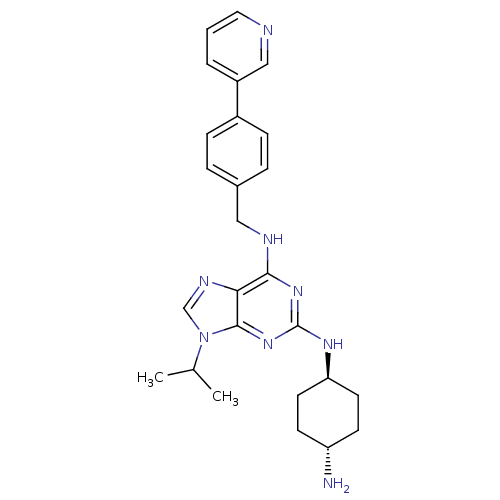
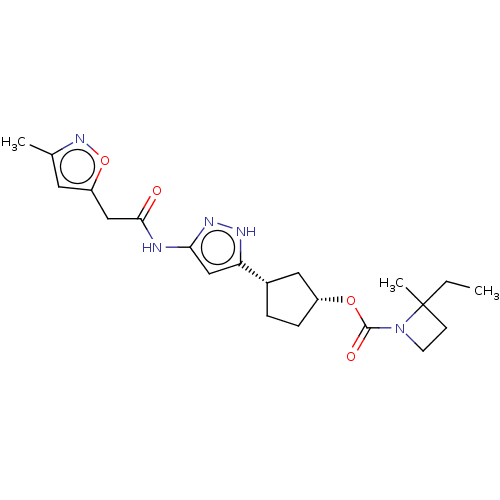
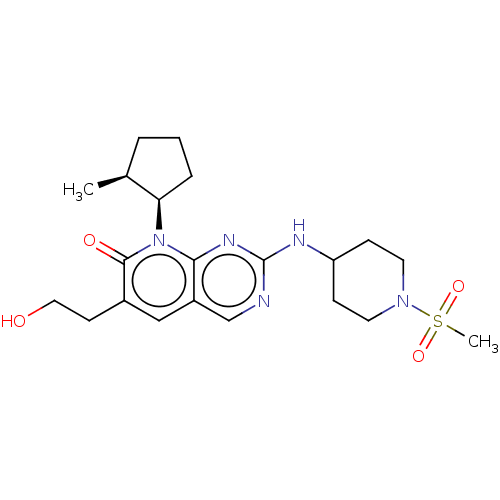
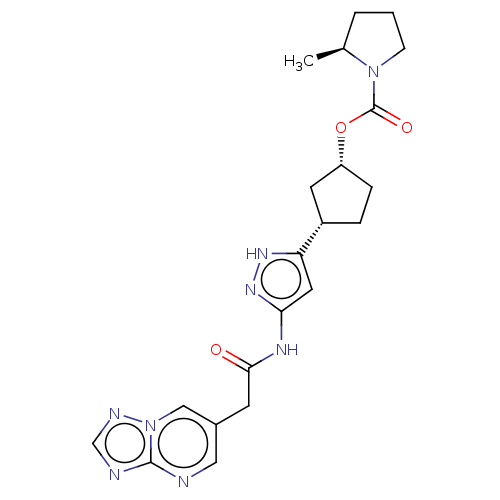
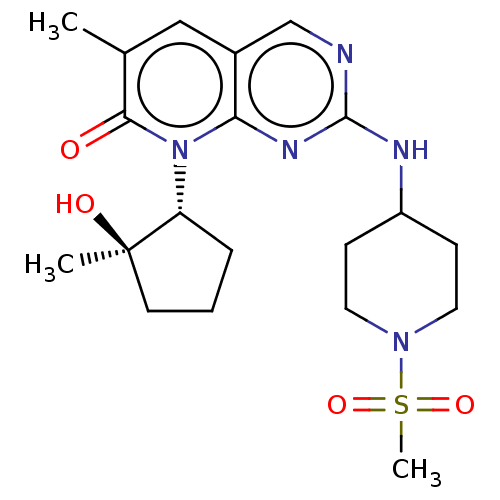
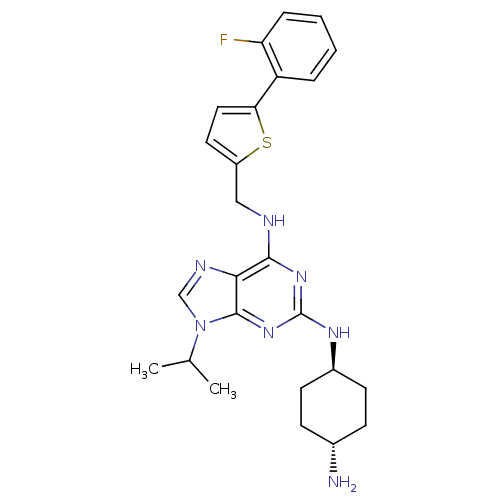
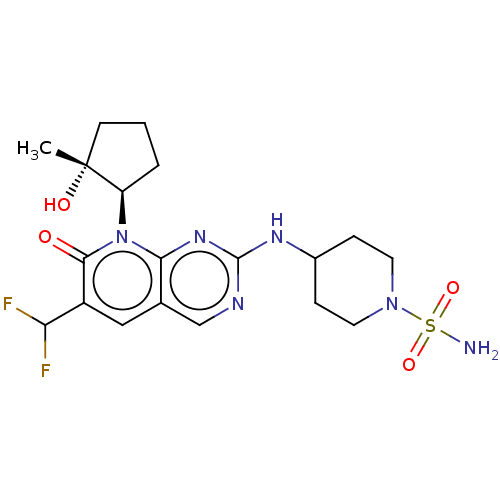
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1/G1/S-specific cyclin-E2(Human)
Amri
Curated by ChEMBL
Amri
Curated by ChEMBL
Affinity DataIC50: 0.0300nMAssay Description:Inhibition of Cdk2/cyclinEMore data for this Ligand-Target Pair
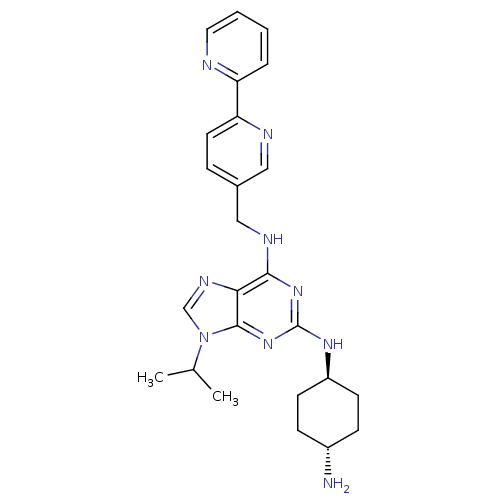
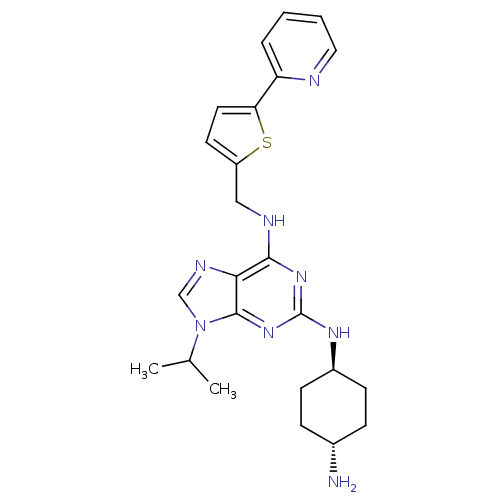
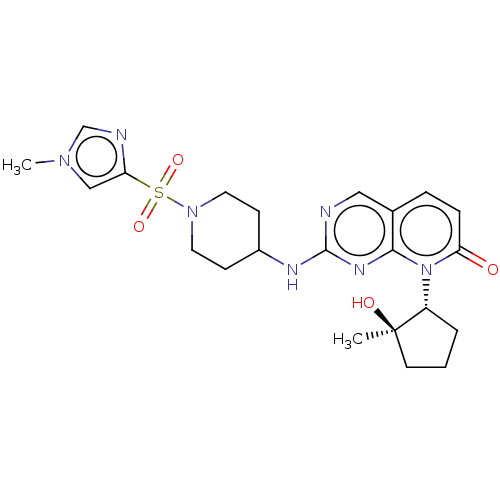
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1/G1/S-specific cyclin-E2(Human)
Amri
Curated by ChEMBL
Amri
Curated by ChEMBL
Affinity DataIC50: 0.0300nMAssay Description:Inhibition of Cdk2/cyclinEMore data for this Ligand-Target Pair
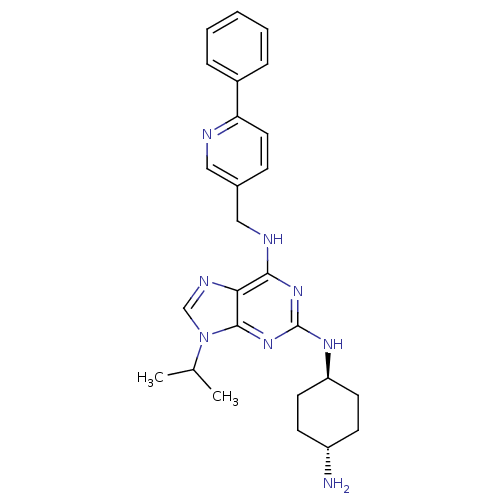
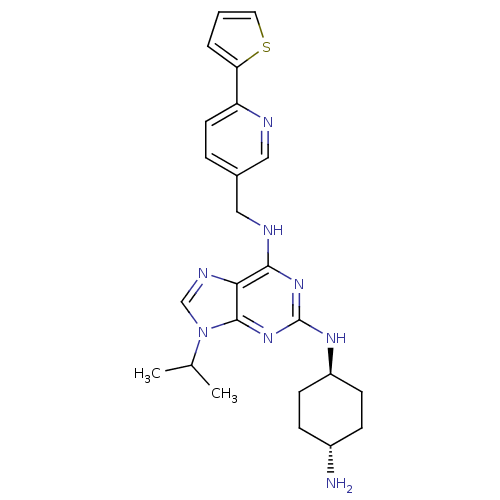
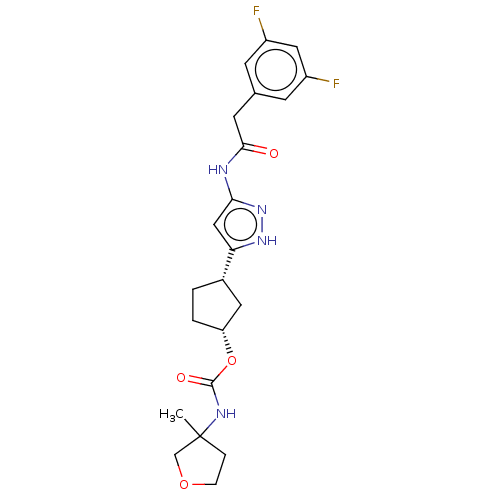
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1/G1/S-specific cyclin-E2(Human)
Amri
Curated by ChEMBL
Amri
Curated by ChEMBL
Affinity DataIC50: 0.0400nMAssay Description:Inhibition of Cdk2/cyclinEMore data for this Ligand-Target Pair
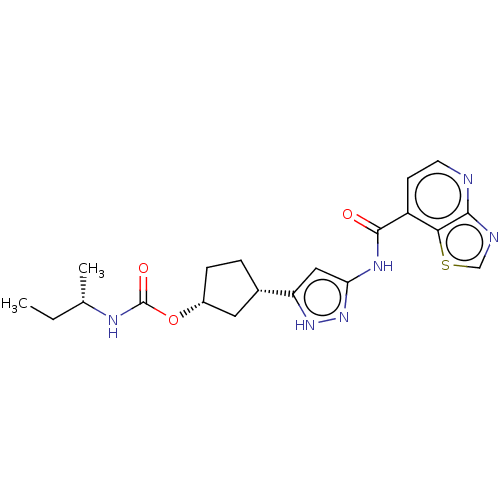
Affinity DataKi: 0.0400nMAssay Description:The purpose of CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a fluores...More data for this Ligand-Target Pair
Affinity DataKi: 0.0400nMAssay Description:The purpose of CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a fluores...More data for this Ligand-Target Pair
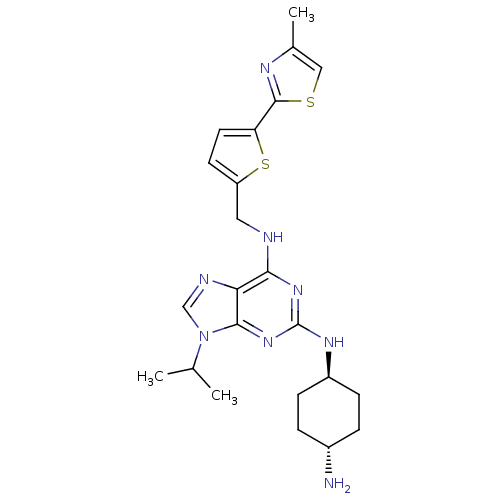
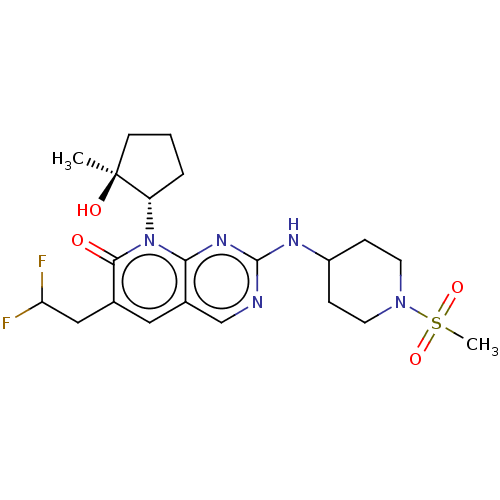
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1/G1/S-specific cyclin-E2(Human)
Amri
Curated by ChEMBL
Amri
Curated by ChEMBL
Affinity DataIC50: 0.0400nMAssay Description:Inhibition of Cdk2/cyclinEMore data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1/G1/S-specific cyclin-E2(Human)
Amri
Curated by ChEMBL
Amri
Curated by ChEMBL
Affinity DataIC50: 0.0400nMAssay Description:Inhibition of Cdk2/cyclinEMore data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1/G1/S-specific cyclin-E2(Human)
Amri
Curated by ChEMBL
Amri
Curated by ChEMBL
Affinity DataIC50: 0.0400nMAssay Description:Inhibition of Cdk2/cyclinEMore data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1/G1/S-specific cyclin-E2(Human)
Amri
Curated by ChEMBL
Amri
Curated by ChEMBL
Affinity DataIC50: 0.0400nMAssay Description:Inhibition of Cdk2/cyclinEMore data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The purpose of CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a fluores...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The purpose of CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a fluores...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1/G1/S-specific cyclin-E2(Human)
Amri
Curated by ChEMBL
Amri
Curated by ChEMBL
Affinity DataIC50: 0.0500nMAssay Description:Inhibition of Cdk2/cyclinEMore data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1/G1/S-specific cyclin-E2(Human)
Amri
Curated by ChEMBL
Amri
Curated by ChEMBL
Affinity DataIC50: 0.0500nMAssay Description:Inhibition of Cdk2/cyclinEMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0500nMAssay Description:Inhibition of recombinant Cdk2/cyclinAMore data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:The purpose of the CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a flu...More data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:The purpose of CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a fluores...More data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:The purpose of CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a fluores...More data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:The mobility shift assay electrophoretically separates the fluorescently labeled peptides (substrate and phosphorylated product) following the kinase...More data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:The mobility shift assay electrophoretically separates the fluorescently labeled peptides (substrate and phosphorylated product) following the kinase...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1/G1/S-specific cyclin-E2(Human)
Amri
Curated by ChEMBL
Amri
Curated by ChEMBL
Affinity DataIC50: 0.0600nMAssay Description:Inhibition of Cdk2/cyclinEMore data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:The purpose of the CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a flu...More data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:The purpose of the CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a flu...More data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:The purpose of the CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a flu...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1/G1/S-specific cyclin-E2(Human)
Amri
Curated by ChEMBL
Amri
Curated by ChEMBL
Affinity DataIC50: 0.0700nMAssay Description:Inhibition of Cdk2/cyclinEMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0700nMAssay Description:Inhibition of recombinant Cdk2/cyclinAMore data for this Ligand-Target Pair
Affinity DataKi: 0.0700nMAssay Description:The purpose of CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a fluores...More data for this Ligand-Target Pair
Affinity DataKi: 0.0700nMAssay Description:The purpose of CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a fluores...More data for this Ligand-Target Pair
Affinity DataKi: 0.0700nMAssay Description:The purpose of CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a fluores...More data for this Ligand-Target Pair
Affinity DataKi: 0.0700nMAssay Description:The purpose of CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a fluores...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The purpose of the CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a flu...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The purpose of CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a fluores...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The purpose of the CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a flu...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1/G1/S-specific cyclin-E2(Human)
Amri
Curated by ChEMBL
Amri
Curated by ChEMBL
Affinity DataIC50: 0.0800nMAssay Description:Inhibition of Cdk2/cyclinEMore data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The purpose of the CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a flu...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataKi: 0.0800nMAssay Description:The mobility shift assay electrophoretically separates the fluorescently labeled peptides (substrate and phosphorylated product) following the kinase...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The purpose of the CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a flu...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The mobility shift assay electrophoretically separates the fluorescently labeled peptides (substrate and phosphorylated product) following the kinase...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataKi: 0.0800nMAssay Description:The purpose of the CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a flu...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The purpose of CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a fluores...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The purpose of the CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a flu...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataKi: 0.0800nMAssay Description:The mobility shift assay electrophoretically separates the fluorescently labeled peptides (substrate and phosphorylated product) following the kinase...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The purpose of CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a fluores...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The purpose of CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a fluores...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0800nMAssay Description:Inhibition of recombinant Cdk2/cyclinAMore data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The purpose of the CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a flu...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The mobility shift assay electrophoretically separates the fluorescently labeled peptides (substrate and phosphorylated product) following the kinase...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The purpose of CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a fluores...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The purpose of CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a fluores...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The purpose of the CDK2/Cyclin E1 assay is to evaluate the inhibition (% inhibition, Kiapp and Ki values) of small molecule inhibitors by using a flu...More data for this Ligand-Target Pair
Affinity DataKi: 0.0900nMAssay Description:Inhibition of recombinant full length wild-type phosphorylated CDK2/Cyclin E1 (unknown origin) expressed in baculovirus expression system assessed as...More data for this Ligand-Target Pair