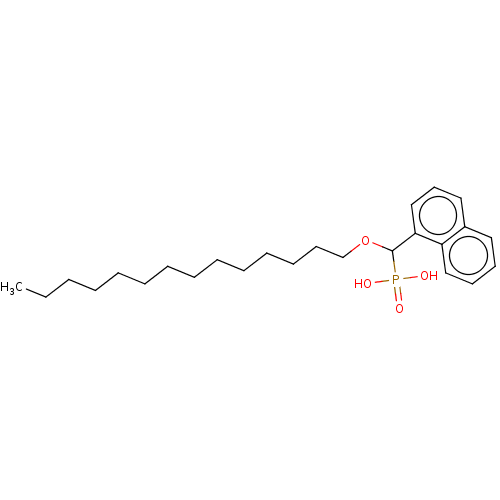
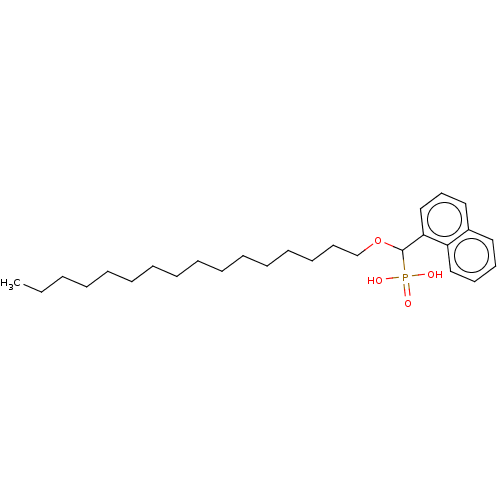
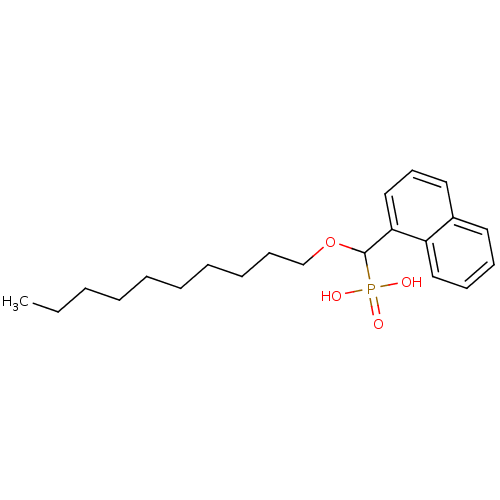
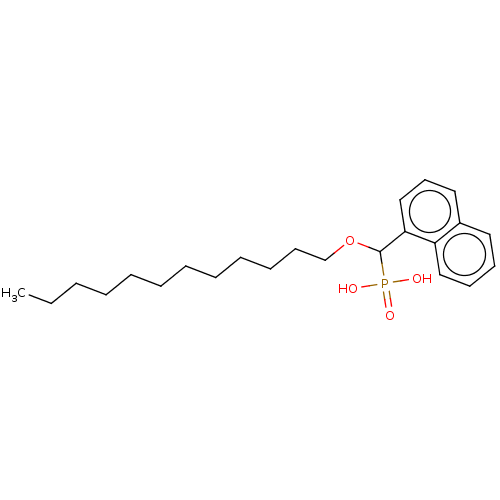
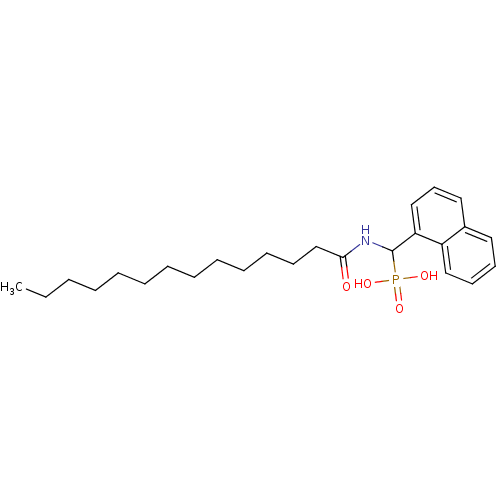
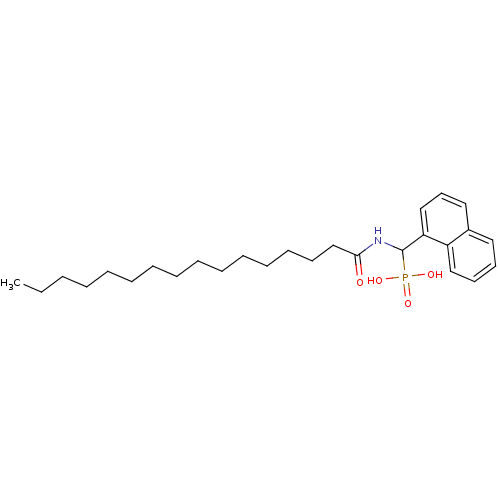
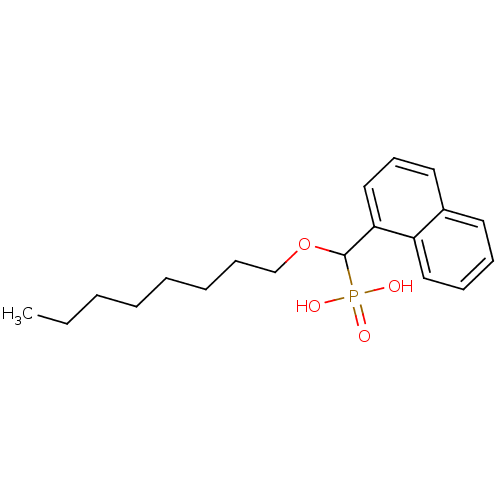
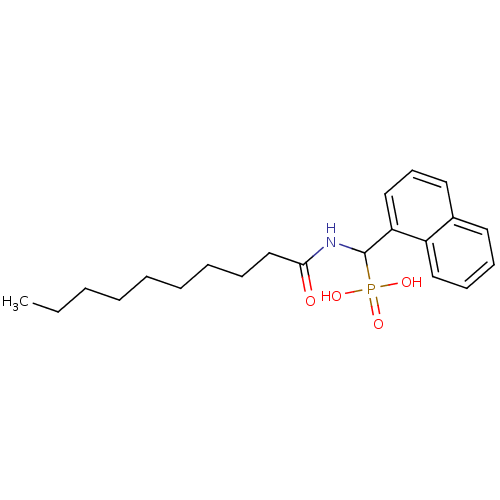
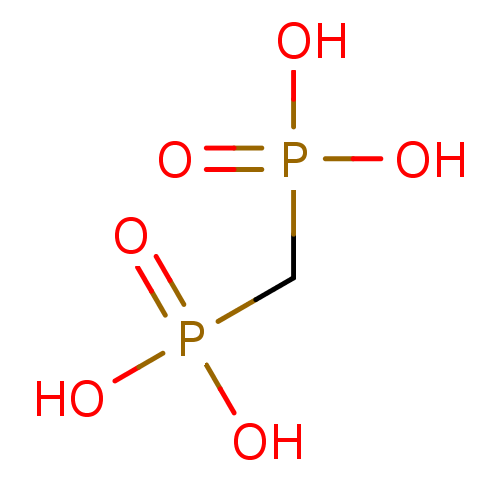
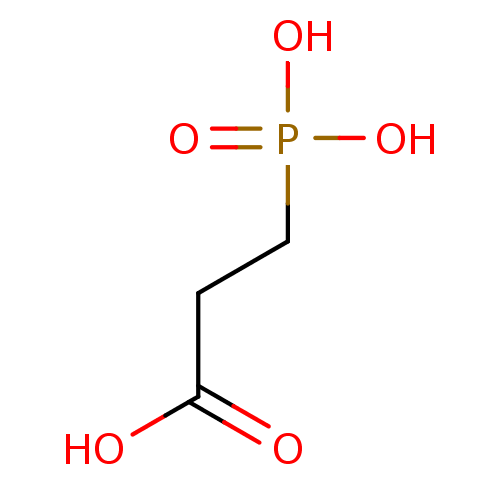
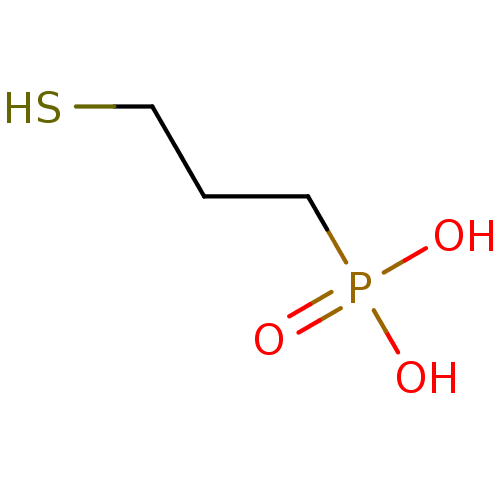
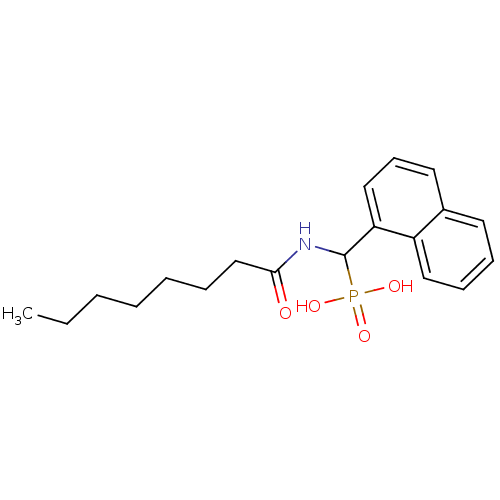
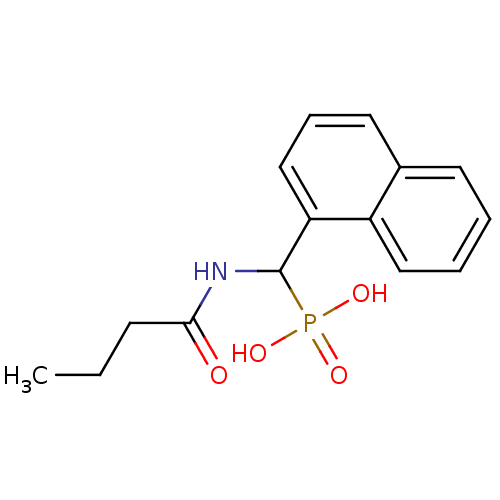
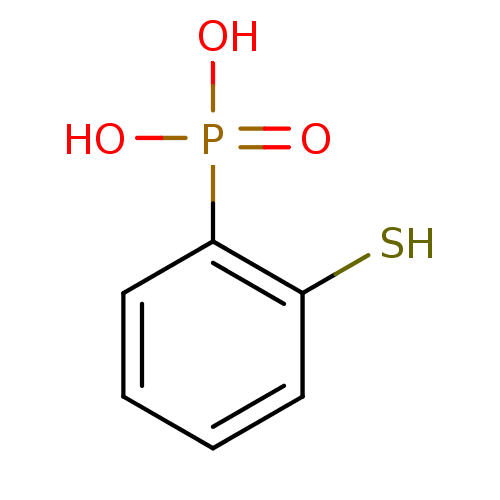
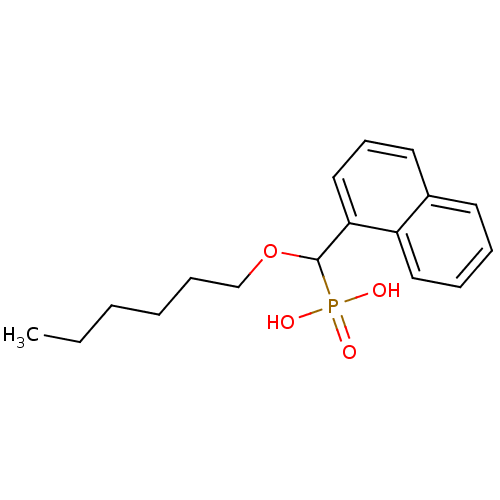
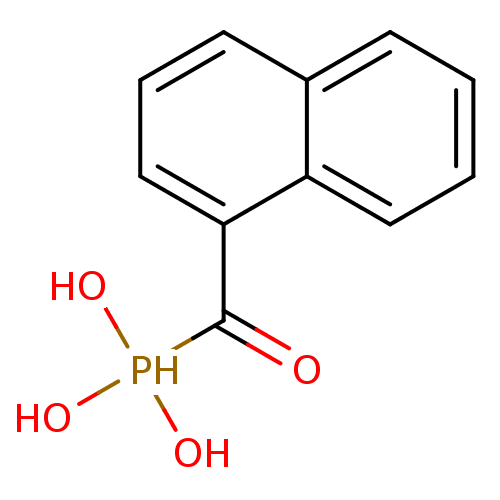
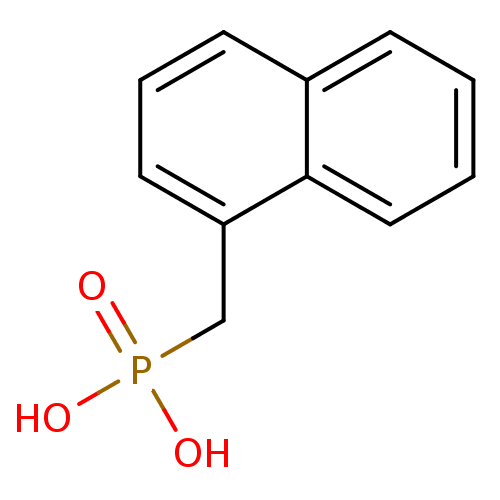
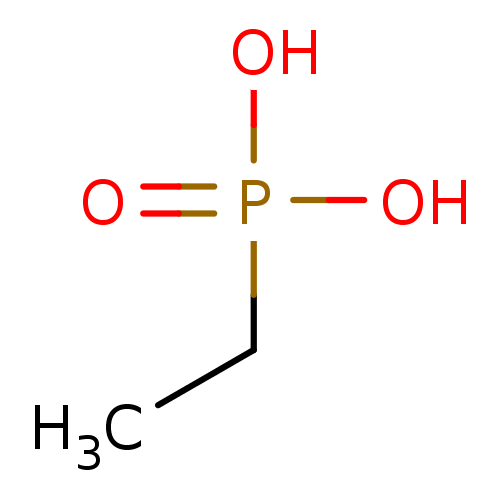
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 200nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 400nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 500nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 800nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 4.00E+3nMpH: 6.2Assay Description:Inhibition of red kidney bean PAP at pH 6.2More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 4.50E+3nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 5.00E+3nMpH: 4.9Assay Description:Competitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometryMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.10E+4nMpH: 4.9Assay Description:Competitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometryMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.40E+4nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 3.10E+4nMpH: 4.9Assay Description:Competitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometryMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 5.00E+4nMpH: 6.2Assay Description:Inhibition of red kidney bean PAP at pH 6.2More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 5.70E+4nMpH: 4.9Assay Description:Competitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometryMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataIC50: 8.00E+4nMAssay Description:Inhibition of red kidney beans purple acid phosphataseMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.02E+5nMpH: 4.9Assay Description:Uncompetitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometr...More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.40E+5nMAssay Description:Inhibition of red kidney beans purple acid phosphataseMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.60E+5nMAssay Description:Inhibition of red kidney beans purple acid phosphataseMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataIC50: 1.60E+5nMAssay Description:Inhibition of red kidney beans purple acid phosphataseMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.60E+5nMAssay Description:Inhibition of red kidney bean PAP using pNPP as substrateMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.95E+5nMpH: 4.9Assay Description:Competitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometryMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 2.22E+5nMpH: 4.9Assay Description:Competitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometryMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 2.38E+5nMpH: 4.9Assay Description:Competitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometryMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataIC50: 3.50E+5nMAssay Description:Inhibition of red kidney beans purple acid phosphataseMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 4.43E+5nMpH: 4.9Assay Description:Uncompetitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometr...More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 4.46E+5nMpH: 4.9Assay Description:Uncompetitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometr...More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 5.90E+5nMAssay Description:Inhibition of red kidney beans purple acid phosphataseMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 6.54E+5nMpH: 4.9Assay Description:Uncompetitive inhibition of red kidney bean PAP using para-nitrophenol as substrate at pH 4.9 preincubated for 10 mins by UV-visible spectrophotometr...More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 1.47E+6nMpH: 6.2Assay Description:Inhibition of red kidney bean PAP at pH 6.2More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataIC50: 3.00E+6nMAssay Description:Inhibition of red kidney beans purple acid phosphataseMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataIC50: 5.00E+6nMpH: 6.2Assay Description:Inhibition of red kidney bean PAP at pH 6.2More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataIC50: 7.00E+6nMpH: 6.2Assay Description:Inhibition of red kidney bean PAP at pH 6.2More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Kidney bean)
The University of Queensland
Curated by ChEMBL
The University of Queensland
Curated by ChEMBL
Affinity DataKi: 7.60E+6nMAssay Description:Inhibition of red kidney beans purple acid phosphataseMore data for this Ligand-Target Pair