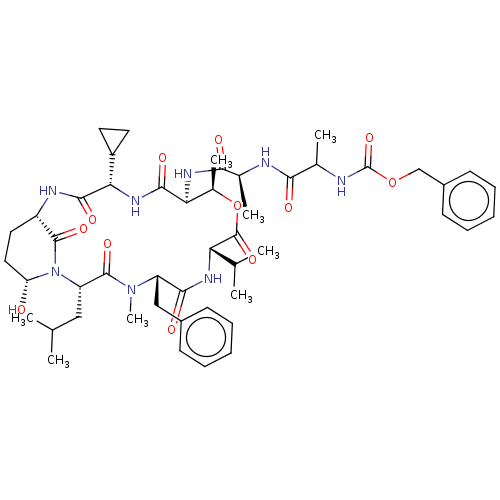
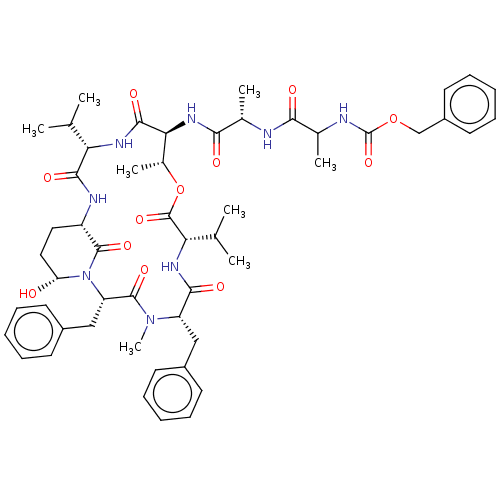
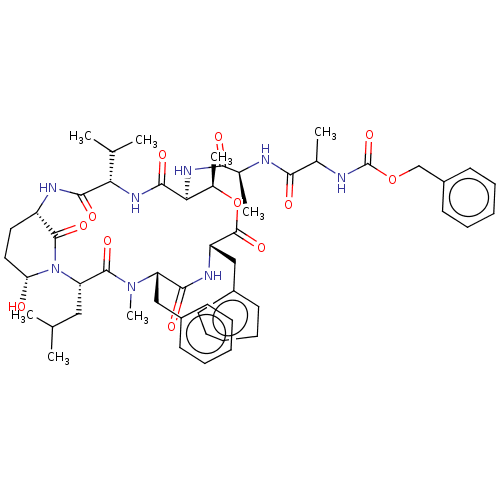
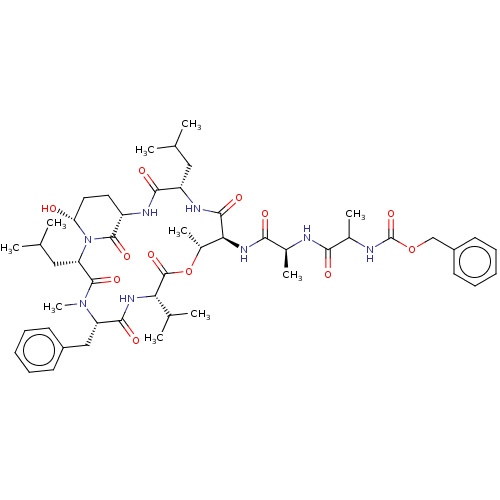
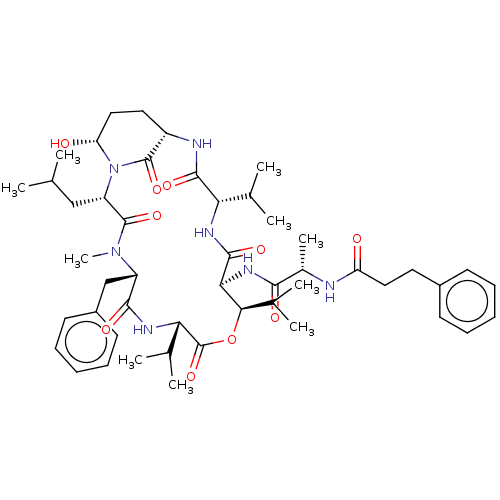
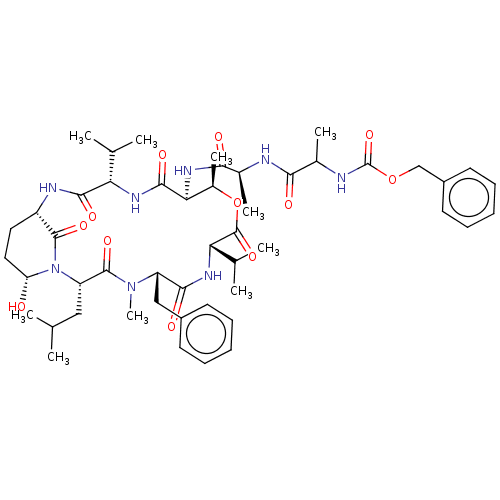
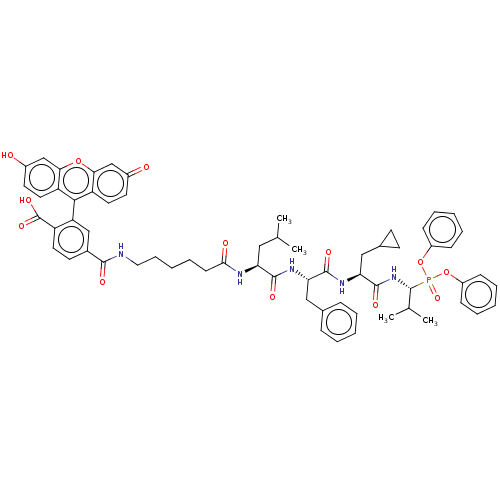
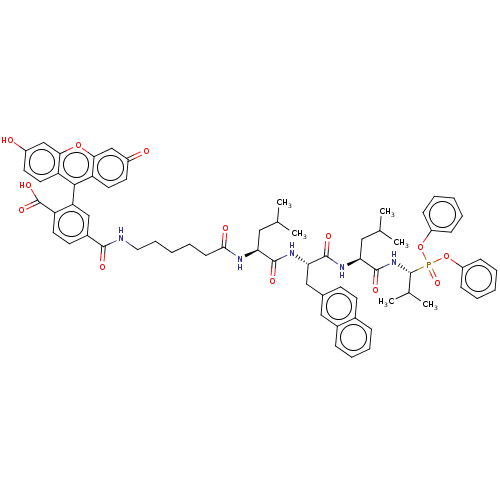
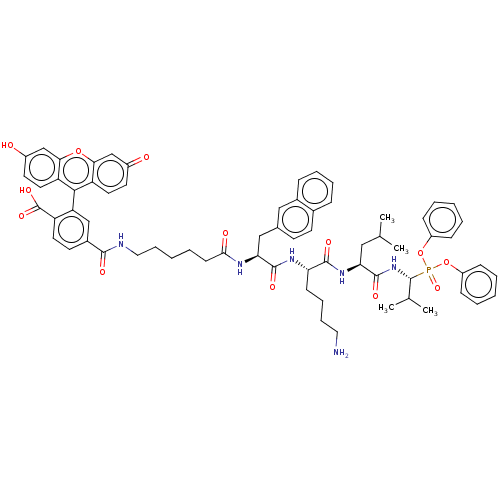
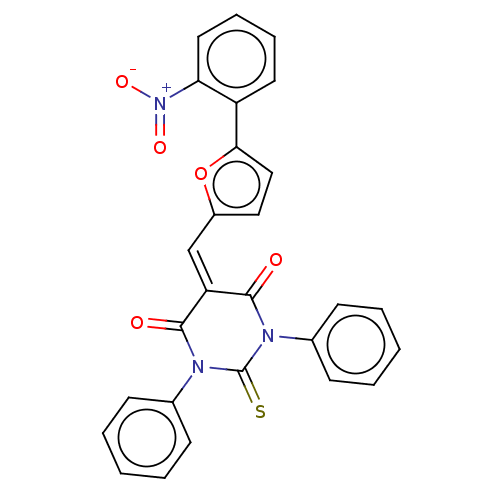
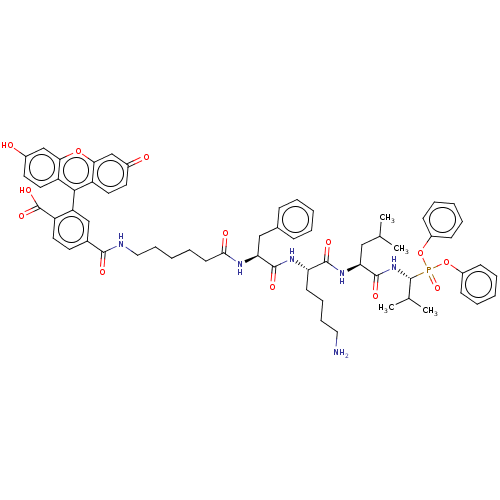
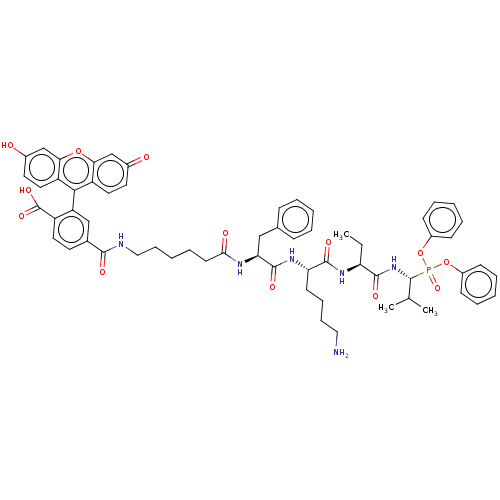
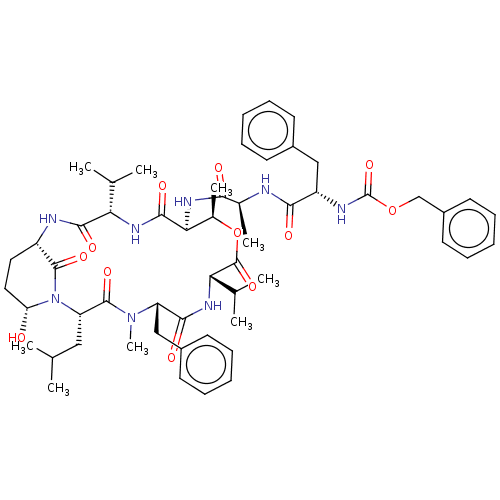
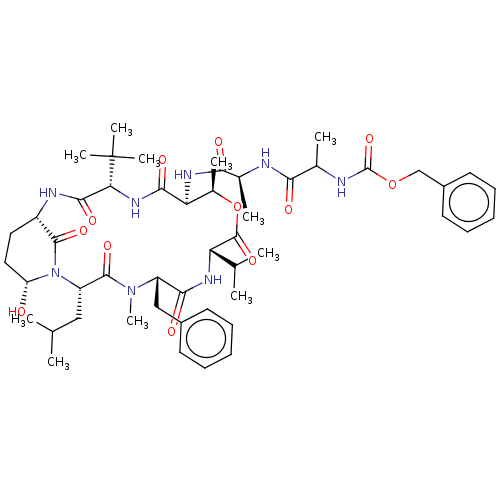
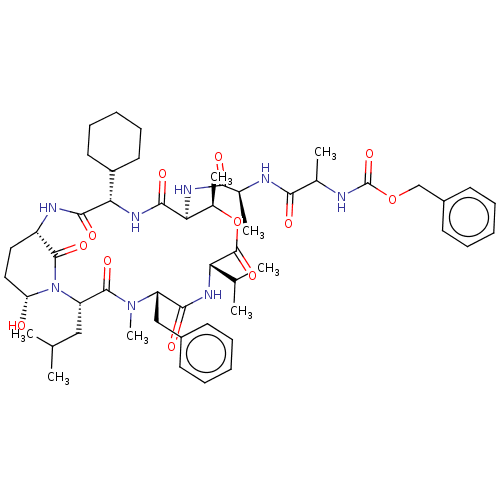
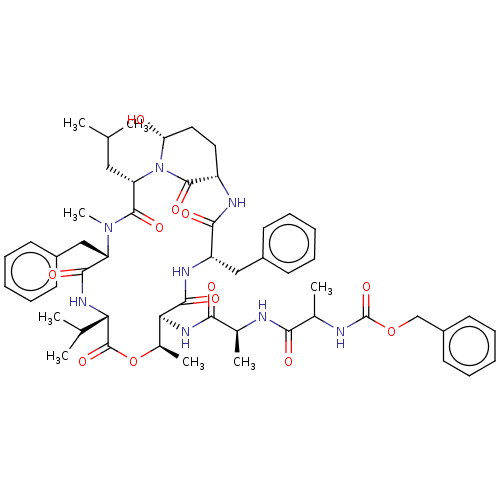
Affinity DataKi: 260nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataKi: 260nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataKi: 270nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataKi: 490nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataKi: 690nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataKi: 700nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataKi: 1.11E+3nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataKi: 1.38E+3nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataKi: 2.10E+3nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataKi: 2.30E+3nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataKi: 2.80E+3nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataKi: 4.00E+3nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataKi: 4.10E+3nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataIC50: 5.20E+3nMAssay Description:Inhibition of recombinant human HTRA2 using H2-OPT as substrate assessed as increase in fluorescence and measured for 2 hrs by fluorescent plate read...More data for this Ligand-Target Pair
Affinity DataIC50: 5.40E+3nMAssay Description:Inhibition of recombinant human HTRA2 using H2-OPT as substrate assessed as increase in fluorescence and measured for 2 hrs by fluorescent plate read...More data for this Ligand-Target Pair
Affinity DataKi: 6.90E+3nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataIC50: 7.80E+3nMAssay Description:Inhibition of recombinant human HTRA2 using H2-OPT as substrate assessed as increase in fluorescence and measured for 2 hrs by fluorescent plate read...More data for this Ligand-Target Pair
Affinity DataIC50: 8.10E+3nMAssay Description:Inhibition of recombinant human HTRA2 using H2-OPT as substrate assessed as increase in fluorescence and measured for 2 hrs by fluorescent plate read...More data for this Ligand-Target Pair
Affinity DataIC50: 8.30E+3nMAssay Description:Inhibition of recombinant human HTRA2 using H2-OPT as substrate assessed as increase in fluorescence and measured for 2 hrs by fluorescent plate read...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibition of recombinant human HTRA2 using H2-OPT as substrate assessed as increase in fluorescence and measured for 2 hrs by fluorescent plate read...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+4nMAssay Description:Inhibition of recombinant human HTRA2 using H2-OPT as substrate assessed as increase in fluorescence and measured for 2 hrs by fluorescent plate read...More data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+4nMAssay Description:Inhibition of recombinant human HTRA2 using H2-OPT as substrate assessed as increase in fluorescence and measured for 2 hrs by fluorescent plate read...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+4nMAssay Description:Inhibition of recombinant human HTRA2 using H2-OPT as substrate assessed as increase in fluorescence and measured for 2 hrs by fluorescent plate read...More data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:In general, proteolytic activity was tested by monitoring the cleavage of the specific chromogenic substrate at 405 nm wavelength for 60 or 120 minut...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of recombinant human HTRA2 using H2-OPT as substrate assessed as increase in fluorescence and measured for 2 hrs by fluorescent plate read...More data for this Ligand-Target Pair