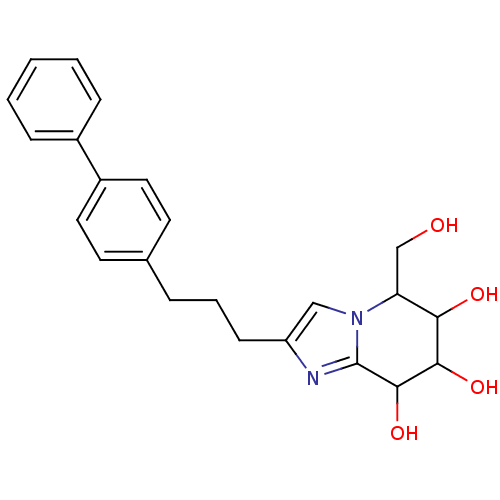
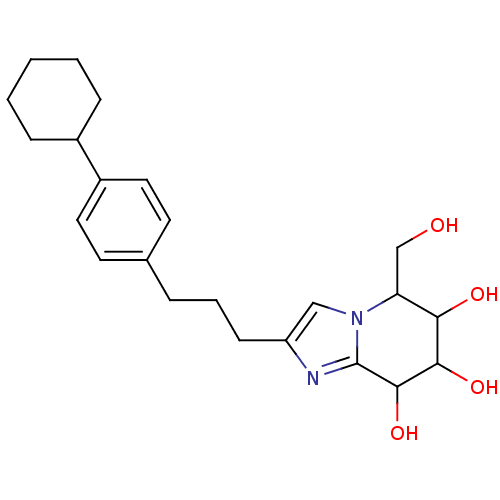
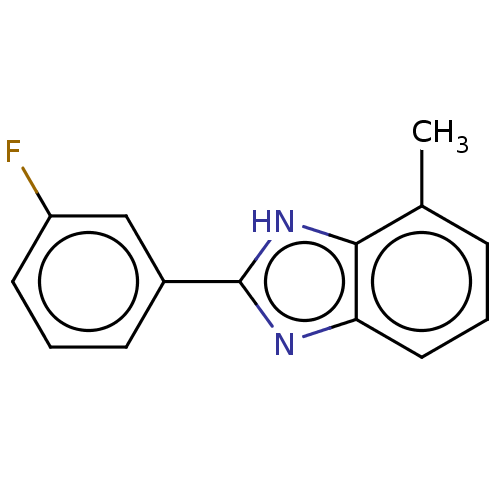
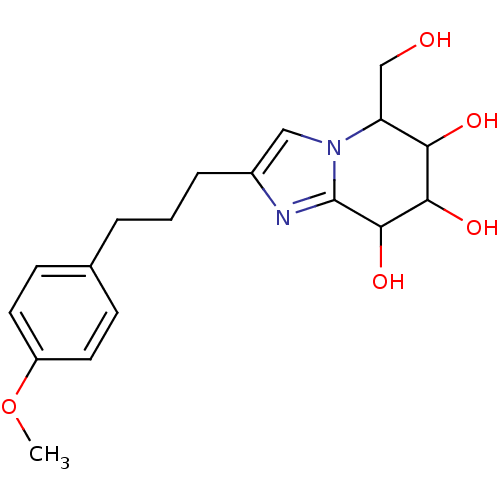
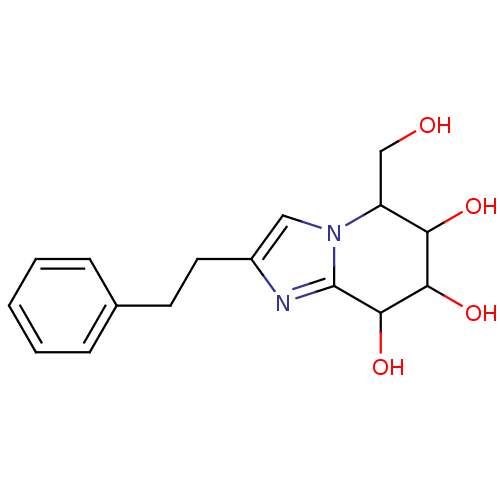
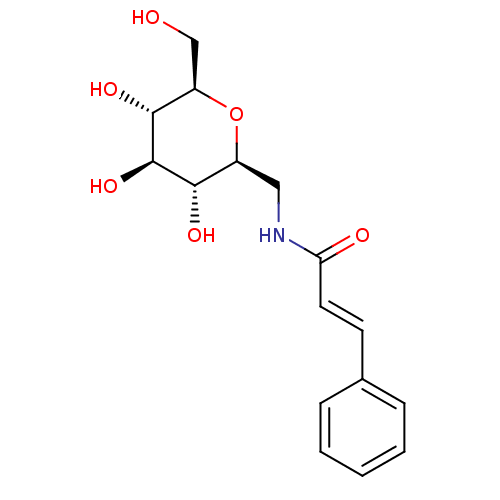
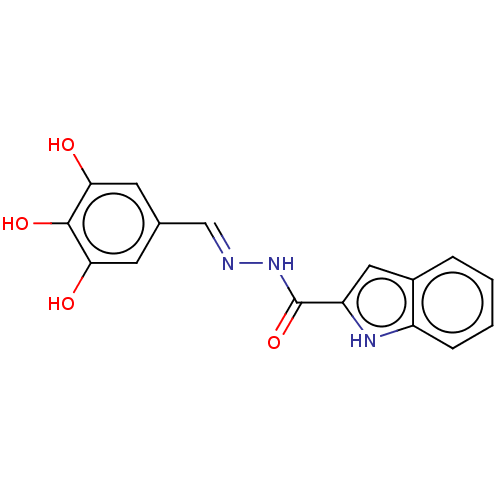
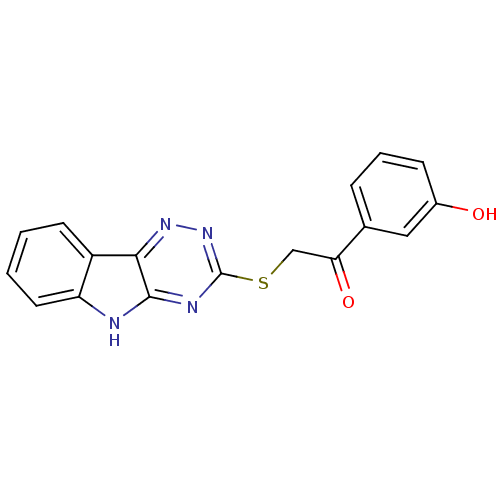
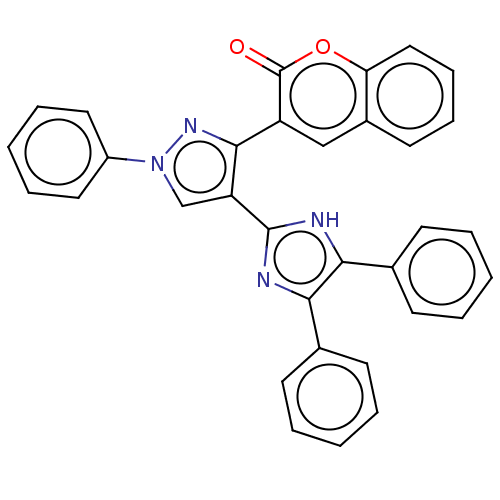
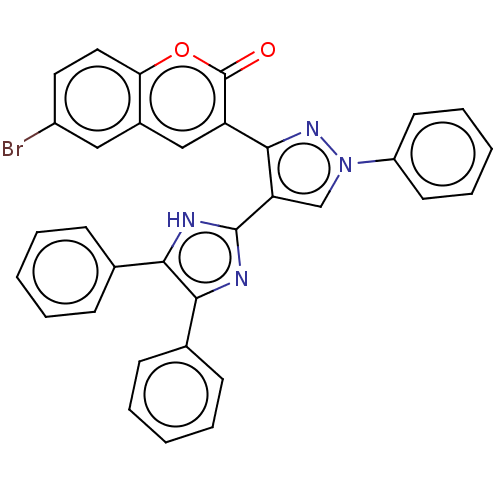
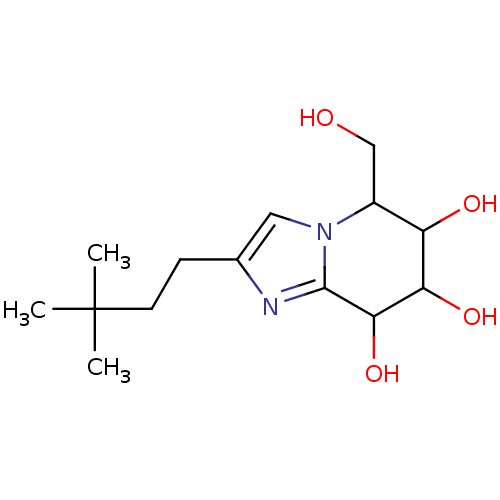
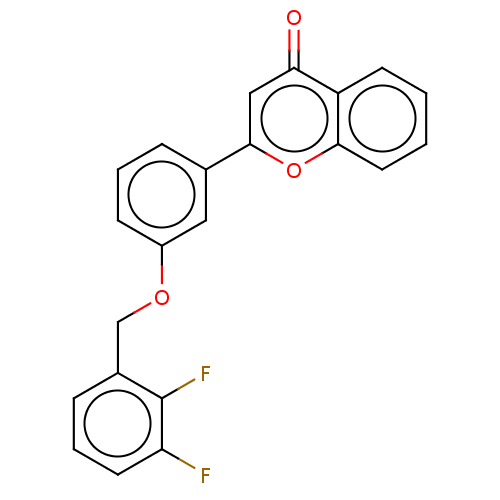
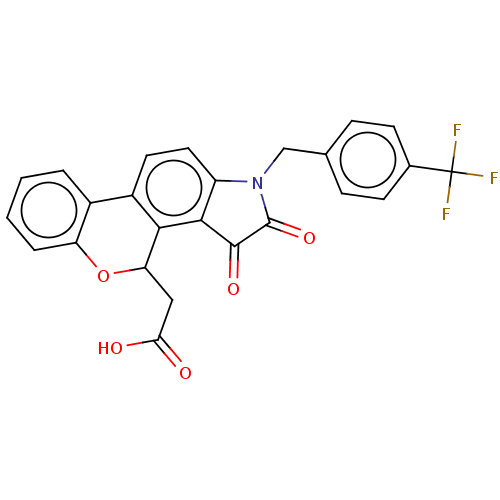
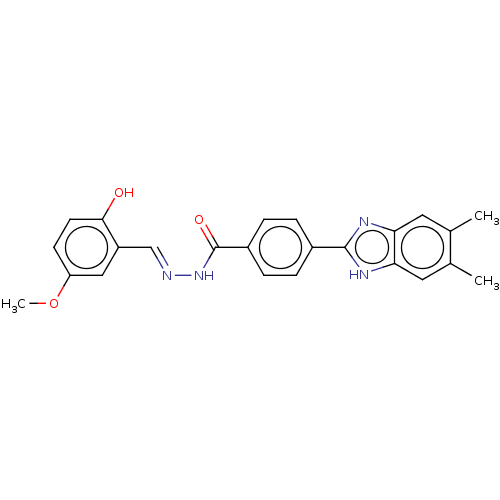
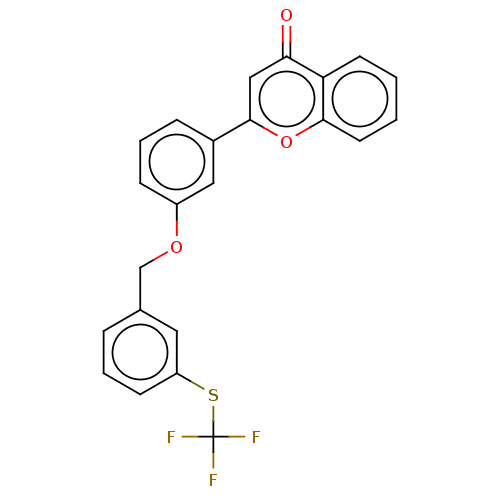
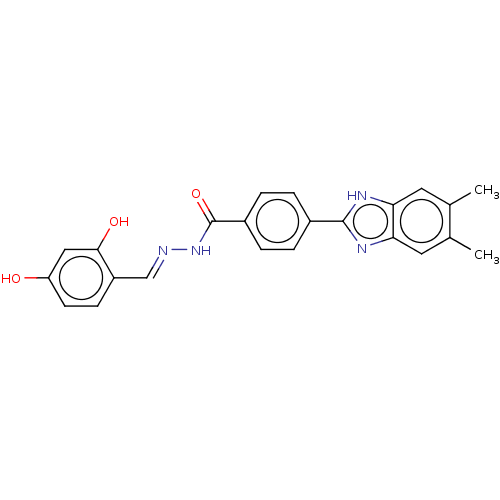
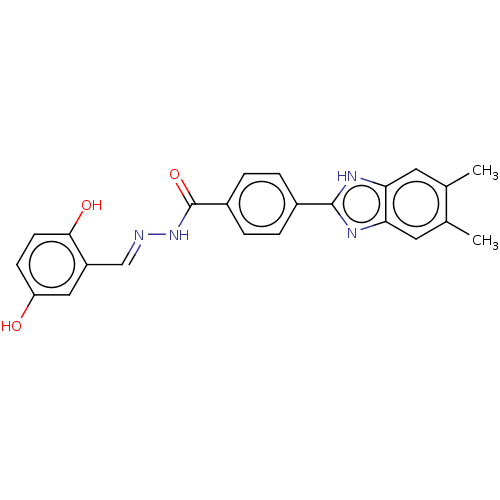
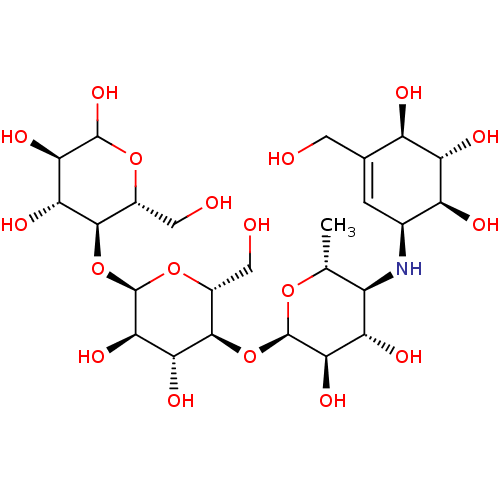
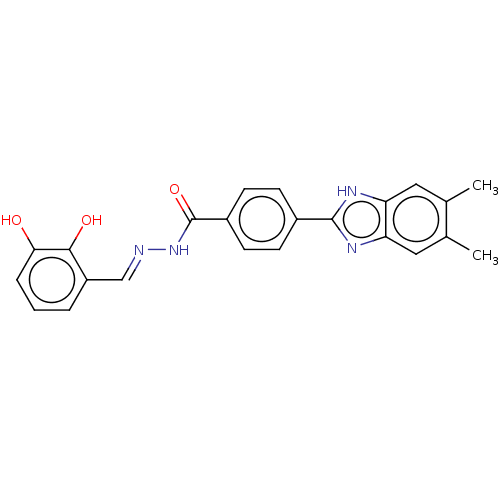
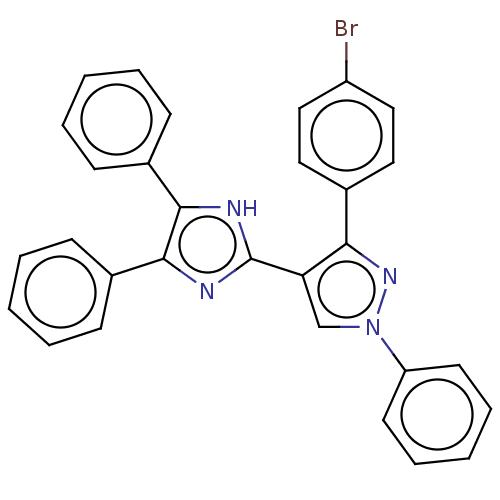
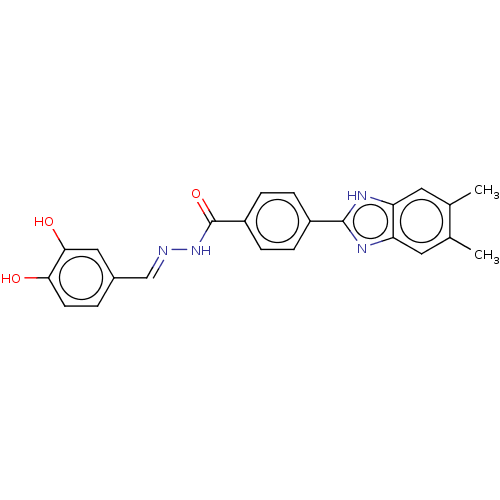
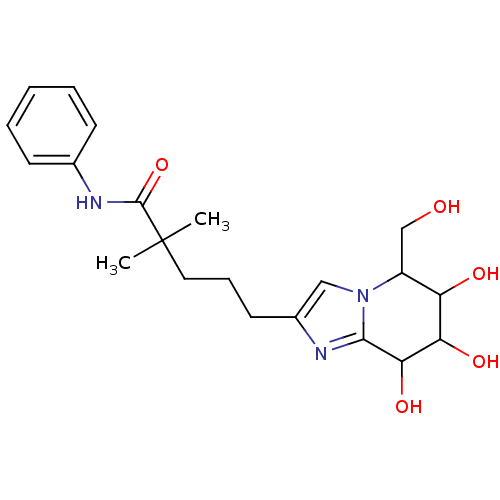
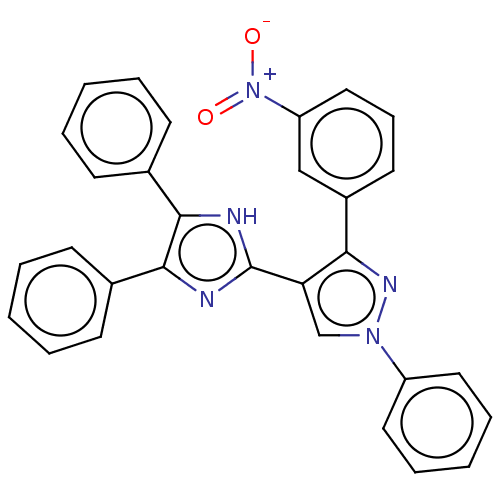
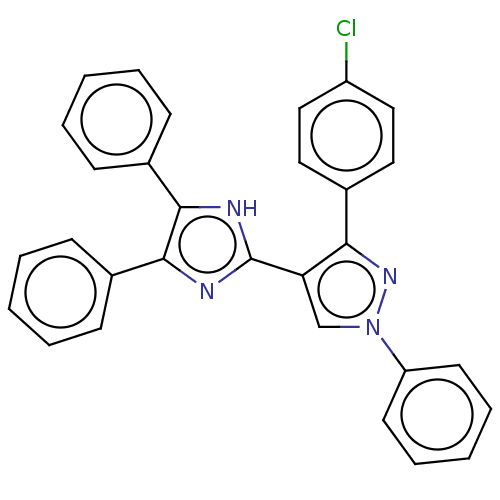
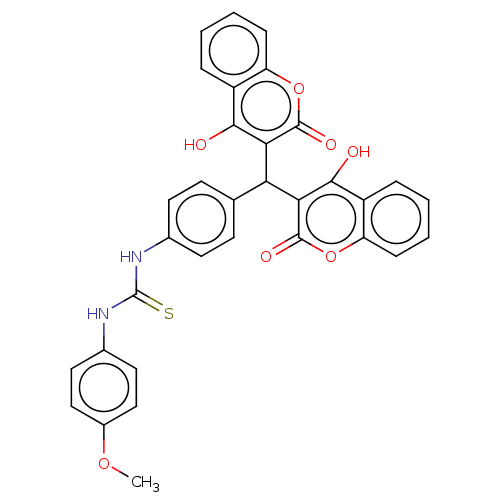
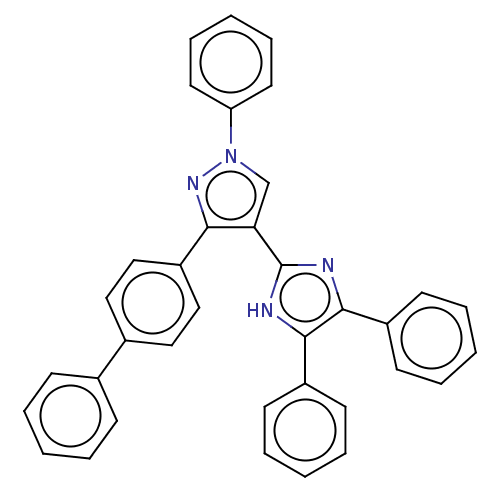
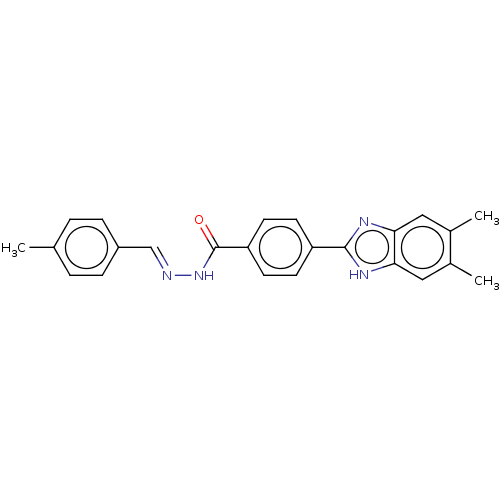
Affinity DataIC50: 6nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
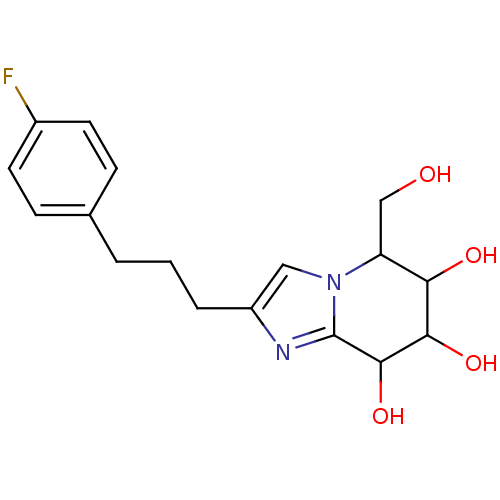
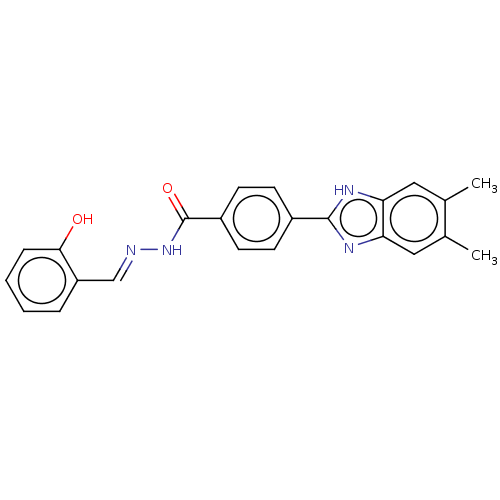
Affinity DataIC50: 52nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
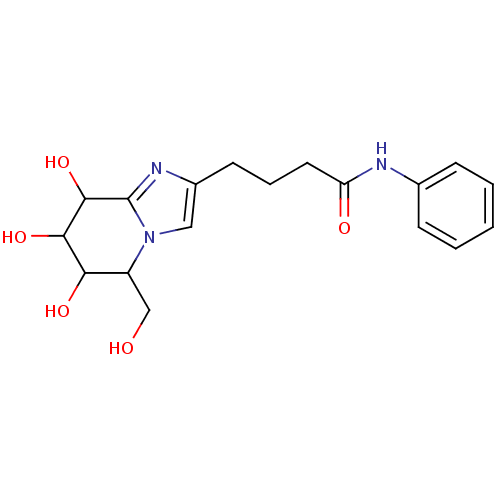
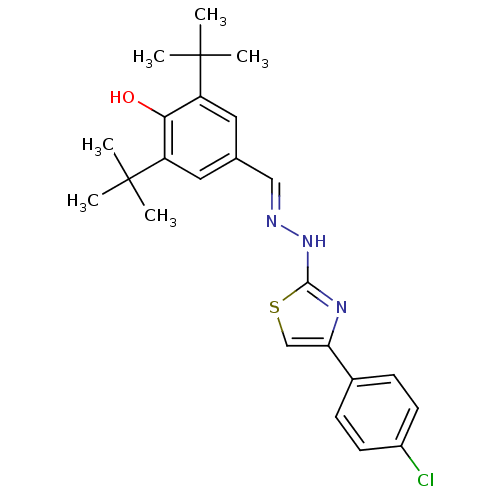
Affinity DataIC50: 165nMAssay Description:The 135 µL of 50 mM phosphate saline buffer pH (6.8), solution of enzyme (20 µL), and 20 µL of test sample with 70% DMSO was added to 96-well plate. ...More data for this Ligand-Target Pair
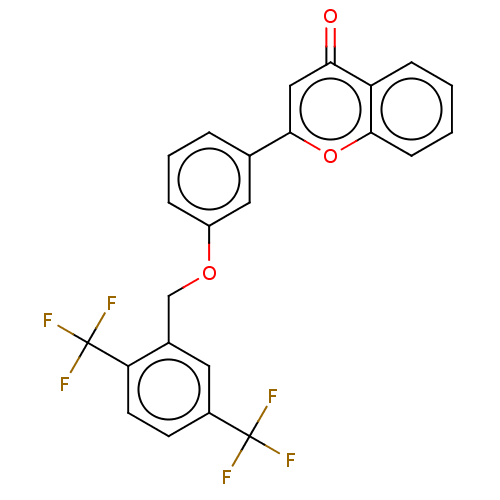
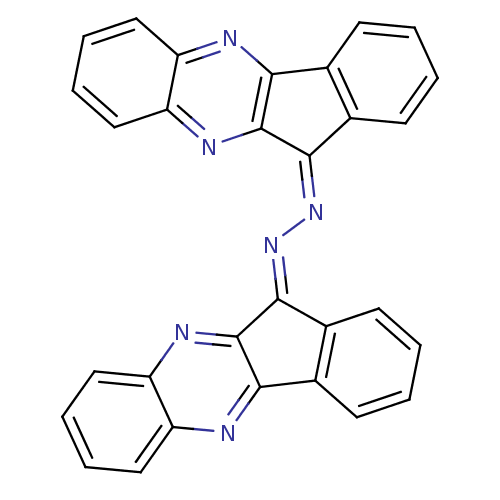
Affinity DataIC50: 280nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 300nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 570nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 730nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 970nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 1.05E+3nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 1.26E+3nMpH: 6.8Assay Description:The alpha-glucosidase inhibition assay had been carried out using baker's yeast alpha-glucosidase (EC 3.2.1.20) and p-nitrophenyl-alpha-D-glucopyrano...More data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+3nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+3nMAssay Description:The 135 µL of 50 mM phosphate saline buffer pH (6.8) was added in the 96-well plate and 20 µL of test sample with 70% DMSO added into the wells. The ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.46E+3nMT: 2°CAssay Description:Typically, α-glucosidase activity was assayed in 50 mM phosphate buffer (pH 6.8) containing 5% v/v DMSO and PNP glycoside was used as a substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 2.78E+3nMAssay Description:At 37 °C, after incubation of 10 min of a reaction mixture solution of 70 μL (50 mM) phosphate buffer of 6.8 pH, 10 μL (0.5 mM) of a test c...More data for this Ligand-Target Pair
Affinity DataIC50: 2.95E+3nMAssay Description:At 37 °C, after incubation of 10 min of a reaction mixture solution of 70 μL (50 mM) phosphate buffer of 6.8 pH, 10 μL (0.5 mM) of a test c...More data for this Ligand-Target Pair
Affinity DataIC50: 3.42E+3nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 3.49E+3nMpH: 6.8Assay Description:The alpha-glucosidase inhibition assay had been carried out using baker's yeast alpha-glucosidase (EC 3.2.1.20) and p-nitrophenyl-alpha-D-glucopyrano...More data for this Ligand-Target Pair
Affinity DataIC50: 4.80E+3nMAssay Description:Inhibition of bakers yeast alpha-glucosidase using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 30 mins by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataKi: 4.86E+3nMAssay Description:Mixed type inhibition of baker's yeast alpha-glucosidase by Dixon plot analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 5.30E+3nMAssay Description:The 135 µL of 50 mM phosphate saline buffer pH (6.8), solution of enzyme (20 µL), and 20 µL of test sample with 70% DMSO was added to 96-well plate. ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.45E+3nMpH: 6.8Assay Description:The alpha-glucosidase inhibition assay had been carried out using baker's yeast alpha-glucosidase (EC 3.2.1.20) and p-nitrophenyl-alpha-D-glucopyrano...More data for this Ligand-Target Pair
Affinity DataIC50: 5.60E+3nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 7.81E+3nMAssay Description:At 37 °C, after incubation of 10 min of a reaction mixture solution of 70 μL (50 mM) phosphate buffer of 6.8 pH, 10 μL (0.5 mM) of a test c...More data for this Ligand-Target Pair
Affinity DataIC50: 8.40E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.44E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.44E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.66E+3nMpH: 6.8Assay Description:The alpha-glucosidase inhibition assay had been carried out using baker's yeast alpha-glucosidase (EC 3.2.1.20) and p-nitrophenyl-alpha-D-glucopyrano...More data for this Ligand-Target Pair
Affinity DataIC50: 8.74E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.74E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 9.37E+3nMAssay Description:α-Glucosidase inhibitory activity was evaluated at 37 °C, by using 0.1M phosphatebuffer (pH6.8) [28]. The enzyme(EC3.2.1.20, Saccharomyces cerev...More data for this Ligand-Target Pair
Affinity DataIC50: 9.99E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMAssay Description:At 37 °C, after incubation of 10 min of a reaction mixture solution of 70 μL (50 mM) phosphate buffer of 6.8 pH, 10 μL (0.5 mM) of a test c...More data for this Ligand-Target Pair
Affinity DataIC50: 1.21E+4nMAssay Description:Inhibition of bakers yeast alpha-glucosidase using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 30 mins by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 1.25E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.28E+4nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 1.28E+4nMAssay Description:At 37 °C, after incubation of 10 min of a reaction mixture solution of 70 μL (50 mM) phosphate buffer of 6.8 pH, 10 μL (0.5 mM) of a test c...More data for this Ligand-Target Pair
Affinity DataIC50: 1.32E+4nMAssay Description:At 37 °C, after incubation of 10 min of a reaction mixture solution of 70 μL (50 mM) phosphate buffer of 6.8 pH, 10 μL (0.5 mM) of a test c...More data for this Ligand-Target Pair
Affinity DataIC50: 1.35E+4nMpH: 6.8 T: 2°CAssay Description:The enzyme inhibition was evaluated according to the method previously reported by Taha et al. with slight modification. Various concentration of tes...More data for this Ligand-Target Pair
Affinity DataKi: 1.48E+4nMAssay Description:Non-competitive inhibition of bakers yeast alpha-glucosidase using p-nitrophenyl-alpha-D-glucopyranoside as substrate by Lineweaver-Burk plot analysi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.51E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.52E+4nMAssay Description:Inhibition of bakers yeast alpha-glucosidase using p-nitrophenyl-alpha-D-glucopyranoside as substrate incubated for 15 mins followed by substrate add...More data for this Ligand-Target Pair
Affinity DataIC50: 1.57E+4nMAssay Description:Inhibition of bakers yeast alpha-glucosidase using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 30 mins by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+4nMAssay Description:Inhibition of bakers yeast alpha-glucosidase using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 30 mins by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 1.61E+4nMAssay Description:The 135 µL of 50 mM phosphate saline buffer pH (6.8), solution of enzyme (20 µL), and 20 µL of test sample with 70% DMSO was added to 96-well plate. ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+4nMAssay Description:Inhibition of bakers yeast alpha-glucosidase using p-nitrophenyl-alpha-D-glucopyranoside as substrate incubated for 15 mins followed by substrate add...More data for this Ligand-Target Pair
Affinity DataIC50: 1.75E+4nMAssay Description:At 37 °C, after incubation of 10 min of a reaction mixture solution of 70 μL (50 mM) phosphate buffer of 6.8 pH, 10 μL (0.5 mM) of a test c...More data for this Ligand-Target Pair
Affinity DataIC50: 1.82E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.82E+4nMpH: 6.8Assay Description:Total volume of 100 µL reaction mixture contained 70 µL 50 mM phosphate buffer pH 6.8, 10 µL test compound (0.5 mM in methanol) followed by the addit...More data for this Ligand-Target Pair
Affinity DataIC50: 1.82E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl-alpha-D-glucopyranoside as substrate preincubated for 10 mins measured a...More data for this Ligand-Target Pair
Affinity DataIC50: 1.83E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair