Report error Found 343 Enz. Inhib. hit(s) with Target = 'Alpha-2A/Alpha-2B/Alpha-2C adrenergic receptor'
Affinity DataKi: 0.25nMAssay Description:Binding affinity for Alpha-2 adrenergic receptor of CHO-C10 membrane preparationMore data for this Ligand-Target Pair
Affinity DataKi: 0.820nMAssay Description:Displacement of [3H]-RX821002 from adrenergic alpha 2 receptor in human brain membrane by liquid scintillation spectroscopyMore data for this Ligand-Target Pair
Affinity DataKi: 0.912nMAssay Description:Displacement of [3H]RX821002 from alpha2 adrenoceptor in human prefrontal cortex neural membrane after 30 mins by liquid scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 0.912nMAssay Description:Displacement of [3H]RX821002 from adrenergic alpha2 receptor in human brain frontal cortexMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Binding affinity for Alpha-2 adrenergic receptor of CHO-C10 membrane preparationMore data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Binding affinity towards Alpha-2 adrenergic receptor in HT-29 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Binding affinity for Alpha-2 adrenergic receptor of CHO-C10 membrane preparationMore data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:Displacement of [3H]RX821002 from alpha-2 adrenergic receptor in human brain pre-frontal cortex neural membranes after 30 mins by liquid scintillatio...More data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:Binding affinity to alpha2 adrenoceptor in human cortical membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 1.5nMAssay Description:Inhibition of [3H]p-aminoclonidine (PAC) binding to alpha-2 adrenergic receptor of purified human platelet plasma membranesMore data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Displacement of [3H]-RX821002 from adrenergic alpha 2 receptor in human brain membrane by liquid scintillation spectroscopyMore data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Displacement of [3H]RX821002 from adrenergic alpha2 receptor in human brain frontal cortexMore data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Displacement of [3H]-RX821002 from adrenergic alpha 2 receptor in human brain membrane by liquid scintillation spectroscopyMore data for this Ligand-Target Pair
Affinity DataKi: 2.40nMAssay Description:Binding affinity towards Alpha-2 adrenergic receptor of HT-29 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2.40nMAssay Description:Binding affinity for Alpha-2 adrenergic receptor of CHO-C10 membrane preparationMore data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of [3H]p-aminoclonidine (PAC) binding to alpha-2 adrenergic receptor of purified human platelet plasma membranesMore data for this Ligand-Target Pair
Affinity DataKi: 3.10nMAssay Description:Binding affinity for Alpha-2 adrenergic receptor of CHO-C10 membrane preparationMore data for this Ligand-Target Pair
Affinity DataKi: 3.80nMAssay Description:Displacement of [3H]RX821002 from human alpha2 adrenoceptor expressed in CHO cell membranes after 60 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of [3H]p-aminoclonidine (PAC) binding to alpha-2 adrenergic receptor of purified human platelet plasma membranesMore data for this Ligand-Target Pair
Affinity DataKi: 4.5nMAssay Description:Displacement of [3H]-RX821002 from adrenergic alpha 2 receptor in post-mortem human brain frontal cortex membrane measured after 30 mins by liquid sc...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Binding affinity against alpha 2-Adrenergic receptor in guinea pig cerebral cortical membranes by displacement of [3H]clonidineMore data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Binding affinity against alpha-2-adrenergic receptor using 10 nM [3H]yohimbine in human platelet membranes from three separate experiments using 10 i...More data for this Ligand-Target Pair
Affinity DataKi: 5.20nMAssay Description:Binding affinity for Alpha-2 adrenergic receptor of CHO-C10 membrane preparationMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Binding affinity towards Alpha-2 adrenergic receptor in HT-29 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 6.10nMAssay Description:Binding affinity against alpha-2-adrenergic receptor using 10 nM [3H]yohimbine in human platelet membranes from three separate experiments using 10 i...More data for this Ligand-Target Pair
Affinity DataKi: 6.80nMAssay Description:Binding affinity towards Alpha-2 adrenergic receptor in HT-29 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Binding affinity towards alpha 2 adrenergic receptorMore data for this Ligand-Target Pair
Affinity DataKi: 7.5nMAssay Description:Binding affinity towards human alpha-2 adrenergic receptorMore data for this Ligand-Target Pair
Affinity DataKi: 7.80nMAssay Description:Binding affinity for Alpha-2 adrenergic receptor of CHO-C10 membrane preparationMore data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:Inhibition of [3H]p-aminoclonidine (PAC) binding to alpha-2 adrenergic receptor of purified human platelet plasma membranesMore data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:Inhibition of [3H]p-aminoclonidine (PAC) binding to alpha-2 adrenergic receptor of purified human platelet plasma membranesMore data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Binding affinity against alpha 2-Adrenergic receptor in guinea pig cerebral cortical membranes by displacement of [3H]clonidineMore data for this Ligand-Target Pair
Affinity DataKi: 9.60nMAssay Description:Binding affinity for Alpha-2 adrenergic receptor of CHO-C10 membrane preparationMore data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Binding affinity towards alpha 2 adrenergic receptorMore data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [3H]-RX821002 from adrenergic alpha 2 receptor in human brain membrane by liquid scintillation spectroscopyMore data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Binding affinity against alpha 2-Adrenergic receptor in guinea pig cerebral cortical membranes by displacement of [3H]clonidineMore data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Binding affinity towards alpha 2 adrenergic receptorMore data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Binding affinity against alpha-2-adrenergic receptor using 10 nM [3H]yohimbine in human platelet membranes from three separate experiments using 10 i...More data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Displacement of [3H]RX821002 from adrenergic alpha2 receptor in human brain frontal cortexMore data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Binding affinity towards alpha 2 adrenergic receptorMore data for this Ligand-Target Pair
Affinity DataKi: 21nMAssay Description:Displacement of [3H]RX821002 from alpha2 adrenoceptor in human prefrontal cortex neural membrane after 30 mins by liquid scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataIC50: 21nMAssay Description:Inhibition of [3H]p-aminoclonidine (PAC) binding to alpha-2 adrenergic receptor of purified human platelet plasma membranesMore data for this Ligand-Target Pair
Affinity DataKi: 21nMAssay Description:Displacement of [3H]RX821002 from adrenergic alpha2 receptor in human brain frontal cortexMore data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:Binding affinity towards alpha 2 adrenergic receptorMore data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Binding affinity against alpha 2-Adrenergic receptor in guinea pig cerebral cortical membranes by displacement of [3H]clonidineMore data for this Ligand-Target Pair
Affinity DataKi: 31nMAssay Description:Displacement of [3H]RX821002 from alpha2 adrenergic receptor in human brain prefrontal cortex membranes after 30 mins by microbeta liquid scintillati...More data for this Ligand-Target Pair
Affinity DataKi: 34nMAssay Description:Displacement of [3H]-RX821002 from adrenergic alpha 2 receptor in human brain membrane by liquid scintillation spectroscopyMore data for this Ligand-Target Pair
Affinity DataKi: 41nMAssay Description:Binding affinity for Alpha-2 adrenergic receptor of CHO-C10 membrane preparationMore data for this Ligand-Target Pair
Affinity DataKi: 41nMAssay Description:Binding affinity for Alpha-2 adrenergic receptor of CHO-C10 membrane preparationMore data for this Ligand-Target Pair
Affinity DataKi: 42nMAssay Description:Displacement of [3H]RX821002 from alpha2 adrenoceptor in human prefrontal cortex neural membrane after 30 mins by liquid scintillation spectrometryMore data for this Ligand-Target Pair
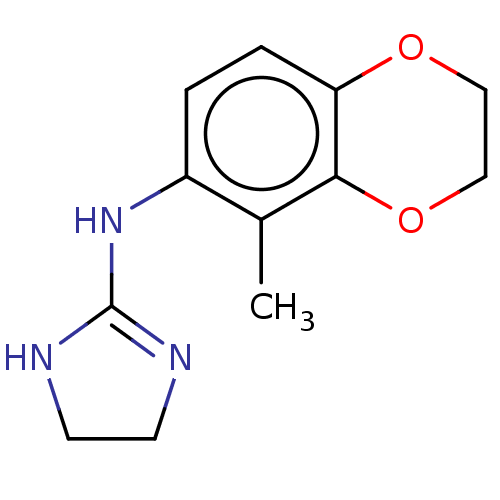
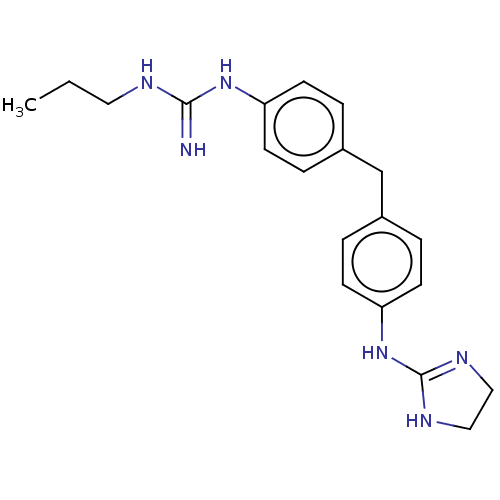
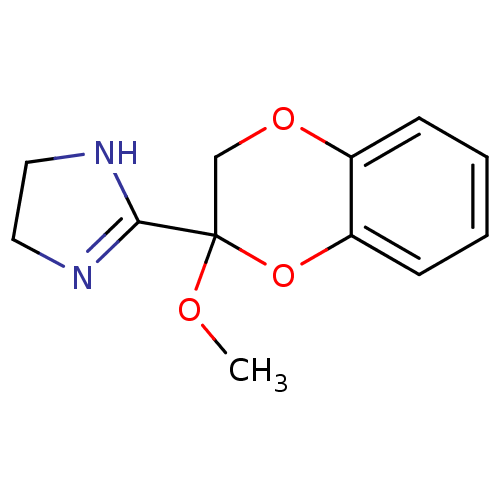


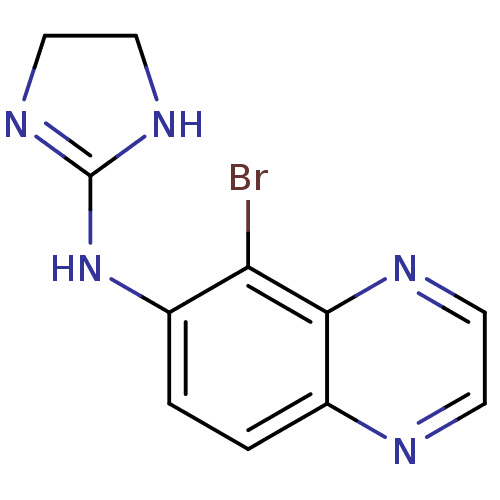
 3D Structure (crystal)
3D Structure (crystal)