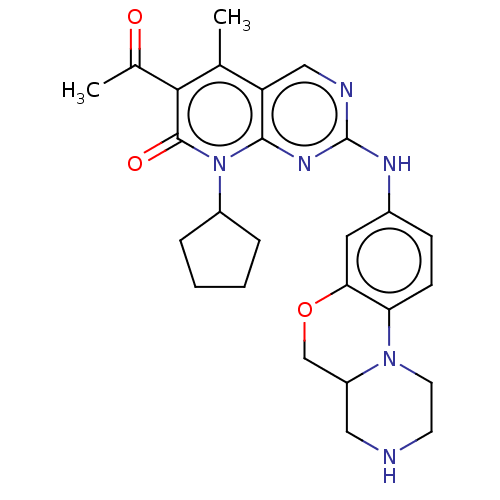
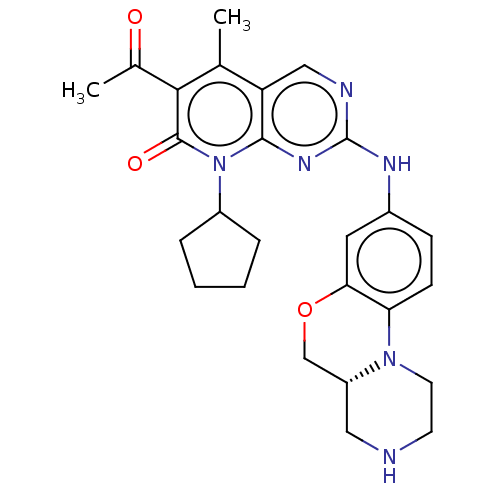
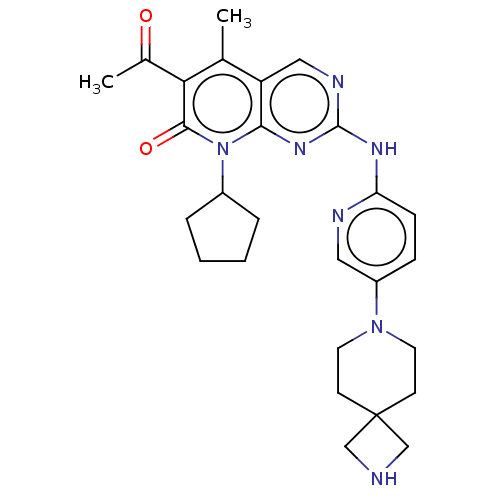
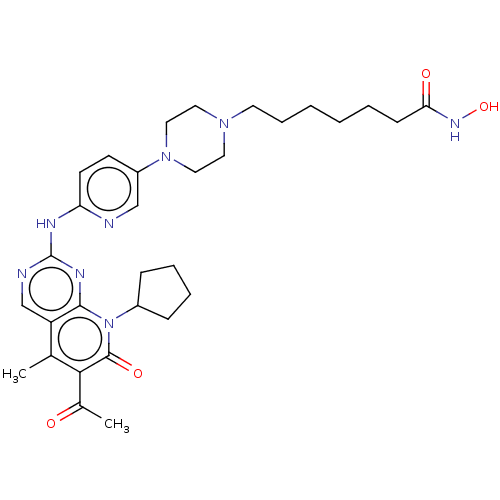
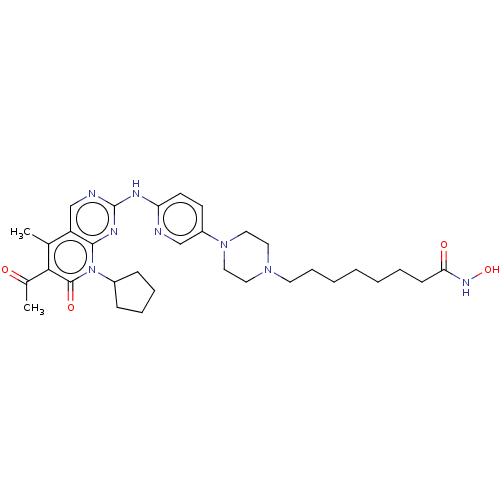
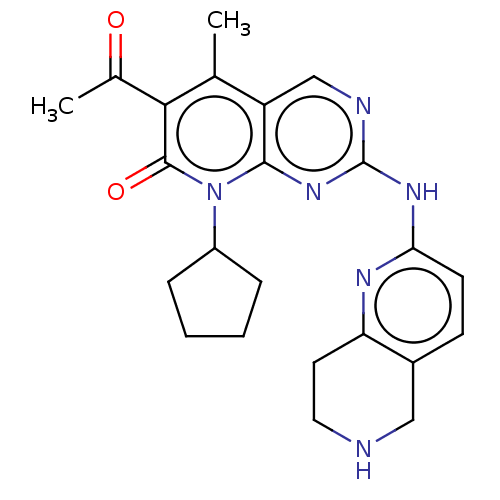
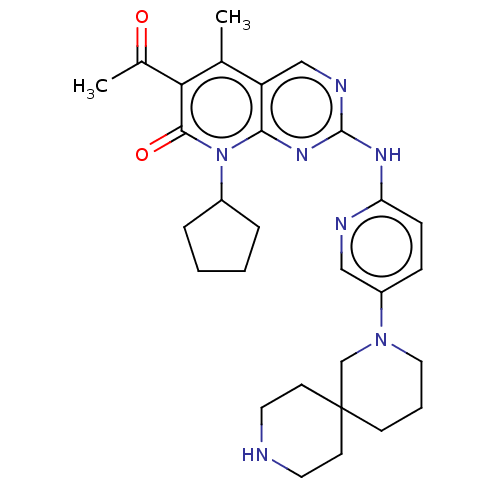
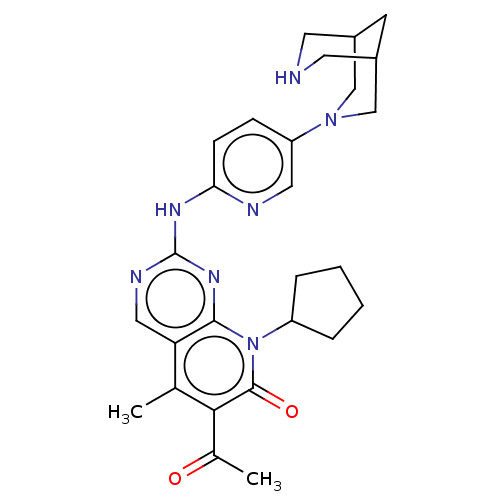
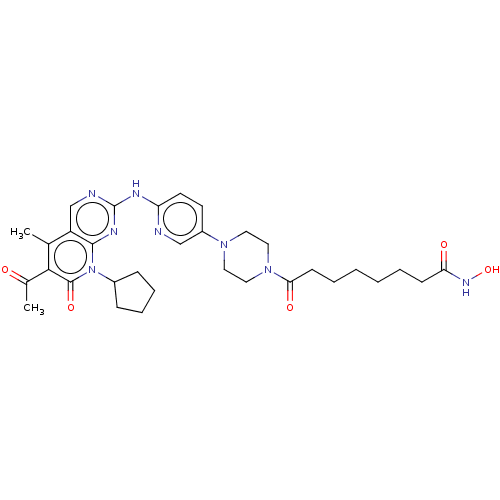
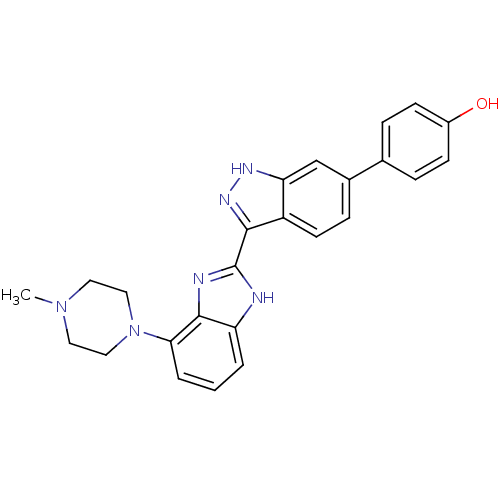
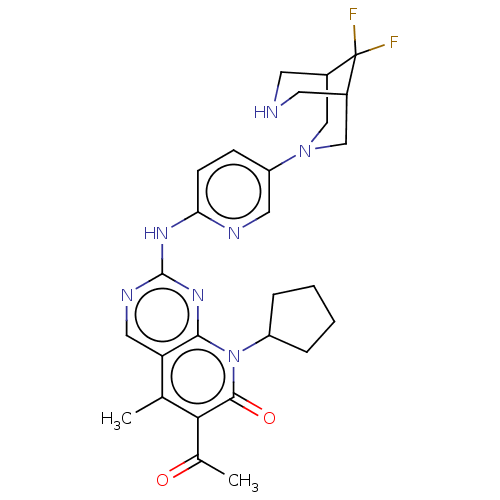
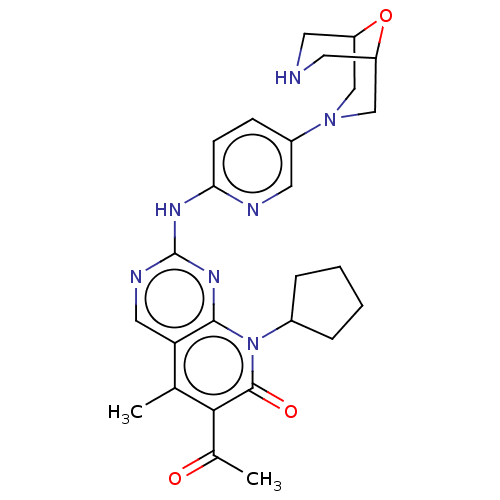
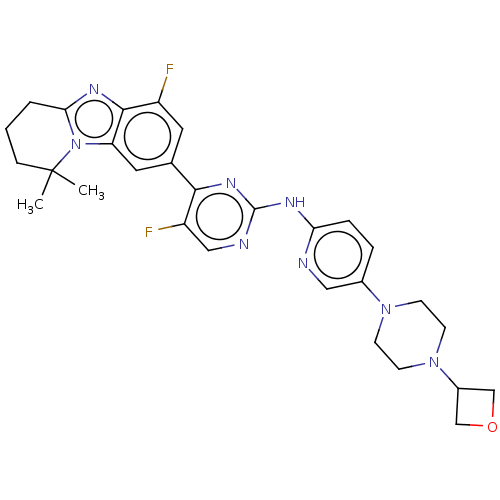
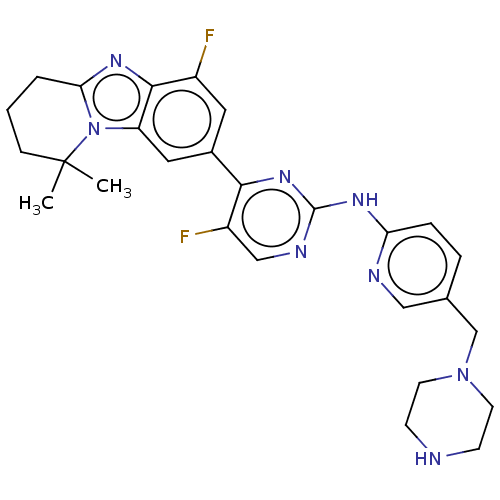
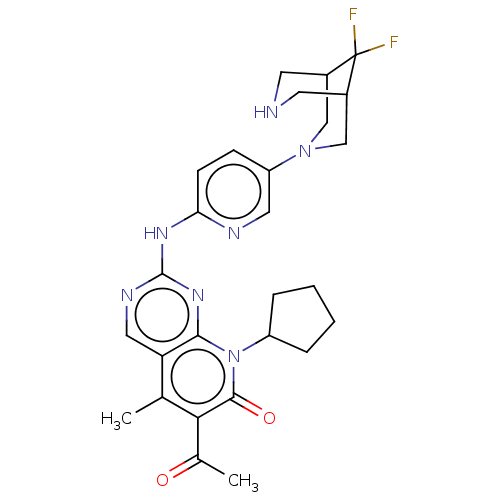
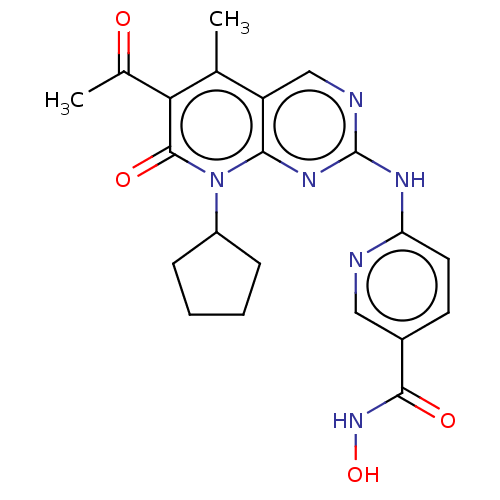
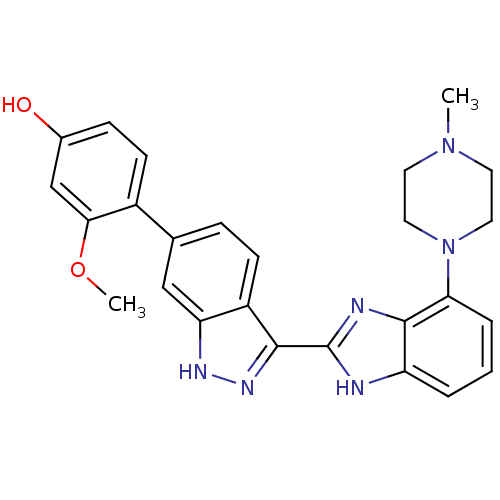
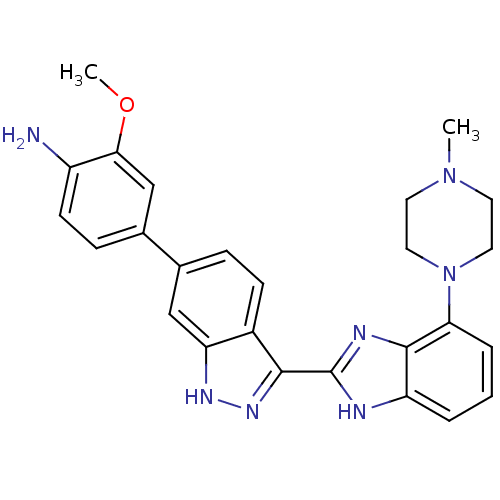
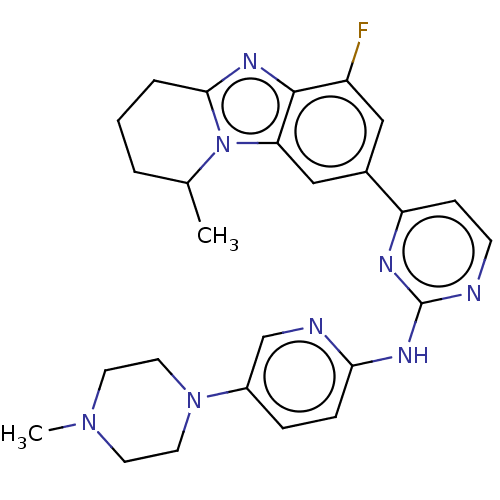
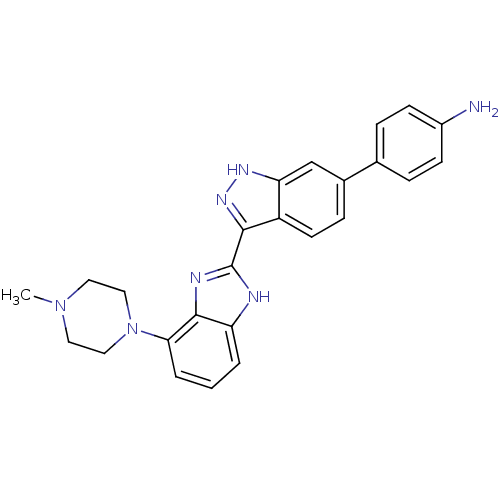
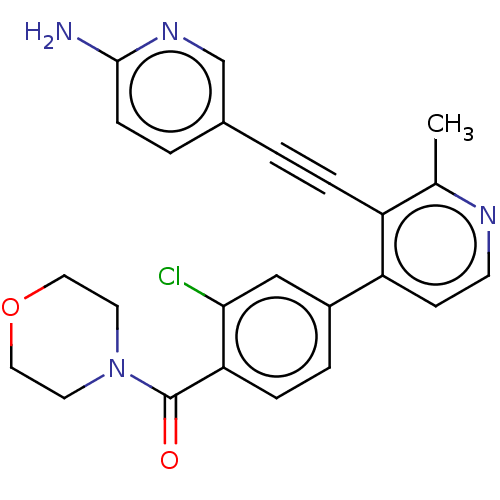
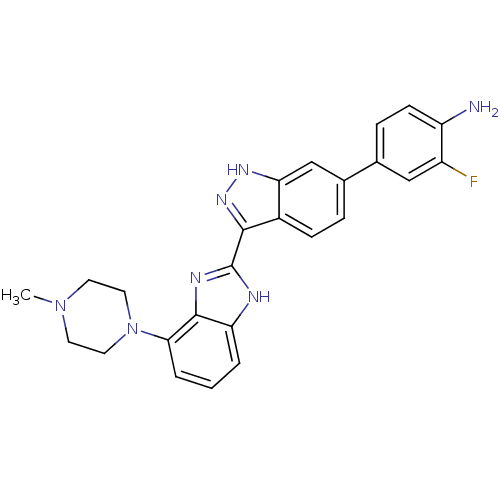
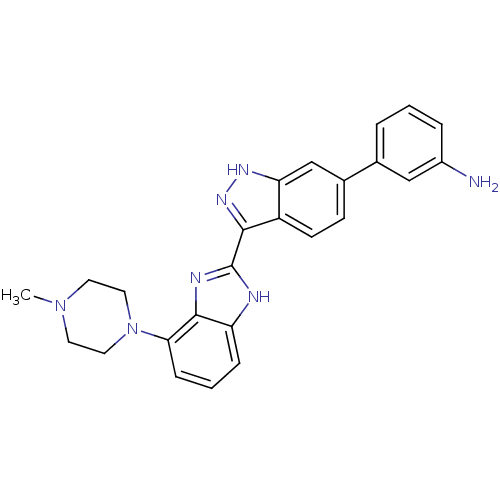
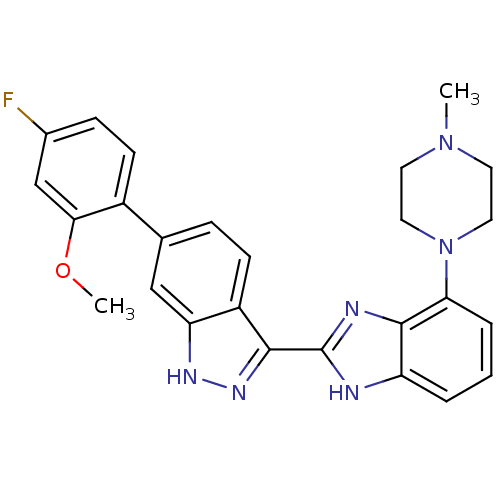
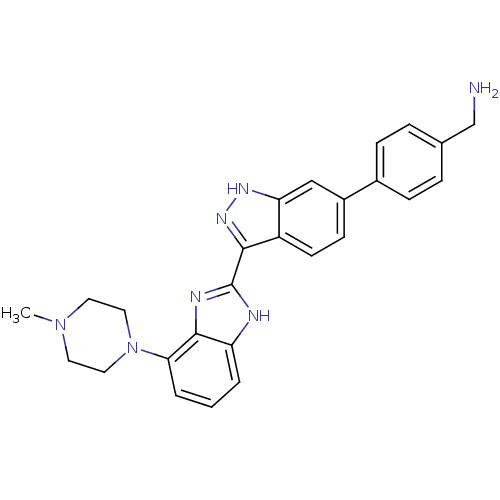
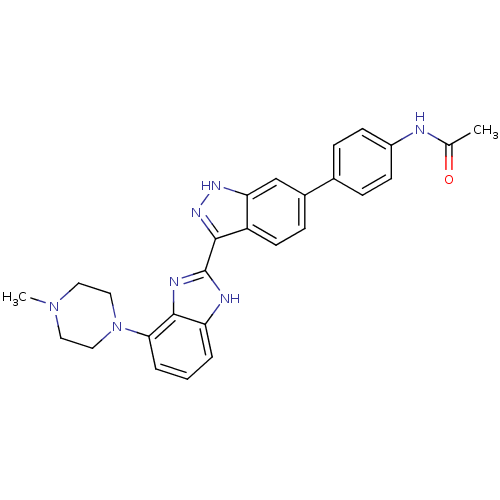
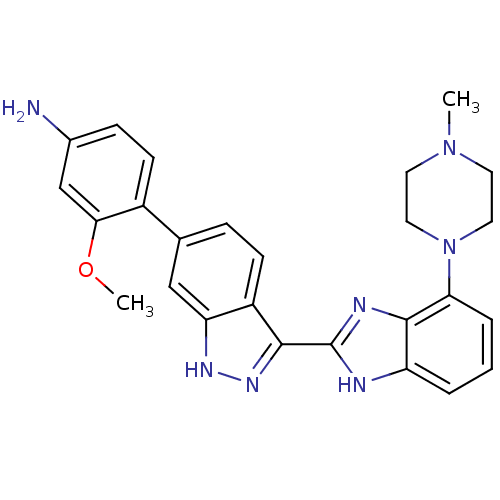
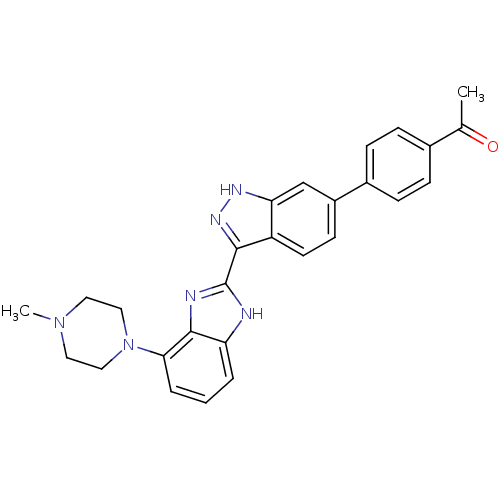
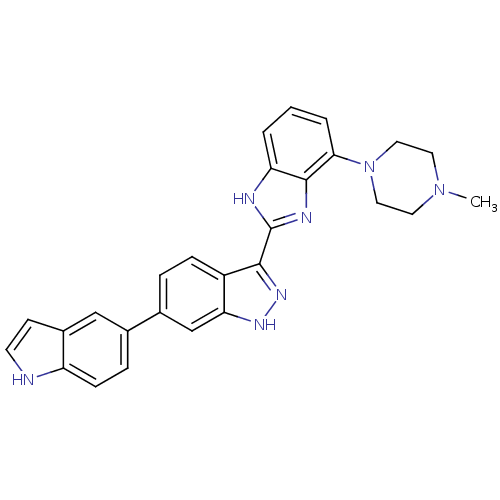
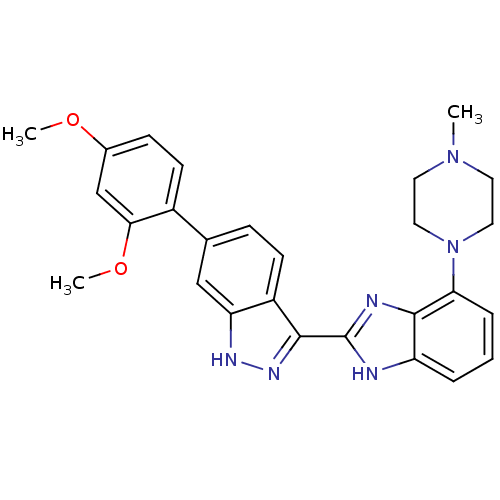
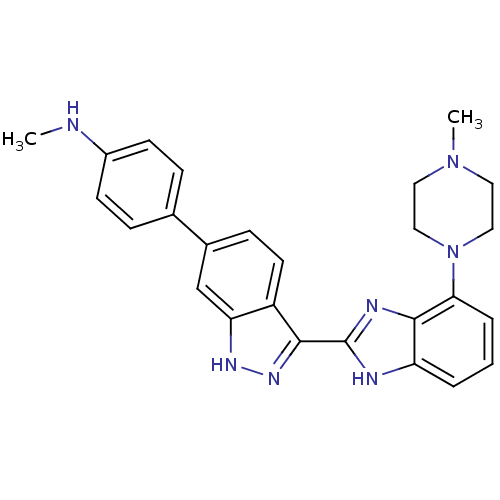
Report error Found 46 Enz. Inhib. hit(s) with Target = 'Cyclin-A2 [177-432]/Cyclin-dependent kinase 2'
Affinity DataIC50: 4.10nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 13.3nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 39nMAssay Description:To demonstrate that the compounds exhibit affinity for CDK kinases (CDK2/CycA2, CDK4/CycD3, CDK6/cycD3), CDK kinase assays were performed.Reaction bu...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:To demonstrate that the compounds exhibit affinity for CDK kinases (CDK2/CycA2, CDK4/CycD3, CDK6/cycD3), CDK kinase assays were performed.Reaction bu...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:To demonstrate that the compounds exhibit affinity for CDK kinases (CDK2/CycA2, CDK4/CycD3, CDK6/cycD3), CDK kinase assays were performed.Reaction bu...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:In vitro enzymatic activity of the CDK isoforms CDK2/CycA2, CDK4/CycD3 and CDK6/cycD3 were measured using Mobility Shift Assay that monitors phosphor...More data for this Ligand-Target Pair
Affinity DataIC50: 102nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 247nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:To demonstrate that the compounds exhibit affinity for CDK kinases (CDK2/CycA2, CDK4/CycD3, CDK6/cycD3), CDK kinase assays were performed.Reaction bu...More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:To demonstrate that the compounds exhibit affinity for CDK kinases (CDK2/CycA2, CDK4/CycD3, CDK6/cycD3), CDK kinase assays were performed.Reaction bu...More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:To demonstrate that the compounds exhibit affinity for CDK kinases (CDK2/CycA2, CDK4/CycD3, CDK6/cycD3), CDK kinase assays were performed.Reaction bu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.04E+3nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMT: 2°CAssay Description:The GSK3beta primary screen was conducted in assay ready 1536 plates (Aurora 29847) that contain 2.5 mL/well of 10 mM compound. Human GSK3beta as a G...More data for this Ligand-Target Pair
Affinity DataIC50: 1.73E+3nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.90E+3nMT: 2°CAssay Description:The GSK3beta primary screen was conducted in assay ready 1536 plates (Aurora 29847) that contain 2.5 mL/well of 10 mM compound. Human GSK3beta as a G...More data for this Ligand-Target Pair
Affinity DataIC50: 6.10E+3nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.15E+4nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.09E+5nMT: 2°CAssay Description:Protein kinase C (PKC), zeta activity was measured by following the phosphorylation of a biotinylated peptide substrate in the presence of ATP using ...More data for this Ligand-Target Pair