Report error Found 36 Enz. Inhib. hit(s) with Target = 'Gamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2'
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
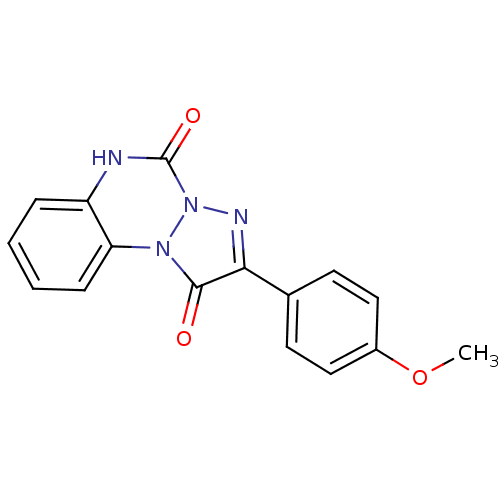
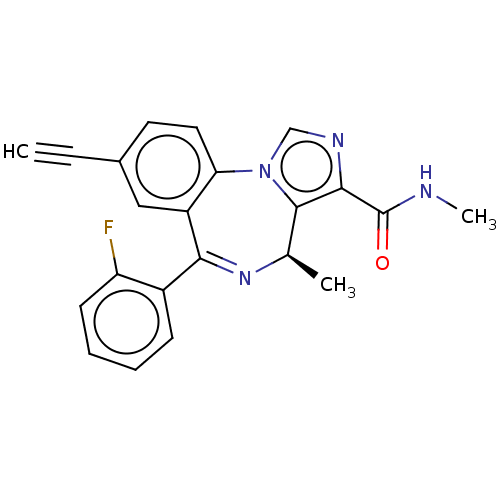
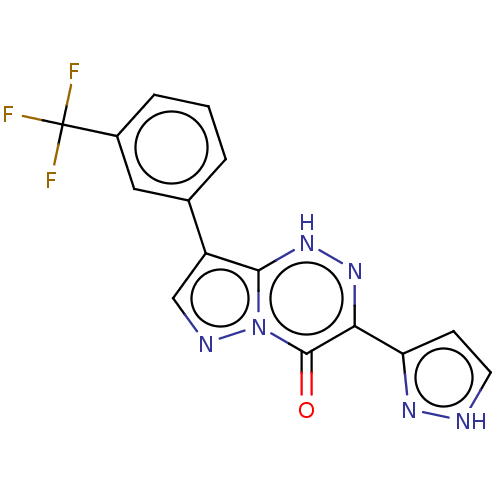
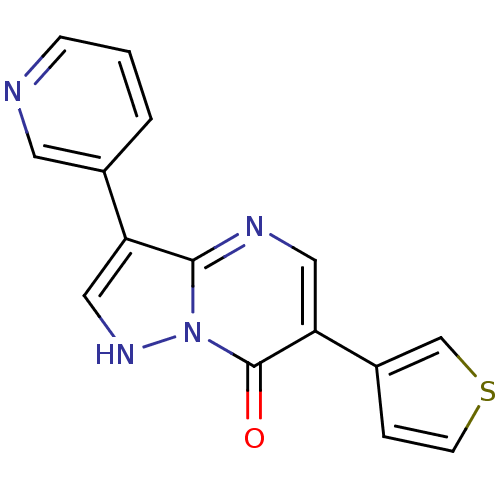
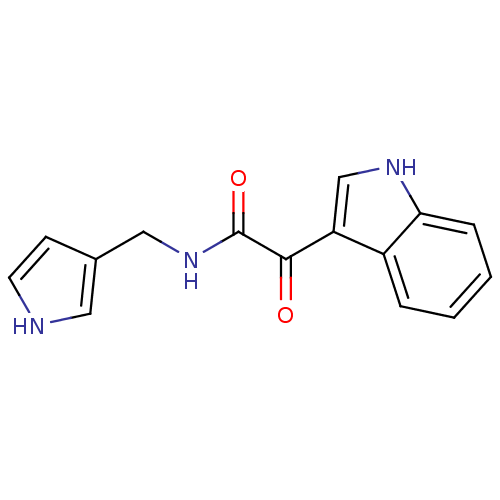
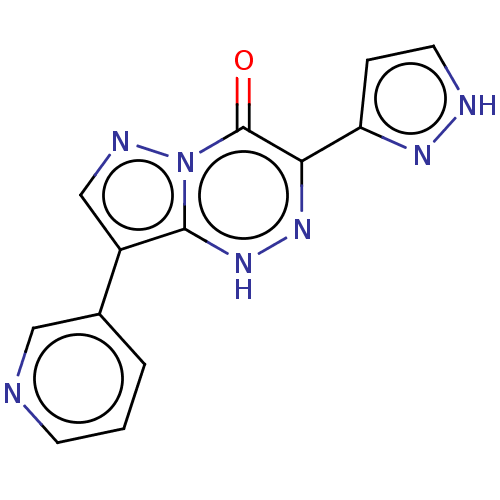
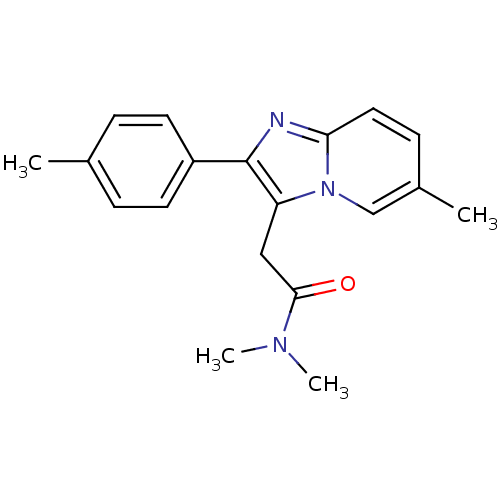
Affinity DataKi: 7.57nMAssay Description:Displacement of [3H]flumazenil from rat GABA-A Gamma-aminobutyric acid A receptor alpha-5-beta-3-gamma-2 expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
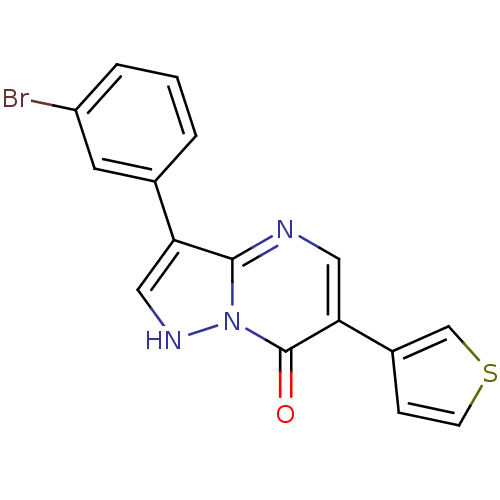
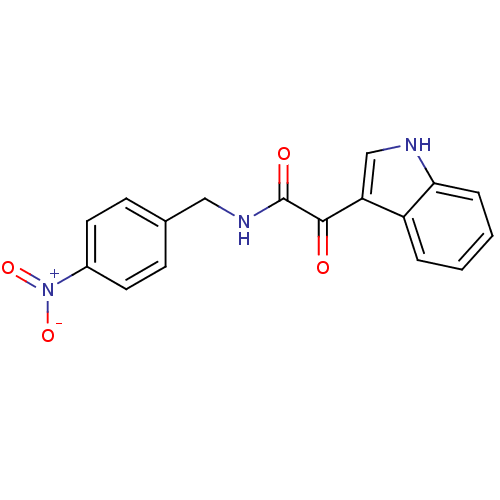
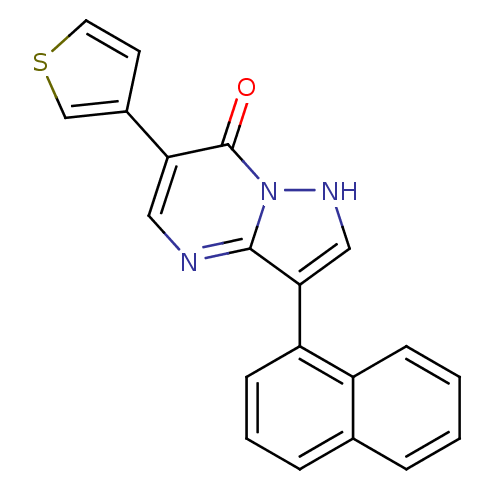
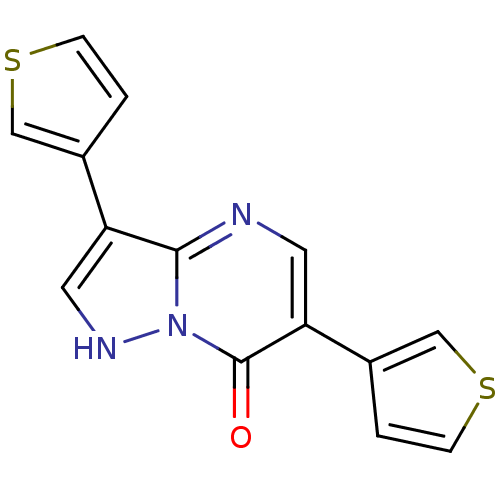
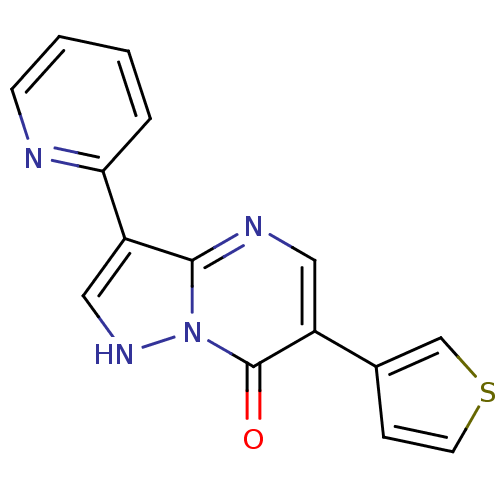
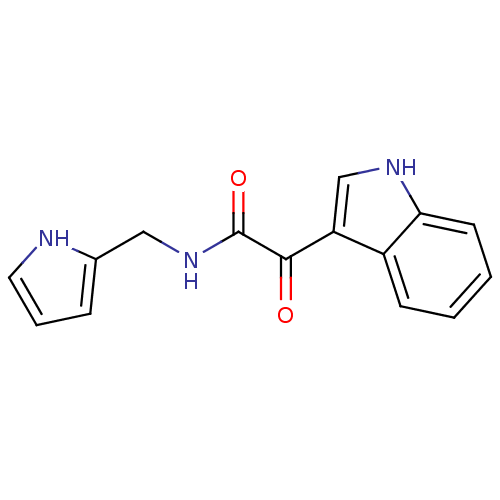
Affinity DataKi: 10.3nMAssay Description:Displacement of [3H]flumazenil from rat GABA-A Gamma-aminobutyric acid A receptor alpha-5-beta-3-gamma-2 expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
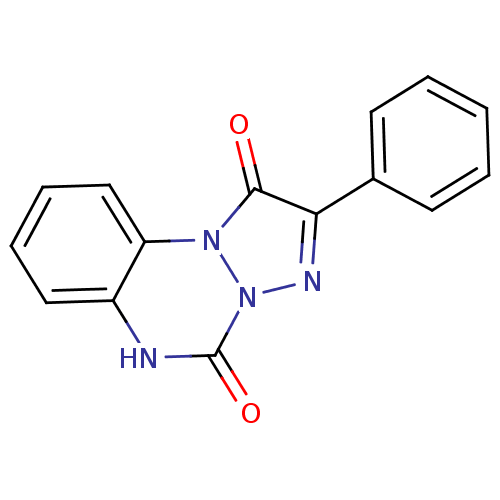
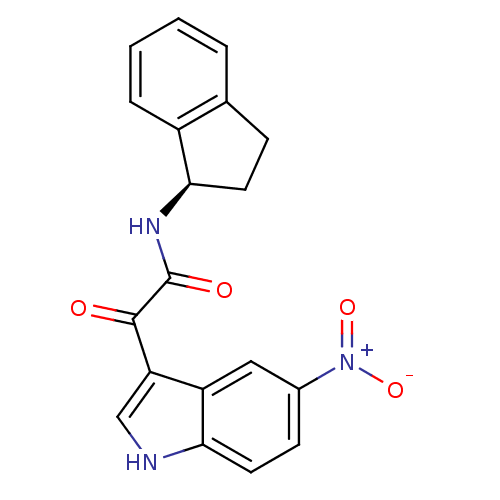
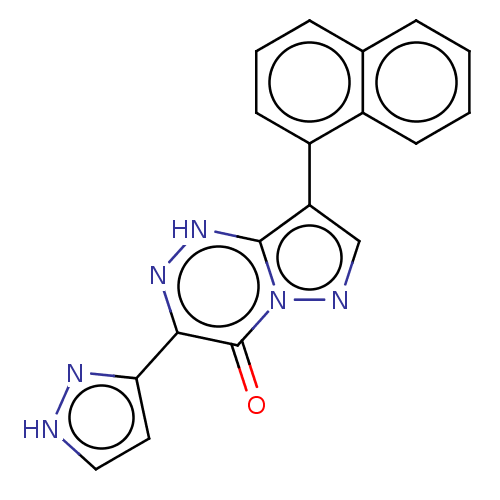
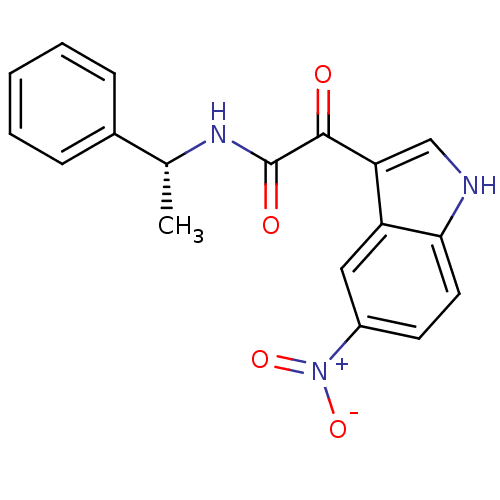
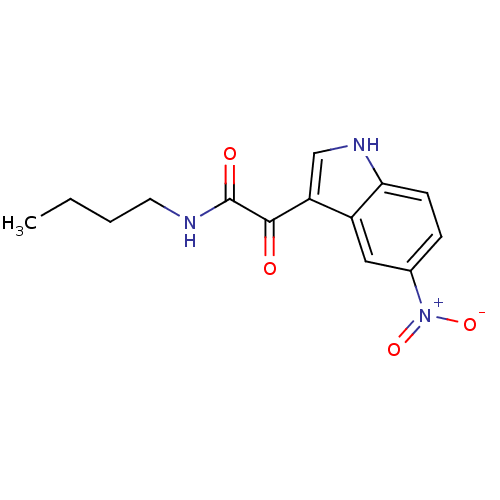
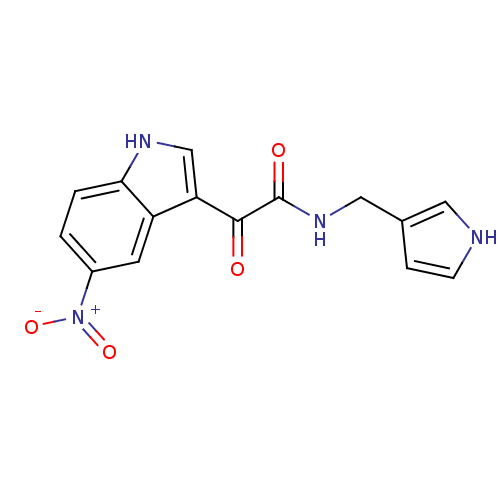
Affinity DataKi: 11nMAssay Description:Binding affinity for recombinant rat GABA-A receptor alpha-5-beta-3-gamma-2 subunits expressed in HEK 293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
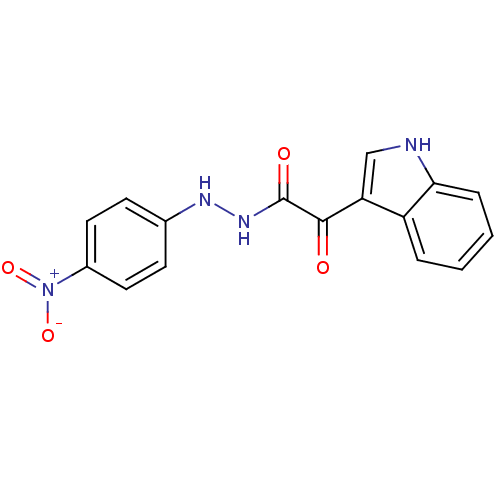
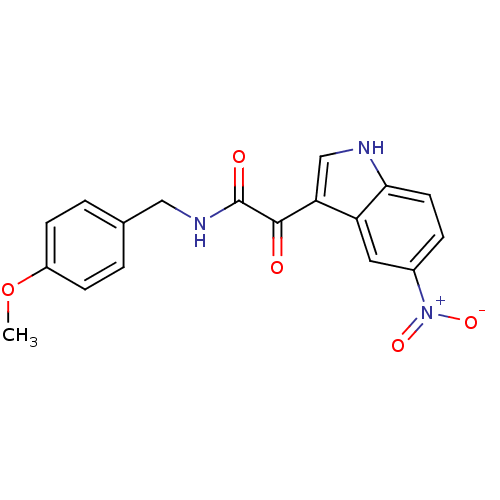
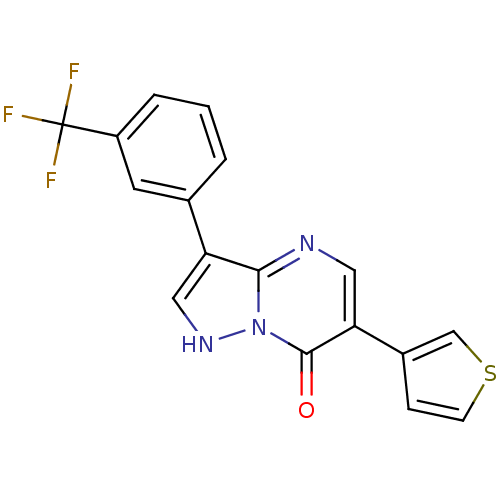
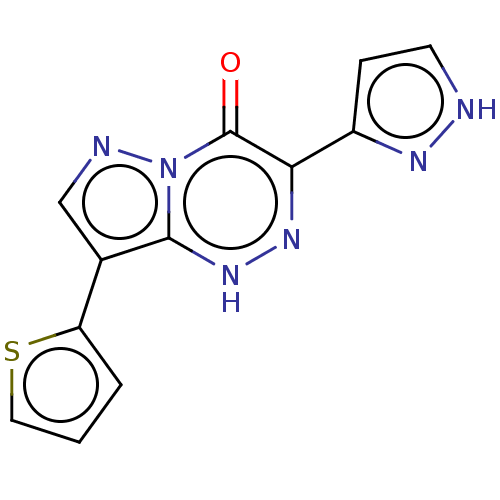
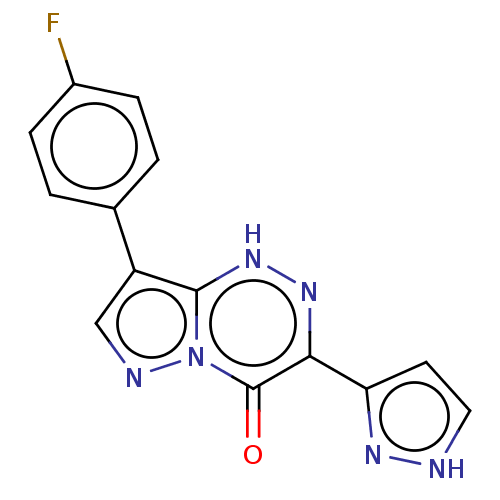
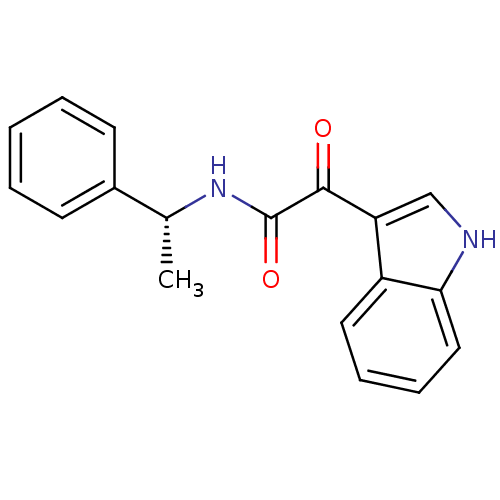
Affinity DataKi: 17.2nMAssay Description:Displacement of [3H]flumazenil from rat GABA-A Gamma-aminobutyric acid A receptor alpha-5-beta-3-gamma-2 expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 22nMAssay Description:Binding affinity for recombinant rat GABA-A receptor alpha-5-beta-3-gamma-2 subunits expressed in HEK 293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 24.1nMAssay Description:Displacement of [3H]flumazenil from rat GABA-A Gamma-aminobutyric acid A receptor alpha-5-beta-3-gamma-2 expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 43nMAssay Description:Inhibition of [3H]flumazenil binding to rat alpha-5-beta-3-gamma-2 GABA-A receptor subunits expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 55nMAssay Description:Displacement of [3H]-flunitrazepam from recombinant rat GABAA receptor alpha5beta3gamma2S expressed in HEK293 cell membranes after 90 mins by liquid ...More data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 58nMAssay Description:Inhibition of [3H]flumazenil binding to rat alpha-5-beta-3-gamma-2 GABA-A receptor subunits expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 95nMAssay Description:Inhibition of [3H]flumazenil binding to rat alpha-5-beta-3-gamma-2 GABA-A receptor subunits expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 126nMAssay Description:Inhibition of [3H]flumazenil binding to rat alpha-5-beta-3-gamma-2 GABA-A receptor subunits expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 144nMAssay Description:Displacement of [3H]Ro15-1788 from recombinant rat GABAAalpha5/beta3/gamma2 receptor after 90 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 156nMAssay Description:Binding affinity for recombinant rat GABA-A receptor alpha-5-beta-3-gamma-2 subunits expressed in HEK 293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 159nMAssay Description:Displacement of [3H]Ro15-1788 from recombinant rat GABAAalpha5/beta3/gamma2 receptor after 90 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 207nMAssay Description:Binding affinity for recombinant rat GABA-A receptor alpha-5-beta-3-gamma-2 subunits expressed in HEK 293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 210nMAssay Description:Binding affinity for recombinant rat GABA-A receptor alpha-5-beta-3-gamma-2 subunits expressed in HEK 293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 223nMAssay Description:Binding affinity for recombinant rat GABA-A receptor alpha-5-beta-3-gamma-2 subunits expressed in HEK 293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 239nMAssay Description:Inhibition of [3H]flumazenil binding to rat alpha-5-beta-3-gamma-2 GABA-A receptor subunits expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 408nMAssay Description:Displacement of [3H]Ro15-1788 from recombinant rat GABAAalpha5/beta3/gamma2 receptor after 90 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 498nMAssay Description:Displacement of [3H]flumazenil from rat GABAA alpha-5-beta-3-gamma-2 expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 662nMAssay Description:Displacement of [3H]Ro15-1788 from recombinant rat GABAAalpha5/beta3/gamma2 receptor after 90 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 681nMAssay Description:Displacement of [3H]flumazenil from rat GABA-Aalpha5 receptor plus beta-3-gamma-2 in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 737nMAssay Description:Binding affinity for recombinant rat GABA-A receptor alpha-5-beta-3-gamma-2 subunits expressed in HEK 293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 740nMAssay Description:Binding affinity for recombinant rat GABA-A receptor alpha-5-beta-3-gamma-2 subunits expressed in HEK 293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 943nMAssay Description:Displacement of [3H]flumazenil from rat GABAA alpha-5-beta-3-gamma-2 expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 1.11E+3nMAssay Description:Displacement of [3H]Ro15-1788 from recombinant rat GABAAalpha5/beta3/gamma2 receptor after 90 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 1.12E+3nMAssay Description:Displacement of [3H]Ro15-1788 from recombinant rat GABAAalpha5/beta3/gamma2 receptor after 90 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 1.15E+3nMAssay Description:Displacement of [3H]flumazenil from rat GABAA alpha-5-beta-3-gamma-2 expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 1.24E+3nMAssay Description:Displacement of [3H]Ro15-1788 from recombinant rat GABAAalpha5/beta3/gamma2 receptor after 90 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 1.45E+3nMAssay Description:Displacement of [3H]flumazenil from rat GABAA alpha-5-beta-3-gamma-2 expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 1.50E+3nMAssay Description:Displacement of [3H]flumazenil from rat GABA-Aalpha5 receptor plus beta-3-gamma-2 in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 1.60E+3nMAssay Description:Displacement of [3H]flumazenil from rat GABA-Aalpha5 receptor plus beta-3-gamma-2 in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 2.16E+3nMAssay Description:Inhibition of [3H]flumazenil binding to rat alpha-5-beta-3-gamma-2 GABA-A receptor subunits expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: 5.50E+3nMAssay Description:Inhibition of [3H]flumazenil binding to rat alpha-5-beta-3-gamma-2 GABA-A receptor subunits expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Binding affinity for recombinant rat GABA-A receptor alpha-5-beta-3-gamma-2 subunits expressed in HEK 293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-5/beta-3/gamma-2(Rat)
Università
Curated by ChEMBL
Università
Curated by ChEMBL
Affinity DataEC50: >2.40E+4nMAssay Description:Displacement of [3H]-flumazenil from rat GABA-A receptor alpha5beta3gamma2 expressed in HEK293 cells by electrophysiology assayMore data for this Ligand-Target Pair


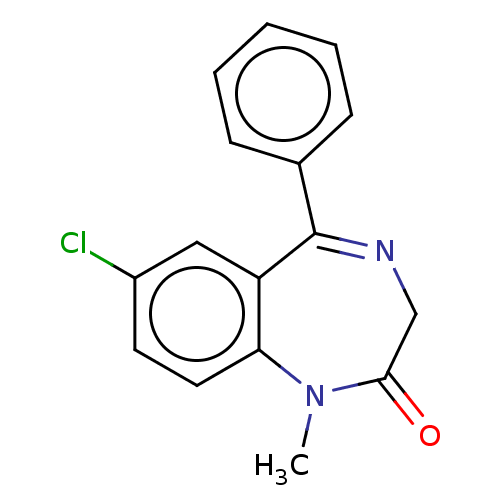
 3D Structure (crystal)
3D Structure (crystal)