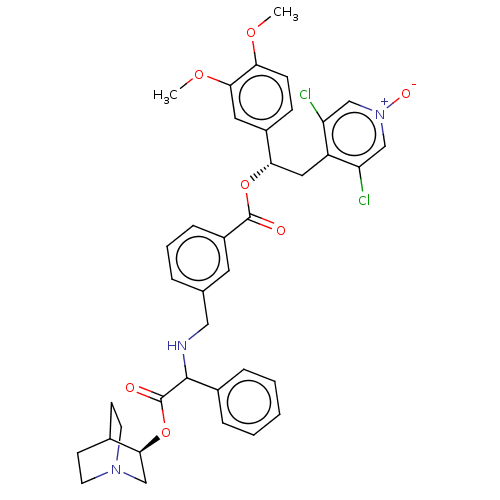
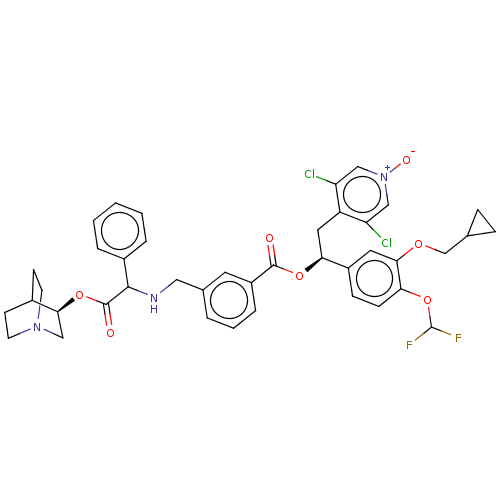
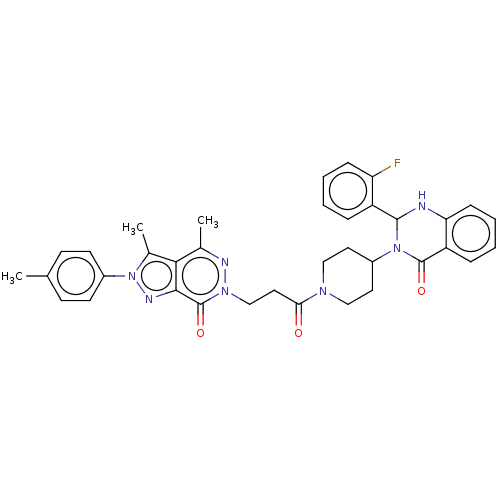
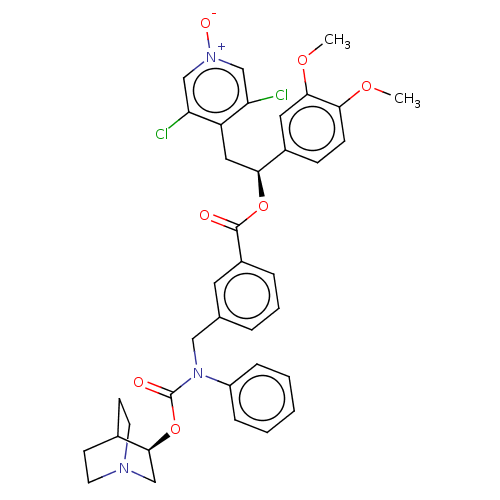
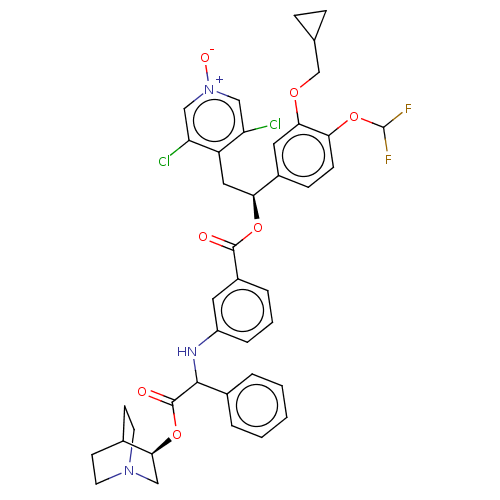
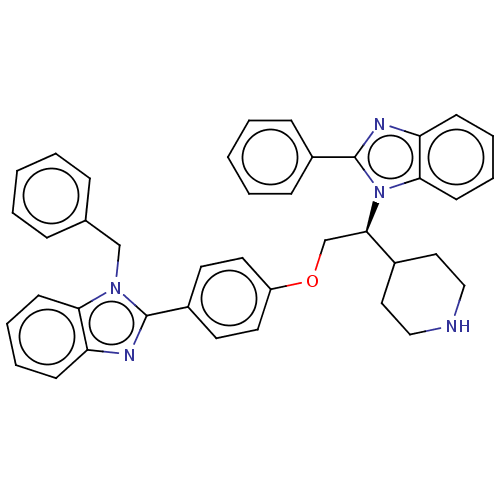
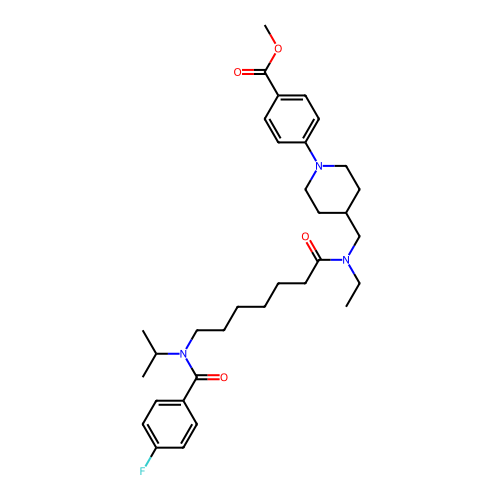
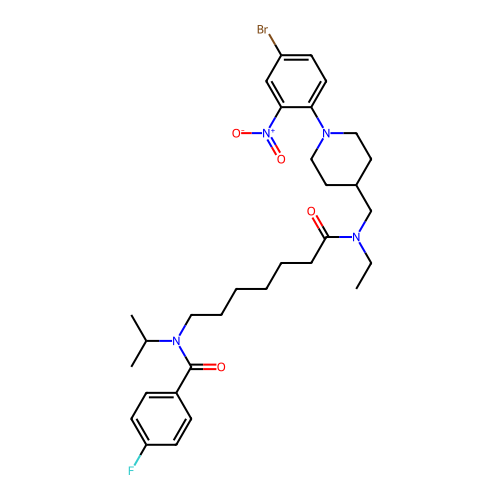
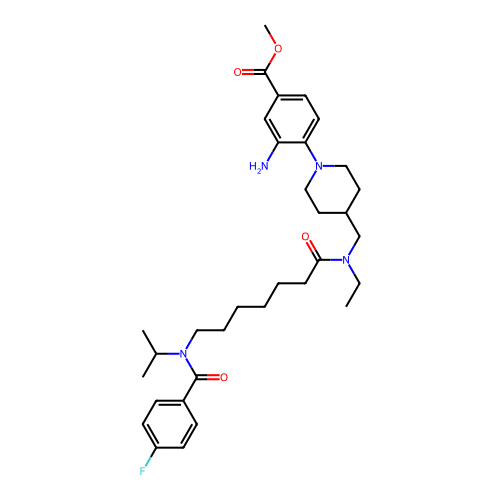
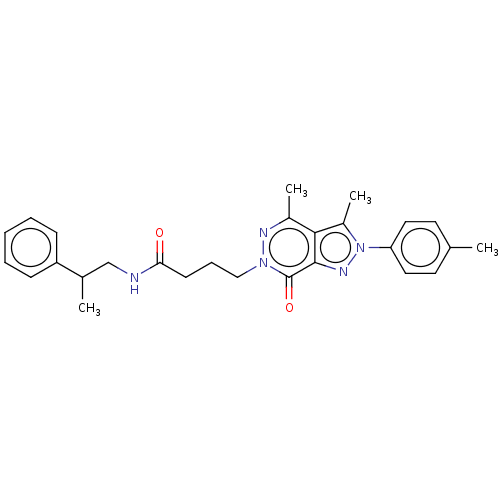
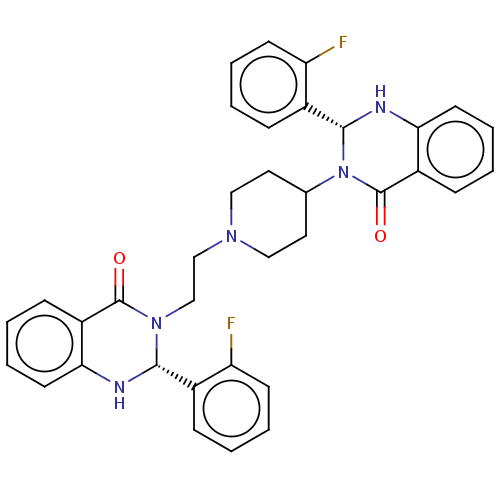
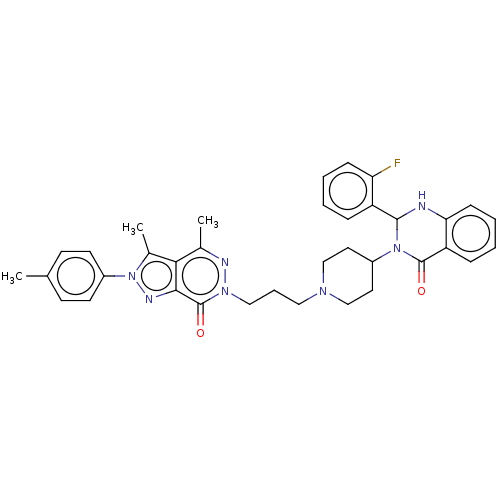
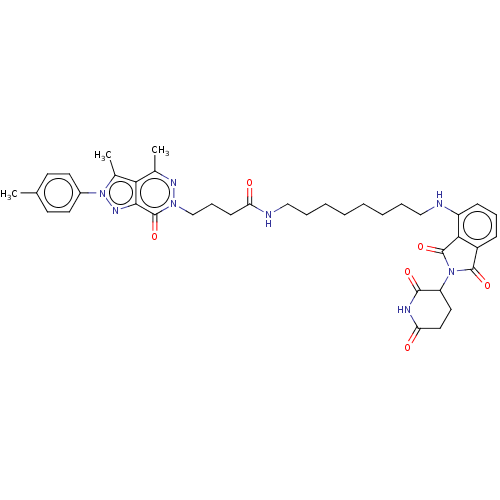
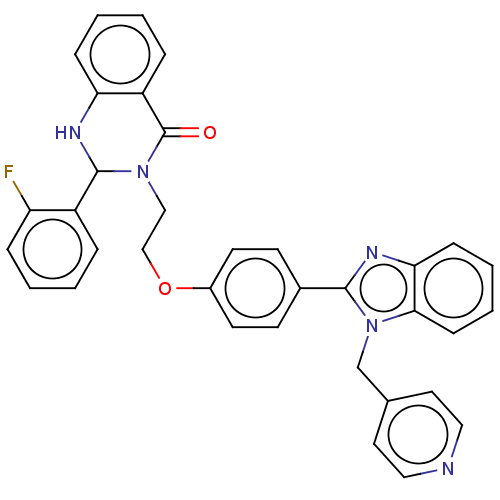
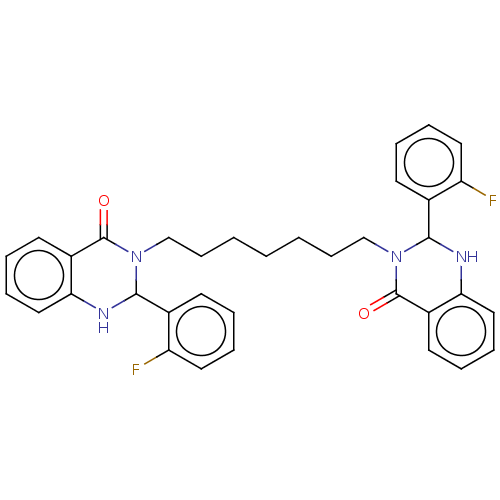
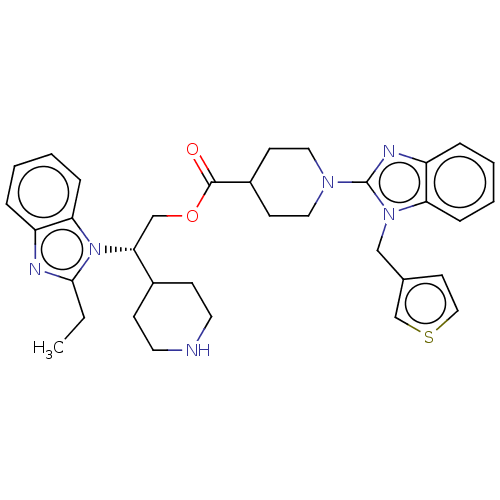
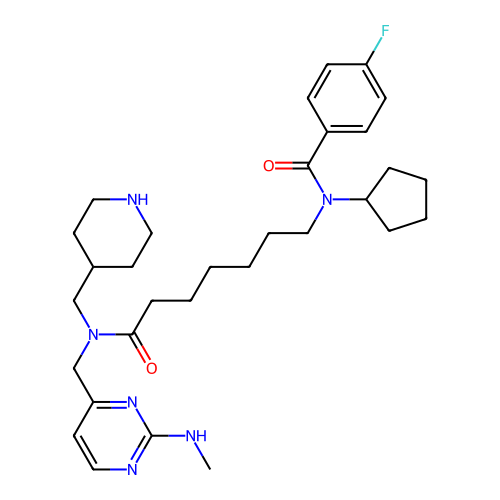
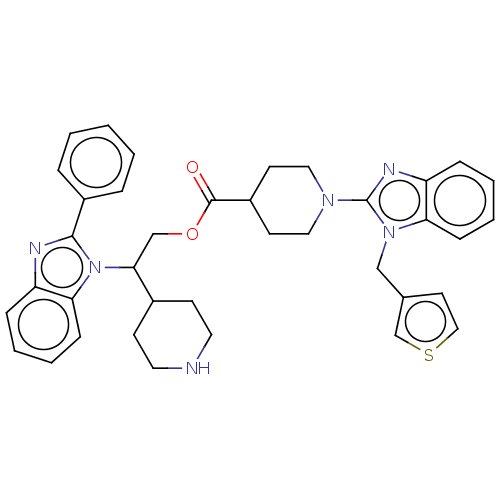
Report error Found 337 Enz. Inhib. hit(s) with Target = 'Retinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta'
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
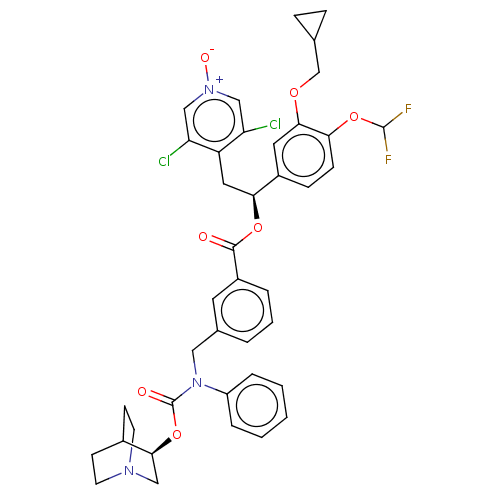
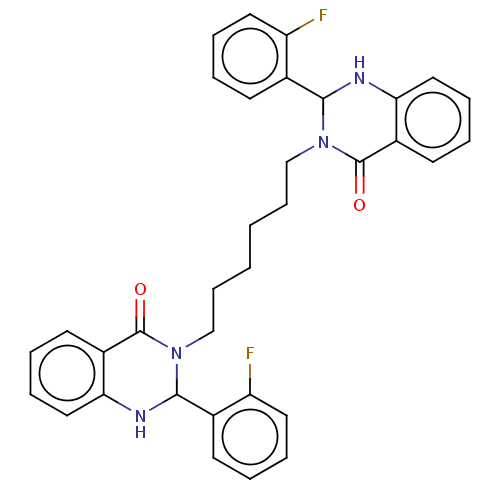
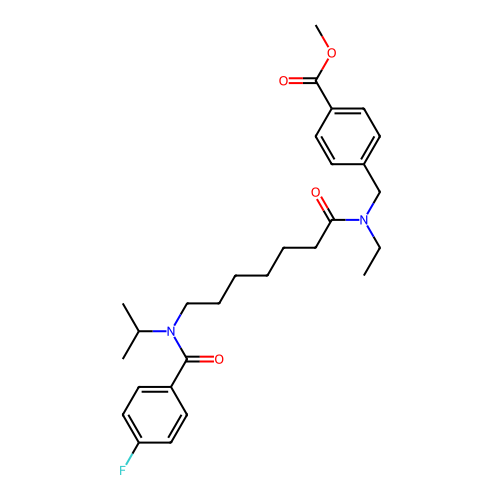
Affinity DataKd: 0.0770nMAssay Description:Binding affinity to PDE-delta (unknown origin) by SPR analysisMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
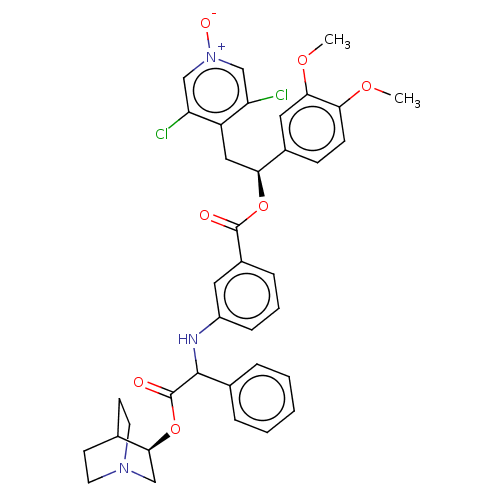
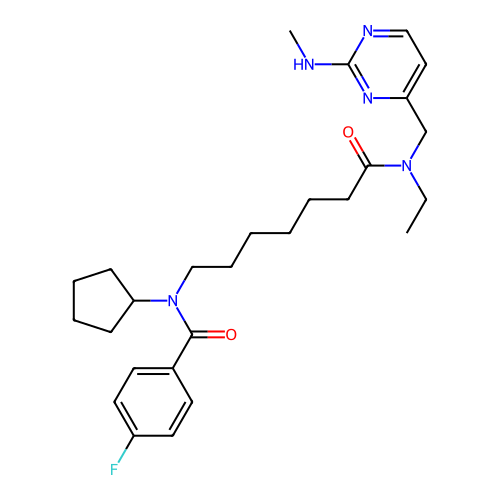
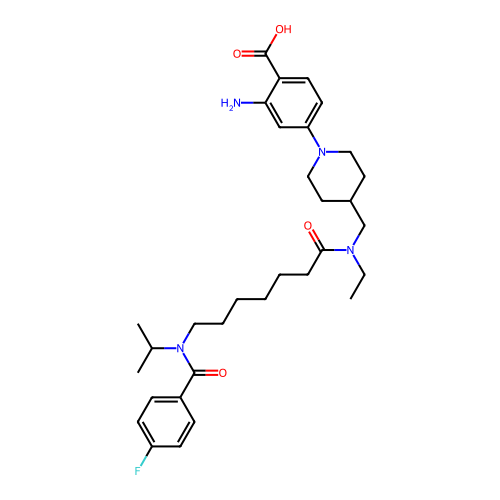
Affinity DataIC50: 0.100nMAssay Description:Inhibition of PDE4 in human U-937 cells assessed as reduction in cAMP level by LANCE cAMP AssayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
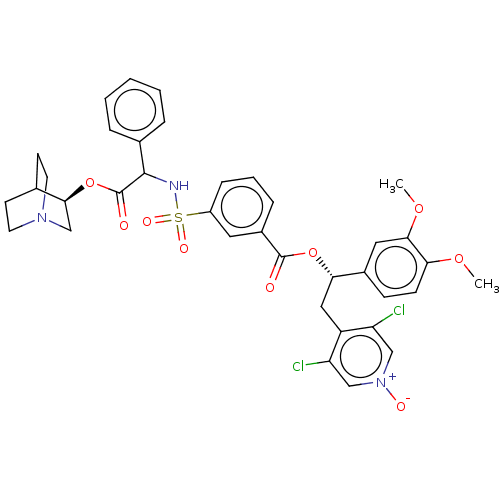
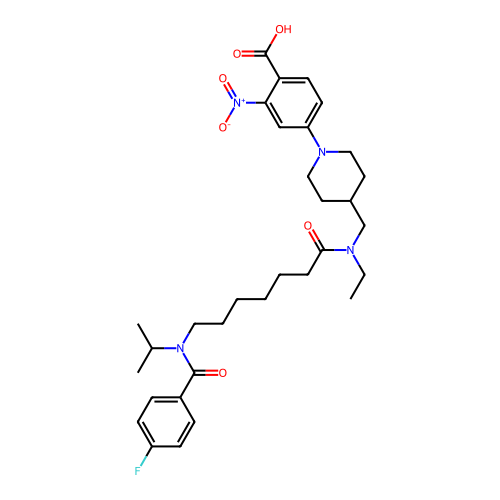
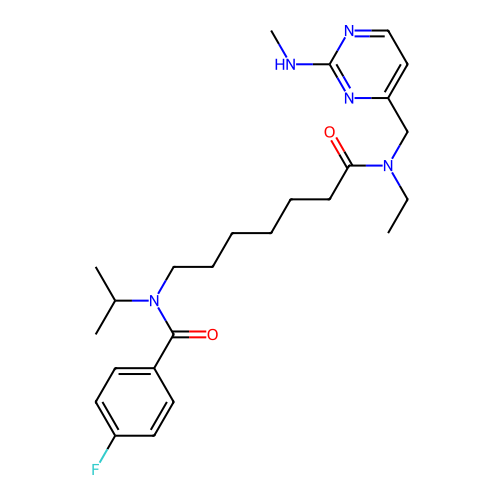
Affinity DataKd: 0.110nMAssay Description:Displacement of fluorescein-labeled Atorvastatin from N-terminal His6 tagged human recombinant PDE6D extracted from Escherichia coli BL21 star (DE3) ...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
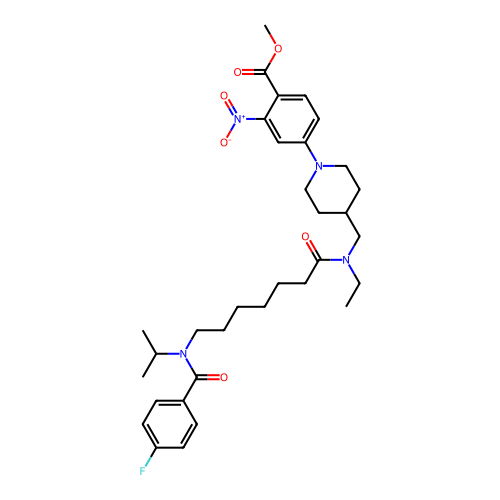
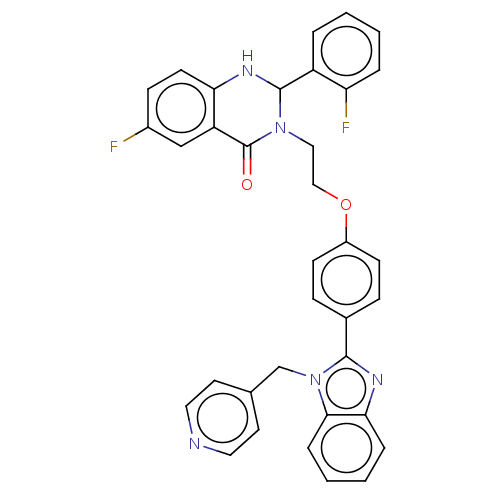
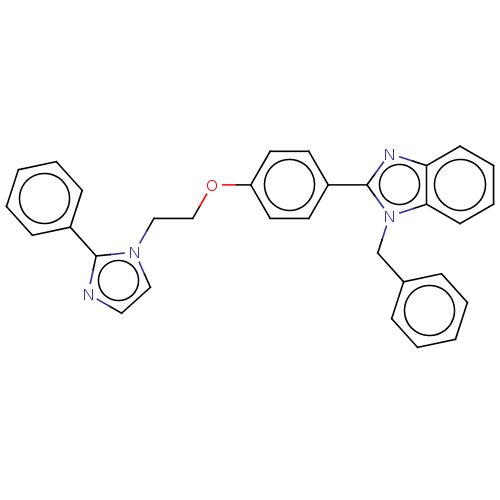
Affinity DataIC50: 0.126nMAssay Description:Inhibition of PDE4 in human U-937 cells assessed as reduction in cAMP level by LANCE cAMP AssayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataIC50: 0.158nMAssay Description:Inhibition of PDE4 in human U-937 cells assessed as reduction in cAMP level by LANCE cAMP AssayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 0.203nMAssay Description:Binding affinity to PDEdelta (unknown origin)More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 0.203nMAssay Description:Binding affinity to PDEdelta (unknown origin)More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataIC50: 0.251nMAssay Description:Inhibition of PDE4 in human U-937 cells assessed as reduction in cAMP level by LANCE cAMP AssayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 0.300nMAssay Description:Displacement of fluorescein-labeled Atorvastatin from N-terminal His6 tagged human recombinant PDE6D extracted from Escherichia coli BL21 star (DE3) ...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataIC50: 0.316nMAssay Description:Inhibition of PDE4 in human U-937 cells assessed as reduction in cAMP level by LANCE cAMP AssayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataIC50: 0.316nMAssay Description:Inhibition of PDE4 in human U-937 cells assessed as reduction in cAMP level by LANCE cAMP AssayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 0.600nMAssay Description:Inhibition of Atrovastatin-PEG3-FITC binding to PDEdelta (unknown origin) incubated for 60 mins by fluorescence anisotropy assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Inhibition of PDE4 in human U-937 cells assessed as reduction in cAMP level by LANCE cAMP AssayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Inhibition of PDE4 in human U-937 cells assessed as reduction in cAMP level by LANCE cAMP AssayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 1.10nMAssay Description:Binding affinity to PDE-delta (unknown origin) by time-resolved fluorescence anisotropic analysisMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 1.30nMAssay Description:Displacement of fluorescein-labeled Atorvastatin from N-terminal His6 tagged human recombinant PDE6D extracted from Escherichia coli BL21 star (DE3) ...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 1.5nMAssay Description:Displacement of fluorescein-labeled Atorvastatin from N-terminal His6 tagged human recombinant PDE6D extracted from Escherichia coli BL21 star (DE3) ...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 1.60nMAssay Description:Displacement of fluorescein-labeled Atorvastatin from N-terminal His6 tagged human recombinant PDE6D extracted from Escherichia coli BL21 star (DE3) ...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 2nMAssay Description:Inhibition of Atrovastatin-PEG3-FITC binding to PDEdelta (unknown origin) incubated for 60 mins by fluorescence anisotropy assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 2nMAssay Description:Inhibition of Atrovastatin-PEG3-FITC binding to PDEdelta (unknown origin) incubated for 60 mins by fluorescence anisotropy assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 2.20nMAssay Description:Inhibition of Atrovastatin-PEG3-FITC binding to PDEdelta (unknown origin) incubated for 60 mins by fluorescence anisotropy assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 2.30nMAssay Description:Inhibition of Atrovastatin-PEG3-FITC binding to PDEdelta (unknown origin) incubated for 60 mins by fluorescence anisotropy assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 3.80nMAssay Description:Displacement of fluorescein-labeled Atorvastatin from N-terminal His6 tagged human recombinant PDE6D extracted from Escherichia coli BL21 star (DE3) ...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 4nMAssay Description:Inhibition of Atrovastatin-PEG3-FITC binding to PDEdelta (unknown origin) incubated for 60 mins by fluorescence anisotropy assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 4nMAssay Description:Inhibition of Atrovastatin-PEG3-FITC binding to PDEdelta (unknown origin) incubated for 60 mins by fluorescence anisotropy assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 4nMAssay Description:Displacement of fluorescein-labeled Atorvastatin from N-terminal His6 tagged human recombinant PDE6D extracted from Escherichia coli BL21 star (DE3) ...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 4.5nMAssay Description:Displacement of fluorescein-labeled Atorvastatin from N-terminal His6 tagged human recombinant PDE6D extracted from Escherichia coli BL21 star (DE3) ...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 4.5nMAssay Description:Inhibition of atrovastatin-PEG3-FITC binding to PDEdelta (unknown origin) incubated for 60 mins by fluorescence anisotropy assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 6nMAssay Description:Displacement of fluorescein-labeled Atorvastatin from N-terminal His6 tagged human recombinant PDE6D extracted from Escherichia coli BL21 star (DE3) ...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 6nMAssay Description:Displacement of fluorescein-labeled Atorvastatin from N-terminal His6 tagged human recombinant PDE6D extracted from Escherichia coli BL21 star (DE3) ...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 6nMAssay Description:Displacement of fluorescein-labeled Atorvastatin from N-terminal His6 tagged human recombinant PDE6D extracted from Escherichia coli BL21 star (DE3) ...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 6nMAssay Description:Binding affinity to His6-tagged PDE-delta (unknown origin) measured every 3 mins by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 7nMAssay Description:Binding affinity to His6-tagged PDE-delta (unknown origin) measured every 3 mins by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 7nMAssay Description:Binding affinity to His6-tagged PDE-delta (unknown origin) measured every 3 mins by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 7nMAssay Description:Binding affinity to His6-tagged PDE-delta (unknown origin) measured every 3 mins by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 7.20nMAssay Description:Displacement of fluorescein-labeled Atorvastatin from N-terminal His6 tagged human recombinant PDE6D extracted from Escherichia coli BL21 star (DE3) ...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 8nMAssay Description:Inhibition of atrovastatin-PEG3-FITC binding to PDEdelta (unknown origin) incubated for 60 mins by fluorescence anisotropy assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 8nMAssay Description:Binding affinity to PDEdelta (unknown origin) assessed as dissociation constant incubated for overnight by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 8nMAssay Description:Inhibition of Atrovastatin-PEG3-FITC binding to PDEdelta (unknown origin) incubated for 60 mins by fluorescence anisotropy assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 8nMAssay Description:Inhibition of Atrovastatin-PEG3-FITC binding to PDEdelta (unknown origin) incubated for 60 mins by fluorescence anisotropy assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKi: 8nMAssay Description:Binding affinity to PDE delta (unknown origin) assessed as dissociation constant measured after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 8nMAssay Description:Inhibition of Atrovastatin-PEG3-FITC binding to PDEdelta (unknown origin) incubated for 60 mins by fluorescence anisotropy assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 8nMAssay Description:Binding affinity to His6-tagged PDE-delta (unknown origin) measured every 3 mins by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKi: 8.60nMAssay Description:Binding affinity to PDE delta (unknown origin) assessed as dissociation constant measured after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 9nMAssay Description:Inhibition of Atrovastatin-PEG3-FITC binding to PDEdelta (unknown origin) incubated for 60 mins by fluorescence anisotropy assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 9nMAssay Description:Inhibition of atrovastatin-PEG3-FITC binding to PDEdelta (unknown origin) incubated for 60 mins by fluorescence anisotropy assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 9nMAssay Description:Inhibition of Atrovastatin-PEG3-FITC binding to PDEdelta (unknown origin) incubated for 60 mins by fluorescence anisotropy assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 9nMAssay Description:Binding affinity to His6-tagged PDE-delta (unknown origin) measured every 3 mins by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 9nMAssay Description:Displacement of fluorescein-labeled Atorvastatin from N-terminal His6 tagged human recombinant PDE6D extracted from Escherichia coli BL21 star (DE3) ...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Max Planck Institute of Molecular Physiology
Curated by ChEMBL
Affinity DataKd: 9nMAssay Description:Binding affinity to His6-tagged PDE-delta (unknown origin) measured every 3 mins by fluorescence polarization assayMore data for this Ligand-Target Pair
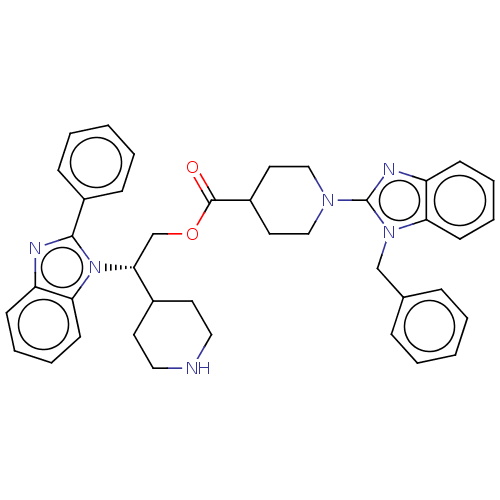
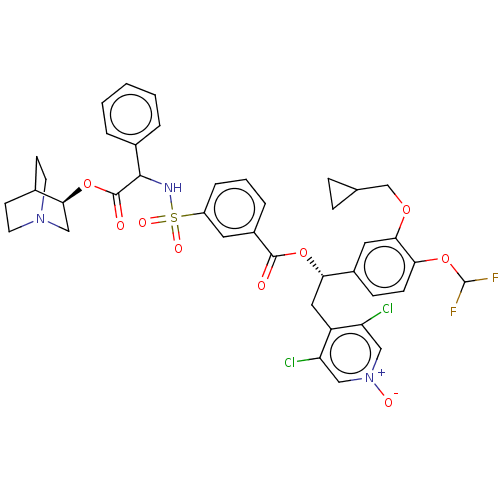
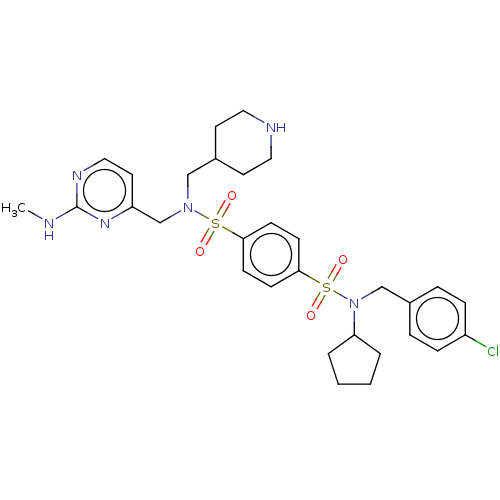
 3D Structure (crystal)
3D Structure (crystal)