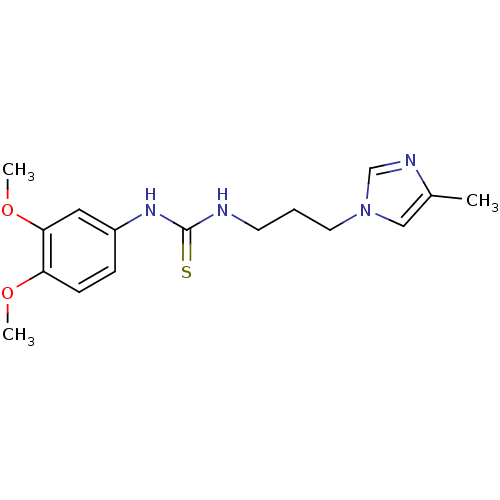
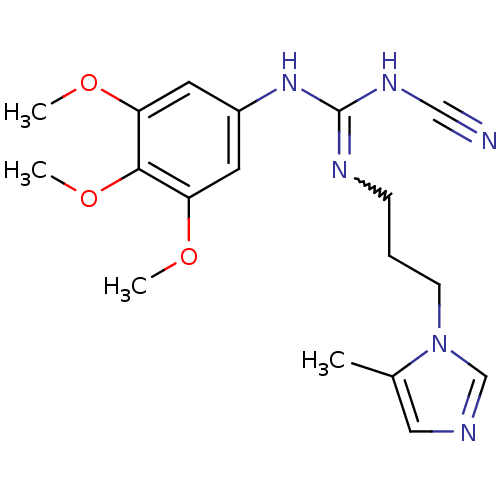
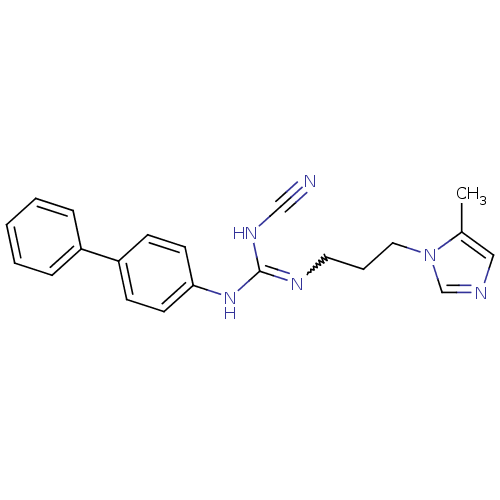
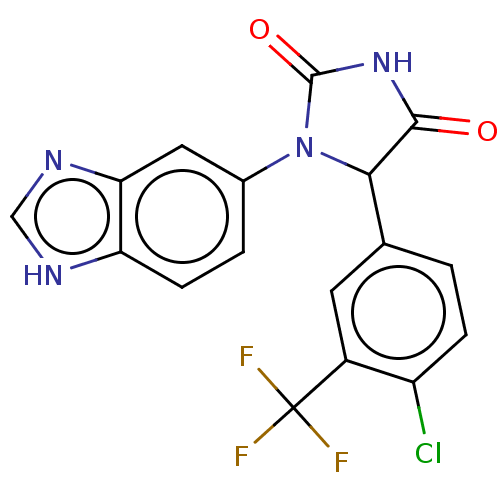
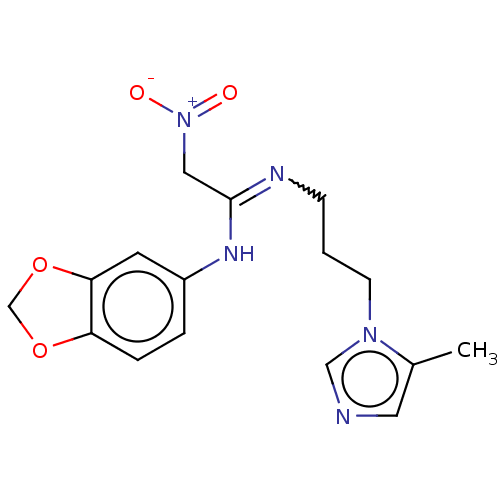
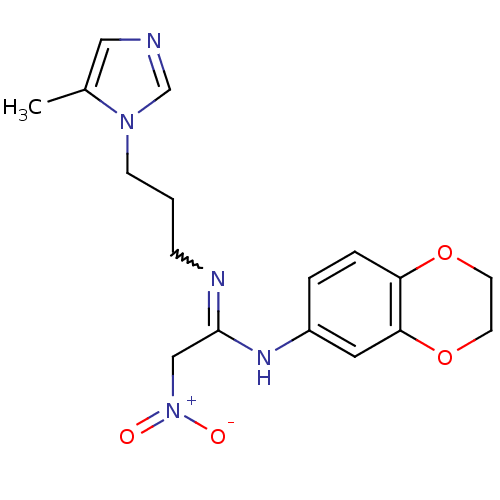
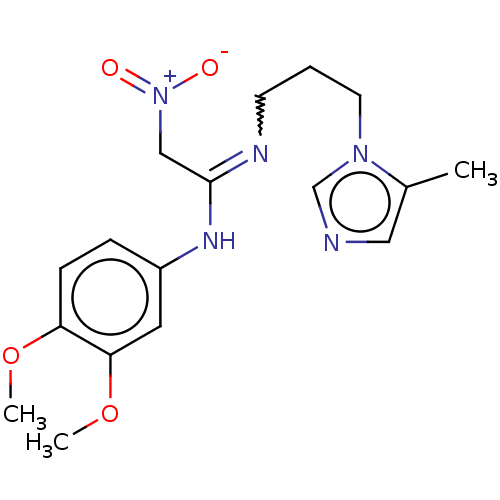
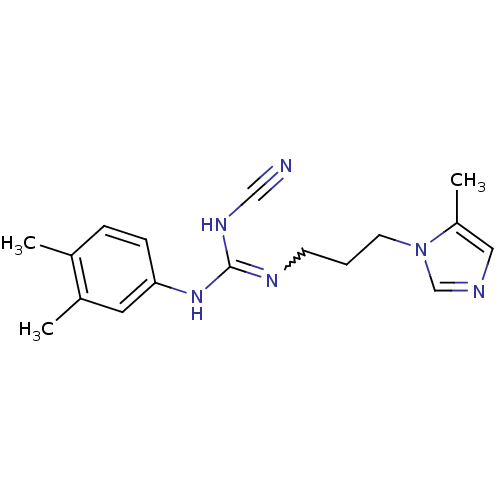
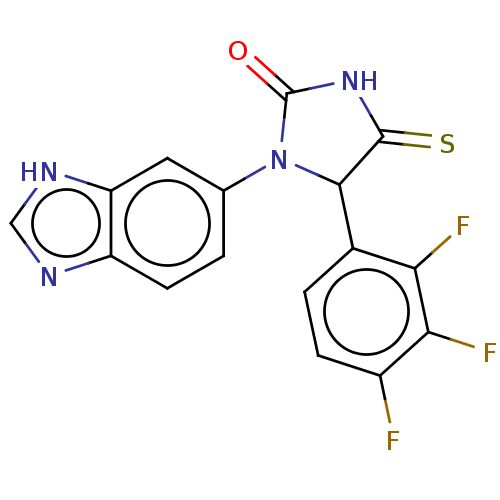
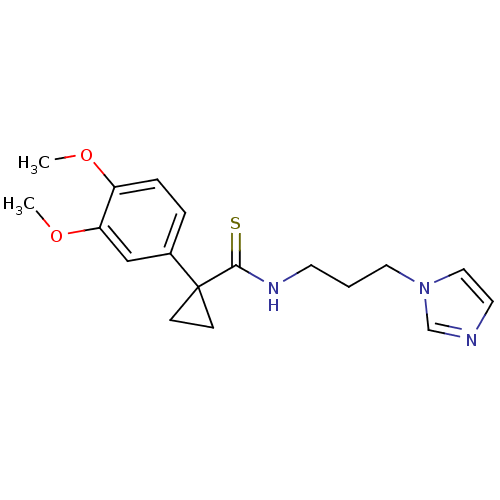
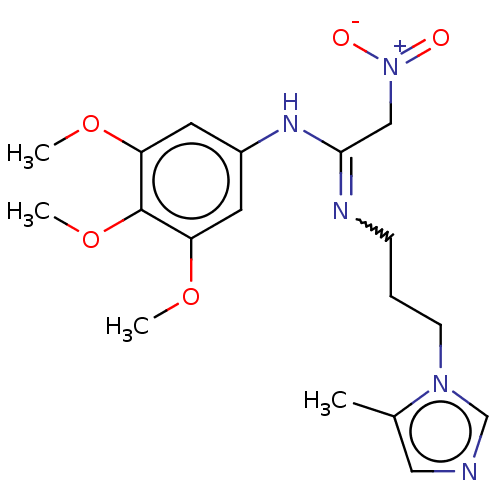
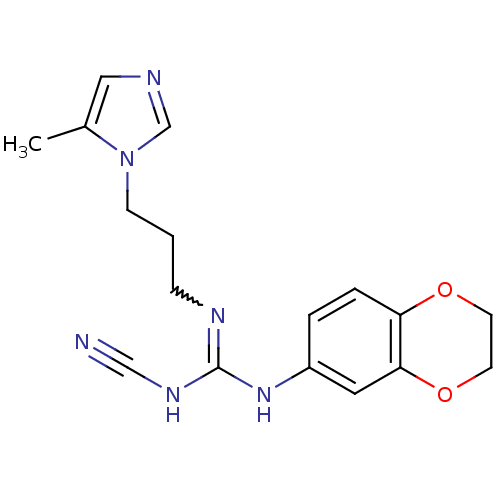
Affinity DataKi: 2.60nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.24nM ΔG°: -49.3kJ/mole IC50: 3.08nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 3.80nM ΔG°: -48.9kJ/mole IC50: 234nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 4.13nM ΔG°: -48.7kJ/mole IC50: 48nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 6.07nM ΔG°: -47.7kJ/mole IC50: 69.7nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 6.30nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.49nM ΔG°: -47.5kJ/mole IC50: 24nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
TargetMeprin A subunit beta(Homo sapiens)
Fraunhofer Institute For Cell Therapy and Immunology Izi
Curated by ChEMBL
Fraunhofer Institute For Cell Therapy and Immunology Izi
Curated by ChEMBL
Affinity DataKi: 8nMAssay Description:Inhibition of recombinant human meprin beta expressed in baculovirus infected Sf9 insect cells using (MCA)-EDEDED-(K-epsilon-dnp) as substrate by flu...More data for this Ligand-Target Pair
Affinity DataKi: 11nM IC50: 580nMpH: 8.0Assay Description:All measurements were performed with a BioAssay Reader HTS-7000Plus for microplates (Perkin Elmer) at 30C. QC activity was evaluated fluorometrically...More data for this Ligand-Target Pair
Affinity DataKi: 13.6nM ΔG°: -45.7kJ/mole IC50: 128nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 17.7nM ΔG°: -45.0kJ/mole IC50: 326nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 21.7nM ΔG°: -44.5kJ/mole IC50: 173nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
TargetMeprin A subunit beta(Homo sapiens)
Fraunhofer Institute For Cell Therapy and Immunology Izi
Curated by ChEMBL
Fraunhofer Institute For Cell Therapy and Immunology Izi
Curated by ChEMBL
Affinity DataKi: 22nMAssay Description:Inhibition of recombinant human meprin beta expressed in baculovirus infected Sf9 insect cells using (MCA)-EDEDED-(K-epsilon-dnp) as substrate by flu...More data for this Ligand-Target Pair
Affinity DataKi: 34nM IC50: 430nMpH: 8.0Assay Description:All measurements were performed with a BioAssay Reader HTS-7000Plus for microplates (Perkin Elmer) at 30C. QC activity was evaluated fluorometrically...More data for this Ligand-Target Pair
Affinity DataKi: 34nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 34.8nM ΔG°: -43.3kJ/mole IC50: 182nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 36nM ΔG°: -43.2kJ/mole IC50: 523nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 39nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 41nM IC50: 230nMpH: 8.0Assay Description:All measurements were performed with a BioAssay Reader HTS-7000Plus for microplates (Perkin Elmer) at 30C. QC activity was evaluated fluorometrically...More data for this Ligand-Target Pair
Affinity DataKi: 41nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 41.3nM ΔG°: -42.9kJ/mole IC50: 741nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 42nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 42.8nM ΔG°: -42.8kJ/mole IC50: 298nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 44nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 45.9nM ΔG°: -42.6kJ/mole IC50: 256nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 48nM ΔG°: -42.5kJ/mole IC50: 34.9nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 51nM IC50: 320nMpH: 8.0Assay Description:All measurements were performed with a BioAssay Reader HTS-7000Plus for microplates (Perkin Elmer) at 30C. QC activity was evaluated fluorometrically...More data for this Ligand-Target Pair
Affinity DataKi: 51nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 51.6nM ΔG°: -42.3kJ/mole IC50: 560nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 55nM IC50: 530nMpH: 8.0Assay Description:All measurements were performed with a BioAssay Reader HTS-7000Plus for microplates (Perkin Elmer) at 30C. QC activity was evaluated fluorometrically...More data for this Ligand-Target Pair
Affinity DataKi: 58.5nM ΔG°: -42.0kJ/mole IC50: 540nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 60nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 60nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 61nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 65nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 65.5nM ΔG°: -41.7kJ/mole IC50: 430nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 67nM IC50: 340nMpH: 8.0Assay Description:All measurements were performed with a BioAssay Reader HTS-7000Plus for microplates (Perkin Elmer) at 30C. QC activity was evaluated fluorometrically...More data for this Ligand-Target Pair
Affinity DataKi: 67nM IC50: 460nMpH: 8.0Assay Description:All measurements were performed with a BioAssay Reader HTS-7000Plus for microplates (Perkin Elmer) at 30C. QC activity was evaluated fluorometrically...More data for this Ligand-Target Pair
Affinity DataKi: 67nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 76nM IC50: 730nMpH: 8.0Assay Description:All measurements were performed with a BioAssay Reader HTS-7000Plus for microplates (Perkin Elmer) at 30C. QC activity was evaluated fluorometrically...More data for this Ligand-Target Pair
Affinity DataKi: 80nM IC50: 340nMpH: 8.0Assay Description:All measurements were performed with a BioAssay Reader HTS-7000Plus for microplates (Perkin Elmer) at 30C. QC activity was evaluated fluorometrically...More data for this Ligand-Target Pair
Affinity DataKi: 82nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 83nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 85.3nM ΔG°: -41.0kJ/mole IC50: 485nMT: 2°CAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 90nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair
Affinity DataKi: 96nM IC50: 620nMpH: 8.0Assay Description:All measurements were performed with a BioAssay Reader HTS-7000Plus for microplates (Perkin Elmer) at 30C. QC activity was evaluated fluorometrically...More data for this Ligand-Target Pair
Affinity DataKi: 130nMAssay Description:Inhibition of human glutaminyl cyclase expressed in Pichia pastoris by pGAP coupled enzyme assayMore data for this Ligand-Target Pair

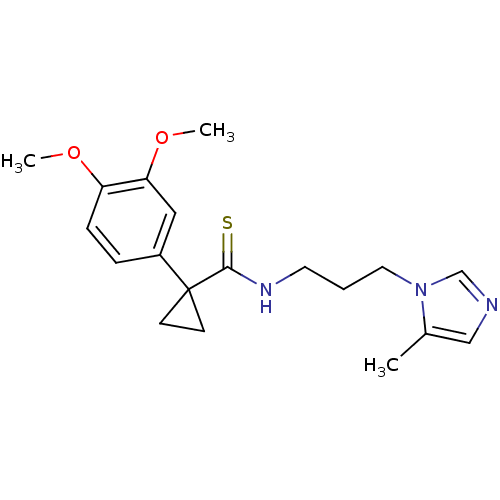
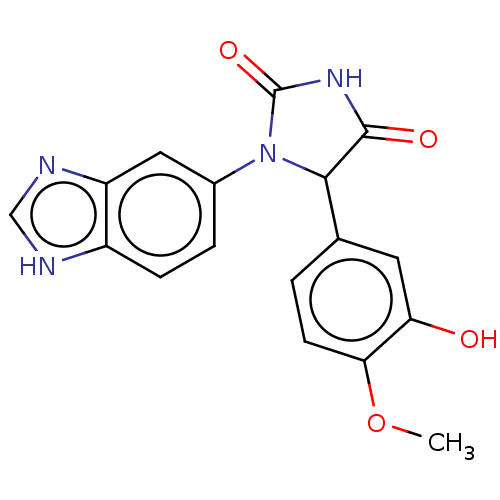
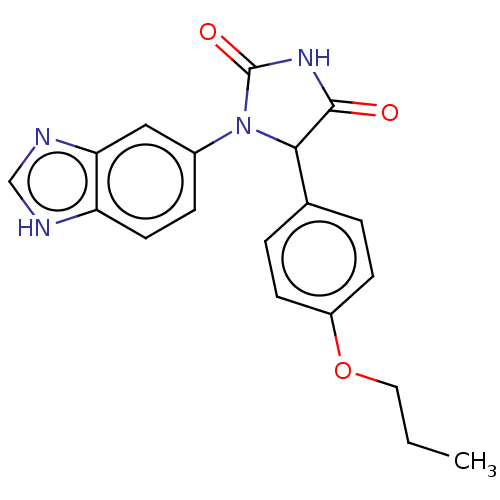
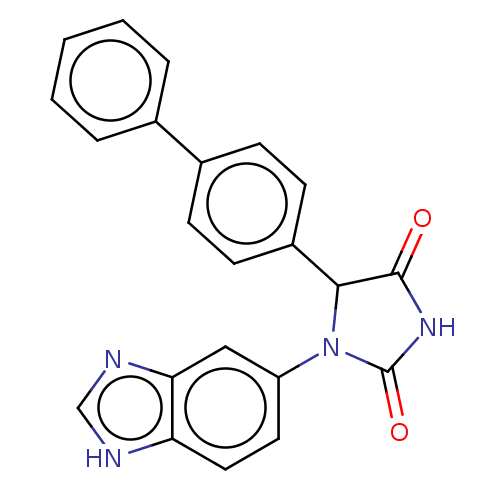
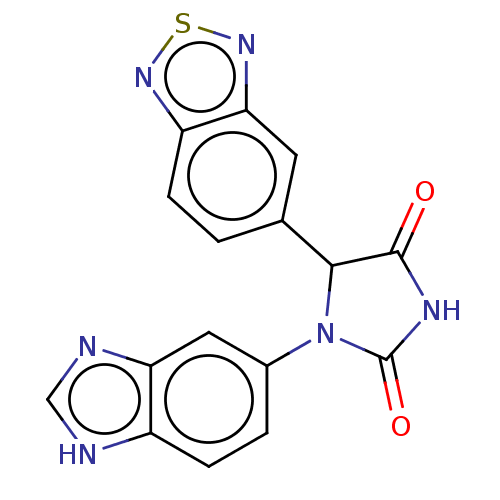
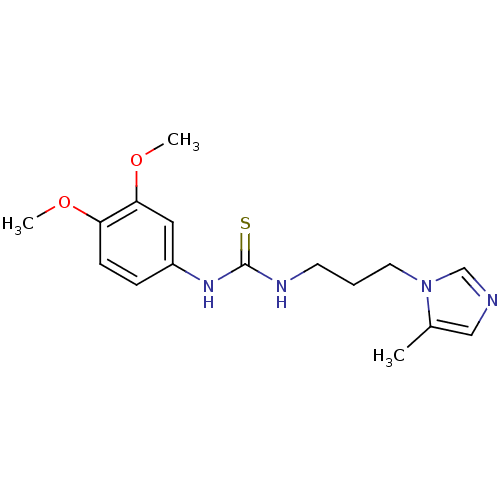
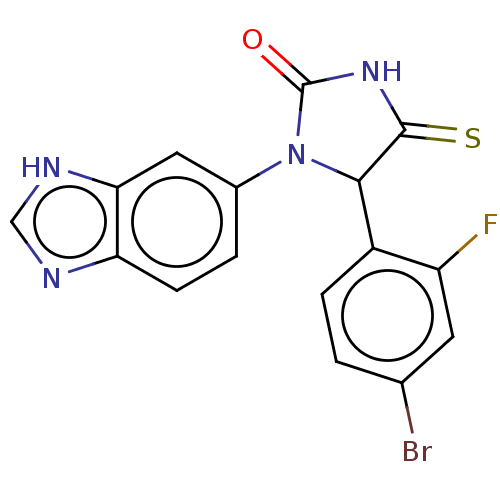
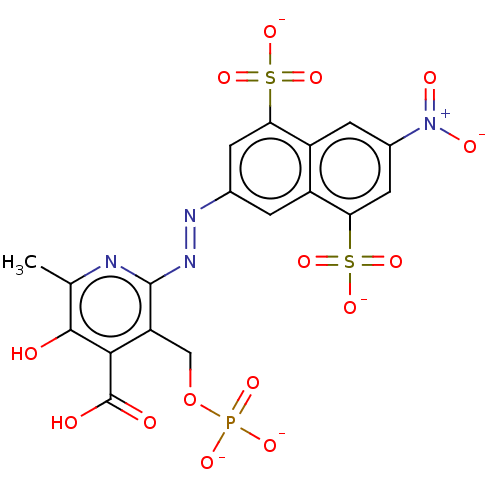
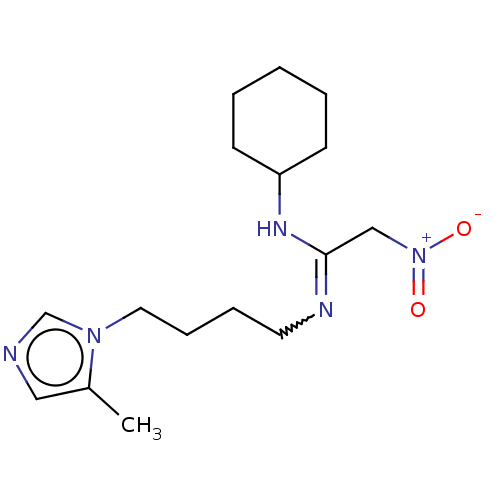
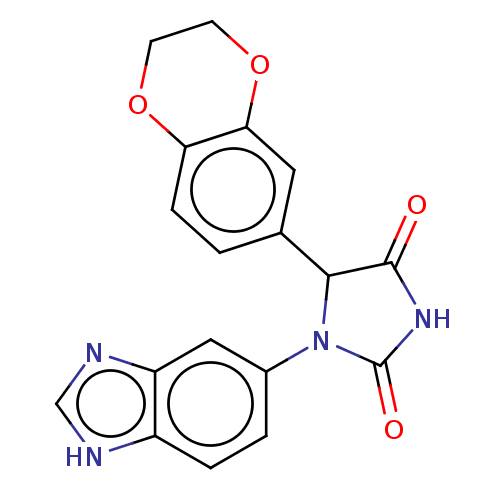
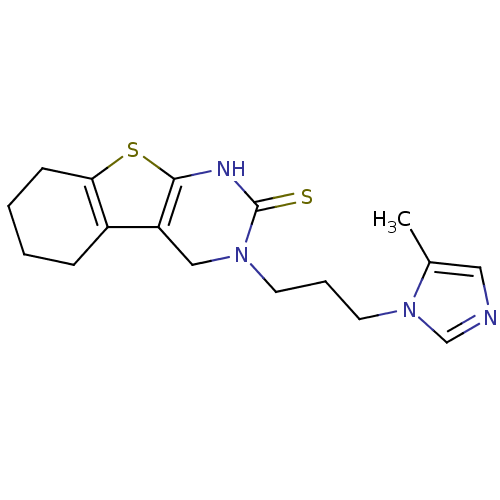
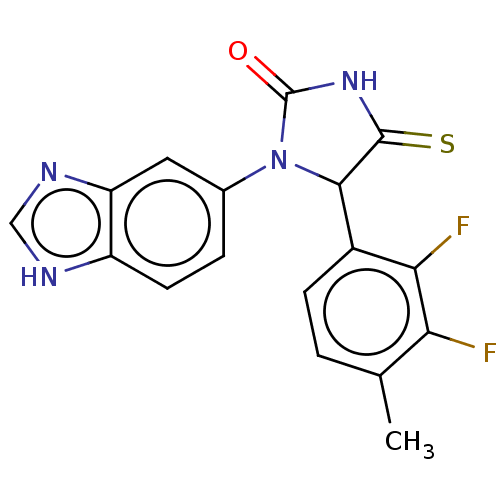
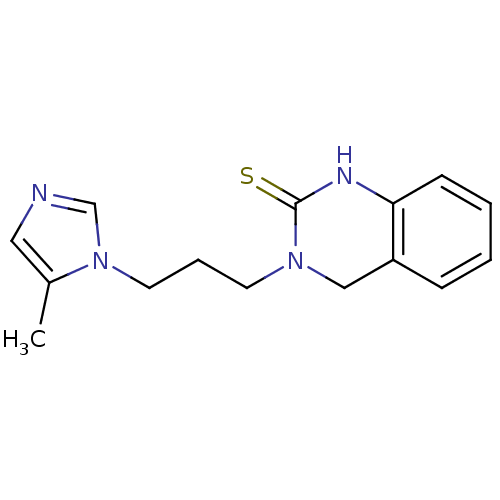

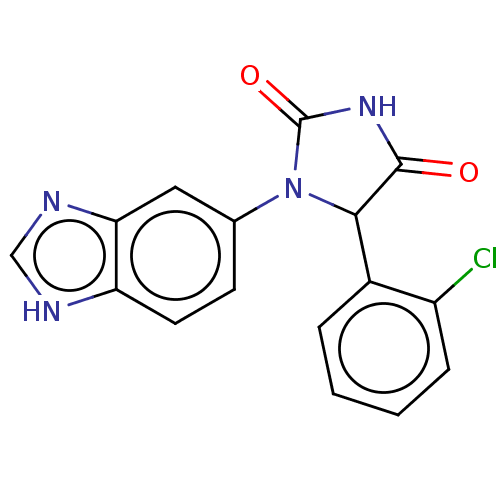
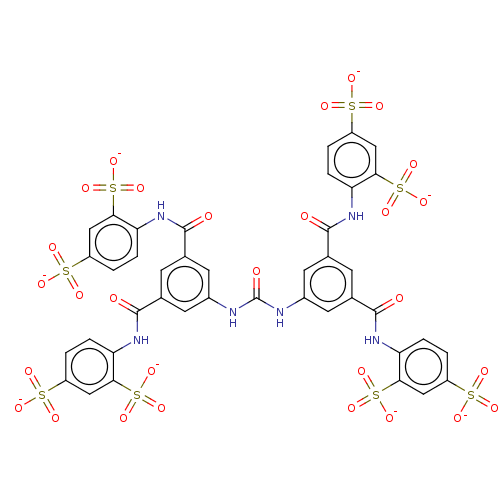
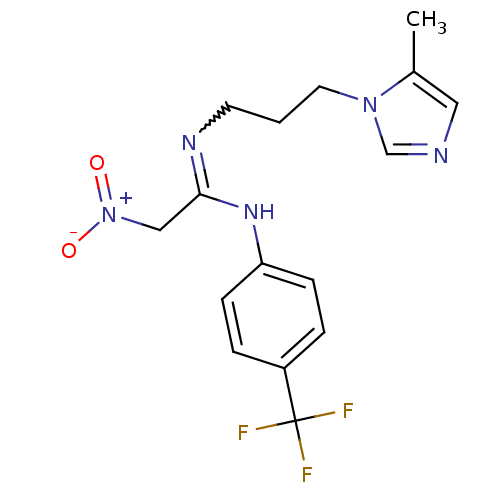
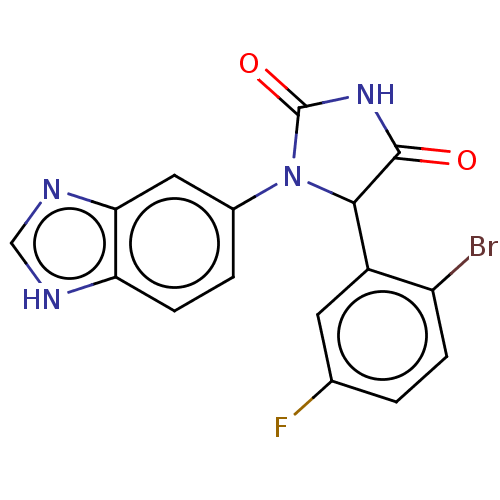
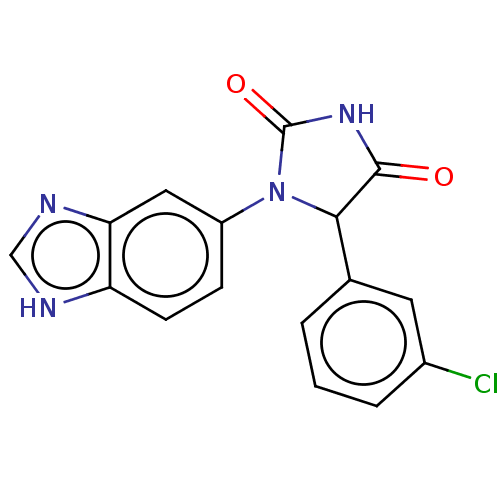
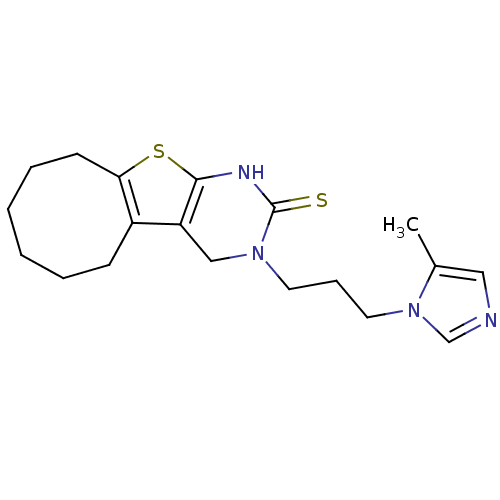
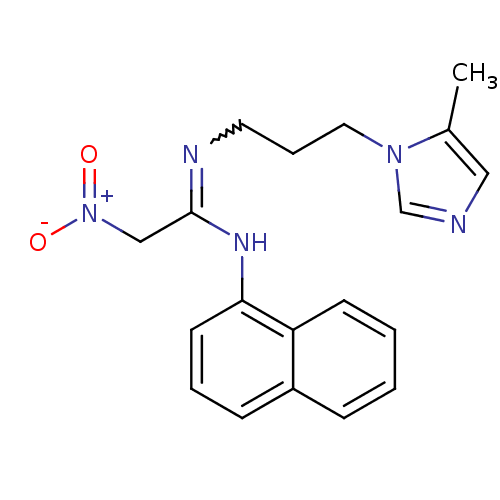

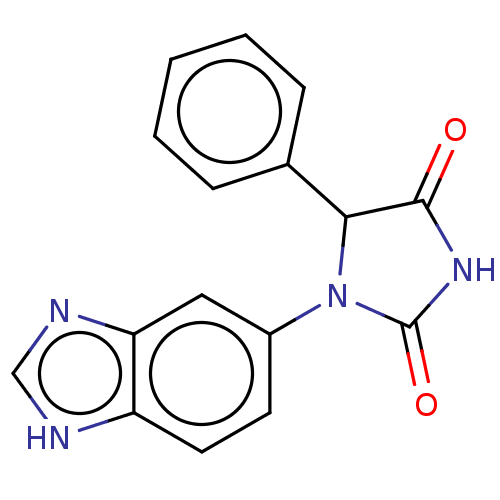
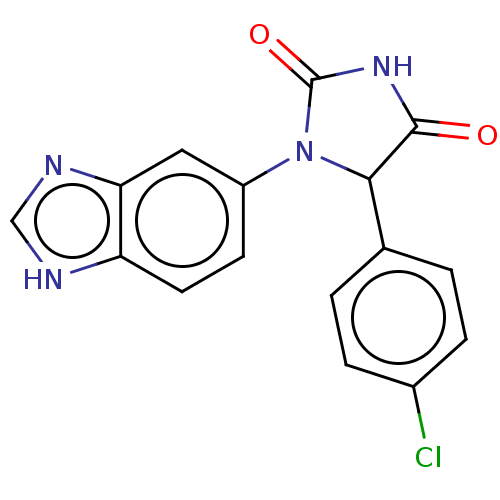
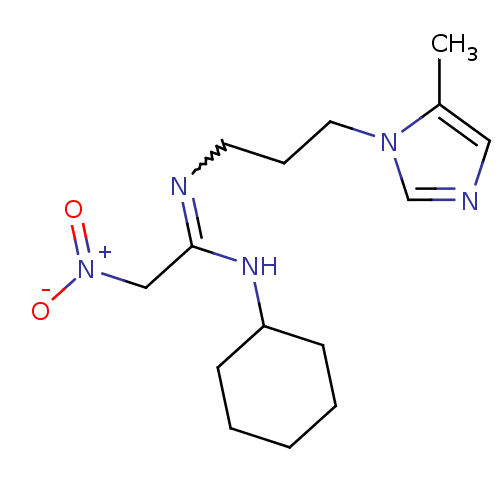
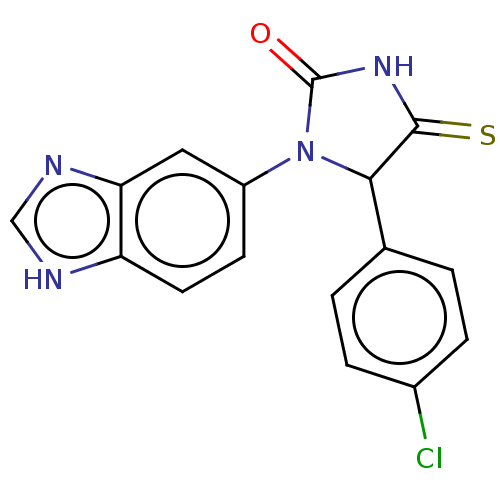
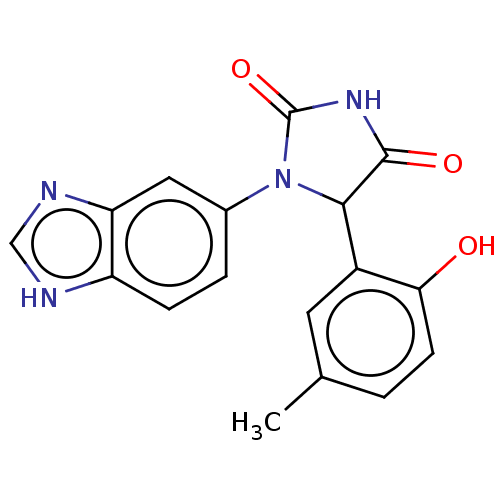
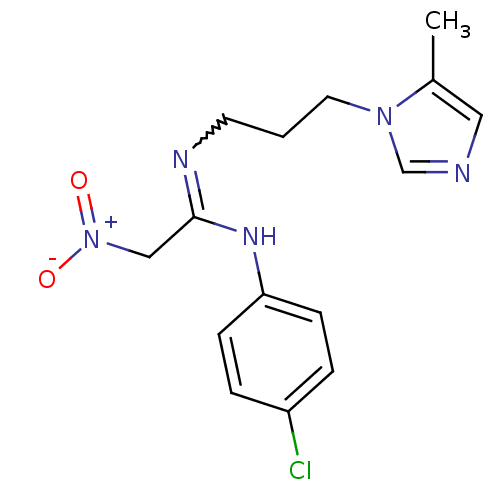
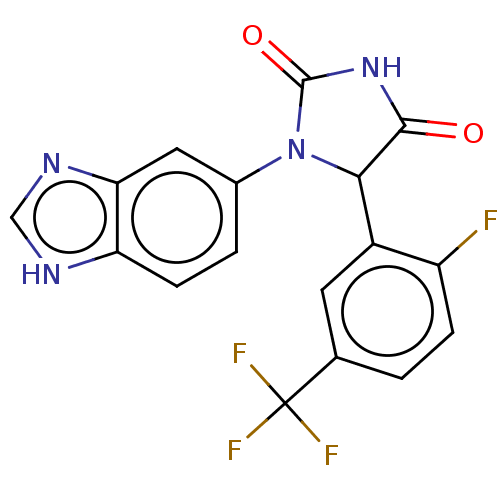
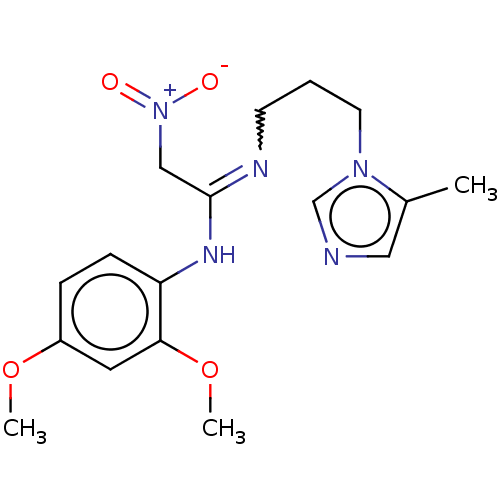
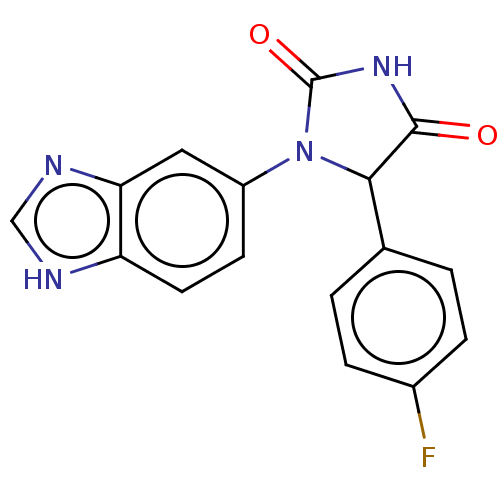
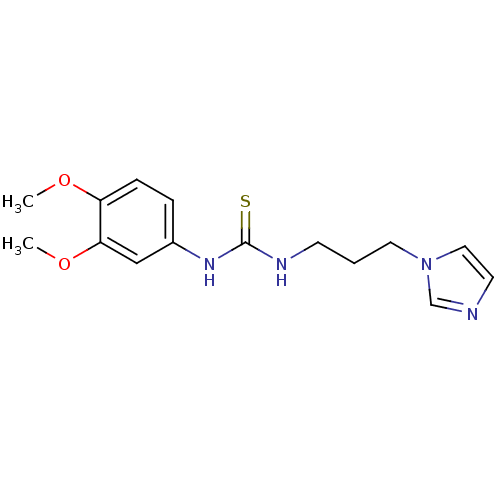
 3D Structure (crystal)
3D Structure (crystal)