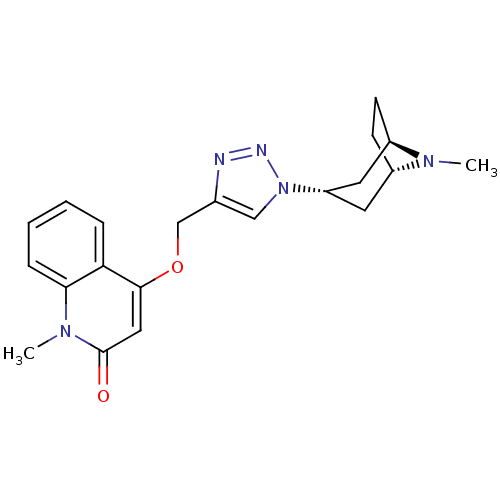
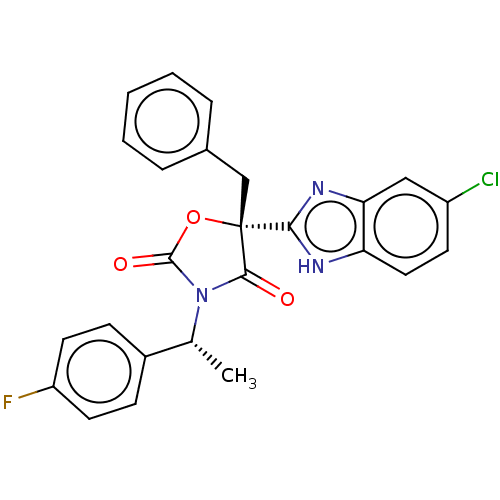
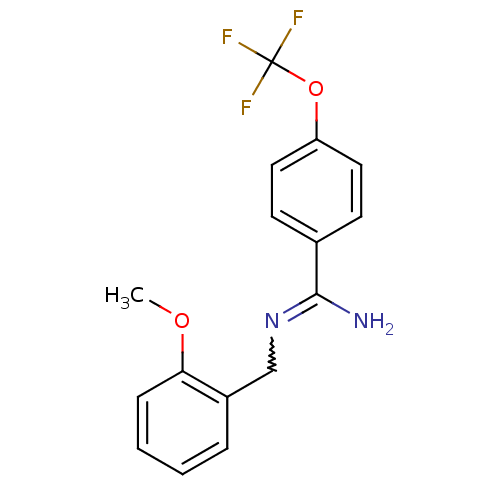
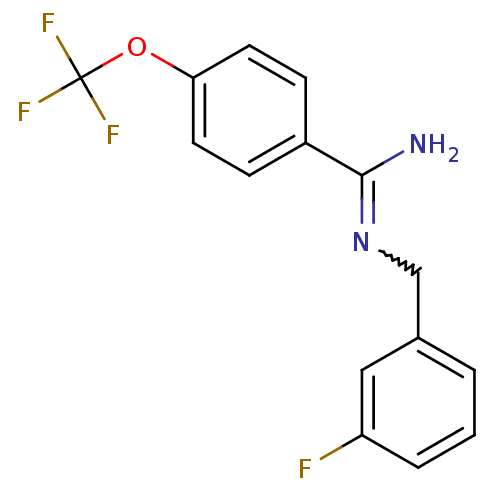
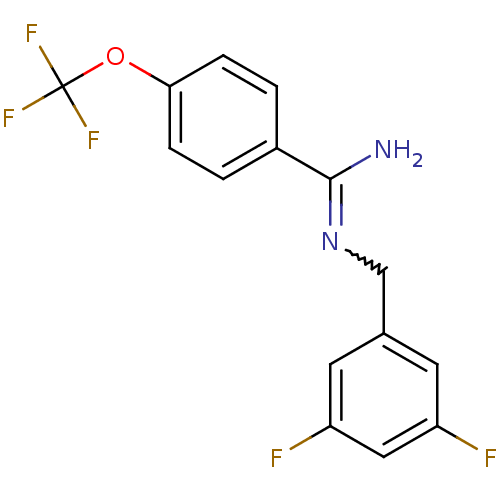
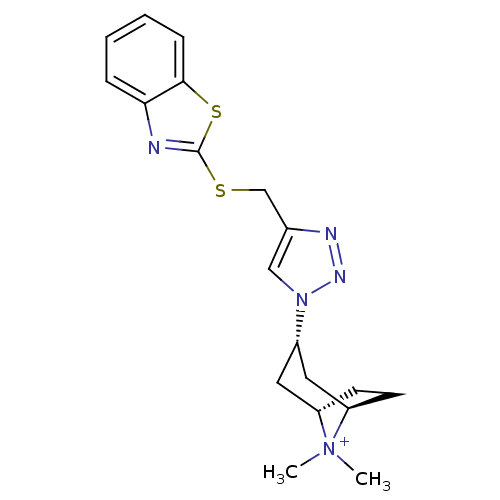
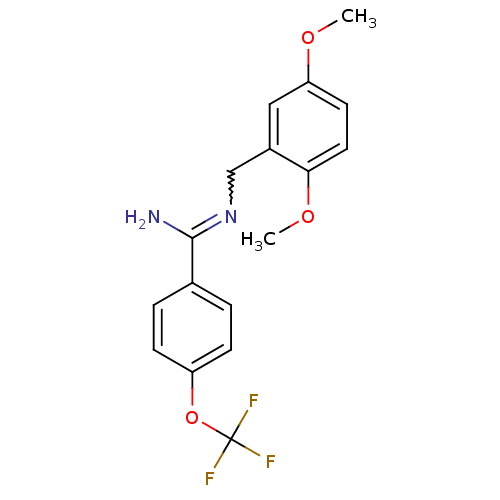
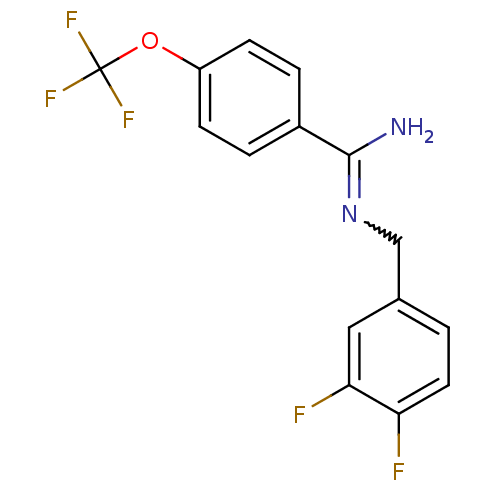
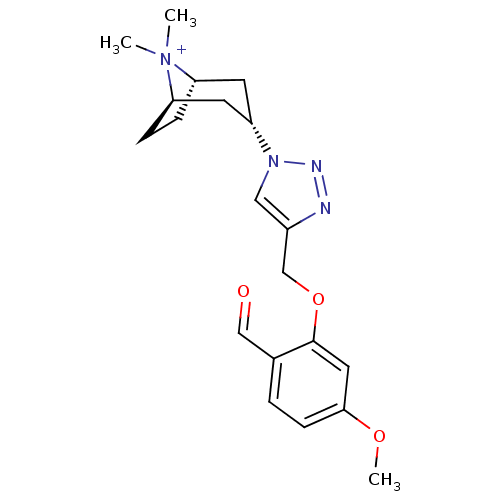
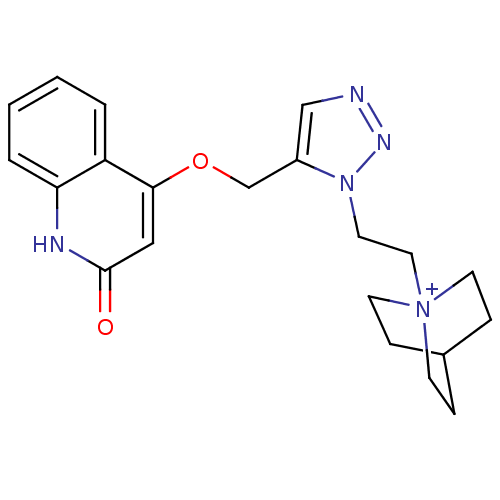
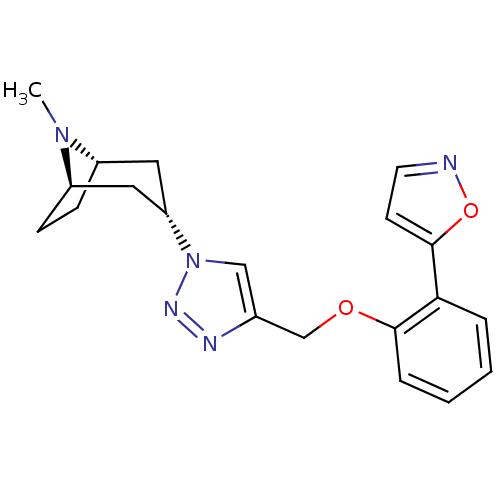
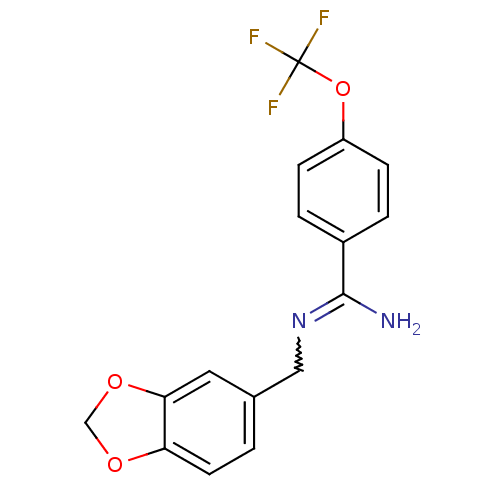
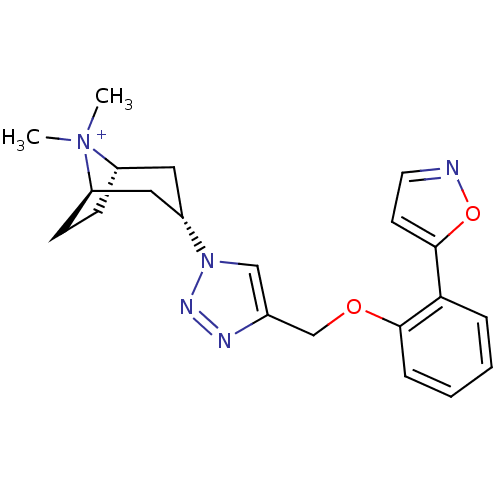
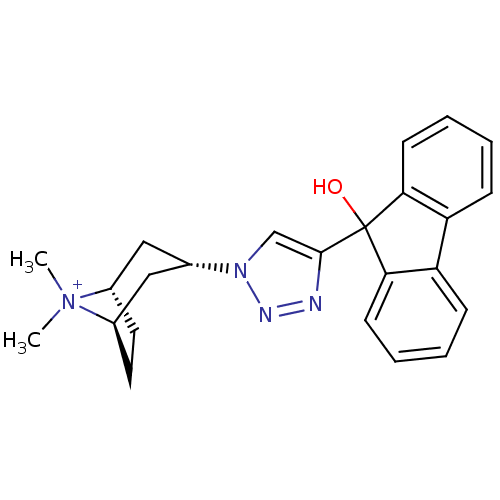
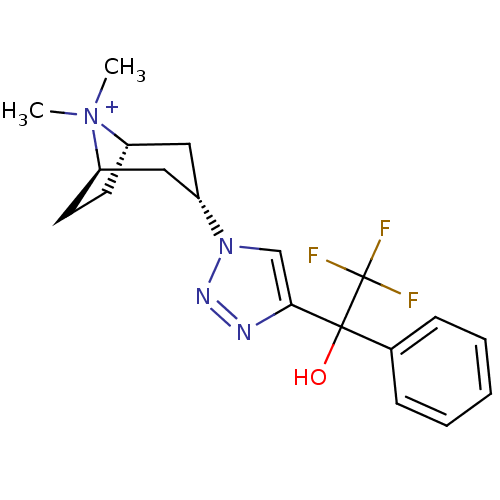
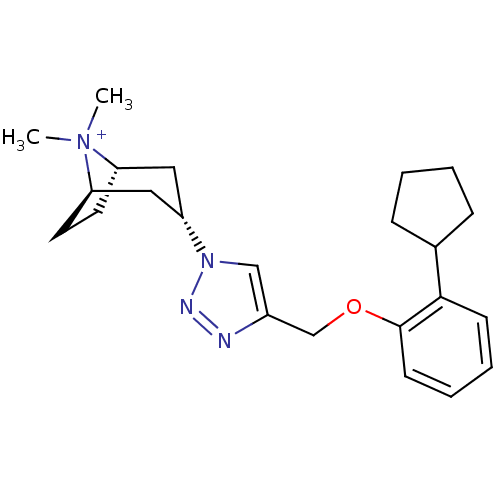
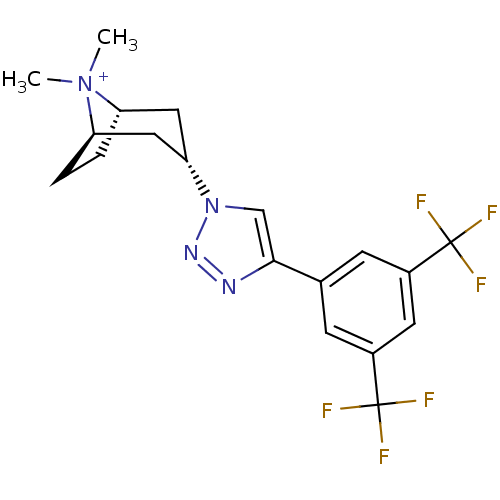
Affinity DataKi: 1.30nM ΔG°: -49.9kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 2nM ΔG°: -48.8kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Binding affinity to mineralocorticoid receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
The University Of Sydney
Curated by ChEMBL
The University Of Sydney
Curated by ChEMBL
Affinity DataKi: 2.10nMAssay Description:Displacement of [3H]ifenprodil from Wistar rat GluN2B receptor after 2 hrs by liquid scintillation spectrometryMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
The University Of Sydney
Curated by ChEMBL
The University Of Sydney
Curated by ChEMBL
Affinity DataKi: 2.90nMAssay Description:Displacement of [3H]ifenprodil from Wistar rat GluN2B receptor after 2 hrs by liquid scintillation spectrometryMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
The University Of Sydney
Curated by ChEMBL
The University Of Sydney
Curated by ChEMBL
Affinity DataKi: 3.60nMAssay Description:Displacement of [3H]ifenprodil from Wistar rat GluN2B receptor after 2 hrs by liquid scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 4nM ΔG°: -47.1kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
The University Of Sydney
Curated by ChEMBL
The University Of Sydney
Curated by ChEMBL
Affinity DataKi: 4.30nMAssay Description:Displacement of [3H]ifenprodil from Wistar rat GluN2B receptor after 2 hrs by liquid scintillation spectrometryMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
The University Of Sydney
Curated by ChEMBL
The University Of Sydney
Curated by ChEMBL
Affinity DataKi: 5.90nMAssay Description:Displacement of [3H]epibatidine from rat alpha7 nAChR transfected in HEK293 cells by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 8nM ΔG°: -45.4kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
The University Of Sydney
Curated by ChEMBL
The University Of Sydney
Curated by ChEMBL
Affinity DataKi: 8.10nMAssay Description:Displacement of [3H]ifenprodil from Wistar rat GluN2B receptor after 2 hrs by liquid scintillation spectrometryMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
The University Of Sydney
Curated by ChEMBL
The University Of Sydney
Curated by ChEMBL
Affinity DataKi: 8.30nMAssay Description:Displacement of [3H]ifenprodil from Wistar rat GluN2B receptor after 2 hrs by liquid scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 13nM ΔG°: -44.3kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 13nM ΔG°: -44.3kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 16nM ΔG°: -43.8kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
The University Of Sydney
Curated by ChEMBL
The University Of Sydney
Curated by ChEMBL
Affinity DataKi: 16nMAssay Description:Displacement of [3H]ifenprodil from Wistar rat GluN2B receptor after 2 hrs by liquid scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 18nM ΔG°: -43.5kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 20nM ΔG°: -43.2kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
The University Of Sydney
Curated by ChEMBL
The University Of Sydney
Curated by ChEMBL
Affinity DataKi: 23nMAssay Description:Displacement of [3H]ifenprodil from Wistar rat GluN2B receptor after 2 hrs by liquid scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 24nM ΔG°: -42.8kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 25nM ΔG°: -42.7kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
The University Of Sydney
Curated by ChEMBL
The University Of Sydney
Curated by ChEMBL
Affinity DataKi: 26nMAssay Description:Displacement of [3H]ifenprodil from Wistar rat GluN2B receptor after 2 hrs by liquid scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 26nM ΔG°: -42.6kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 30nM ΔG°: -42.2kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 30nM ΔG°: -42.2kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
The University Of Sydney
Curated by ChEMBL
The University Of Sydney
Curated by ChEMBL
Affinity DataKi: 34nMAssay Description:Displacement of [3H]epibatidine from rat alpha7 nAChR transfected in HEK293 cells by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 40nM ΔG°: -41.5kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 40nM ΔG°: -41.5kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 40nM ΔG°: -41.5kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 50nM ΔG°: -41.0kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 50nM ΔG°: -41.0kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 50nM ΔG°: -41.0kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 60nM ΔG°: -40.5kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 60nM ΔG°: -40.5kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 70nM ΔG°: -40.2kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 80nM ΔG°: -39.8kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
The University Of Sydney
Curated by ChEMBL
The University Of Sydney
Curated by ChEMBL
Affinity DataKi: 85nMAssay Description:Displacement of [3H]epibatidine from rat alpha7 nAChR transfected in HEK293 cells by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 96nM ΔG°: -39.4kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 100nM ΔG°: -39.3kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 109nM ΔG°: -39.1kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 110nM ΔG°: -39.1kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 110nM ΔG°: -39.1kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 110nM ΔG°: -39.1kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 120nM ΔG°: -38.8kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 120nM ΔG°: -38.8kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 120nM ΔG°: -38.8kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 130nM ΔG°: -38.6kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-7(Rattus norvegicus (Rat))
The University Of Sydney
Curated by ChEMBL
The University Of Sydney
Curated by ChEMBL
Affinity DataKi: 140nMAssay Description:Displacement of [3H]epibatidine from rat alpha7 nAChR transfected in HEK293 cells by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 140nM ΔG°: -38.5kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair
Affinity DataKi: 140nM ΔG°: -38.5kJ/molepH: 7.4 T: 2°CAssay Description:Assay Description 2 Assays used to generate Ki or EC50 values. 1) SPA Assay - Quick screen binding assays were performed using 100 ?l of 0.2 mg/ml an...More data for this Ligand-Target Pair