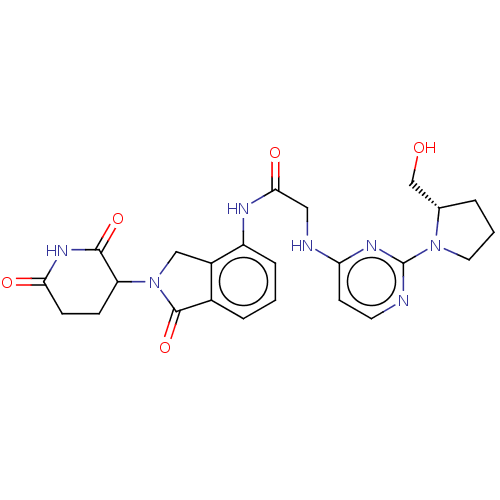
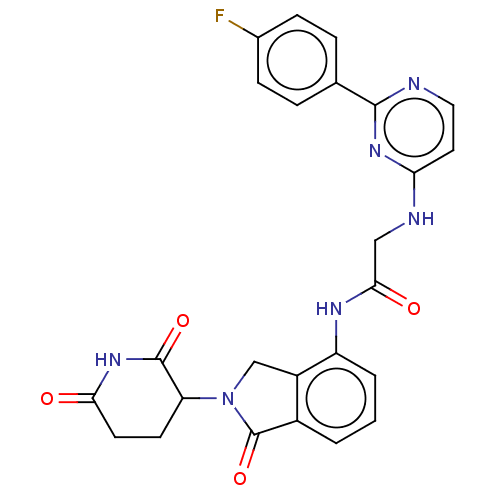
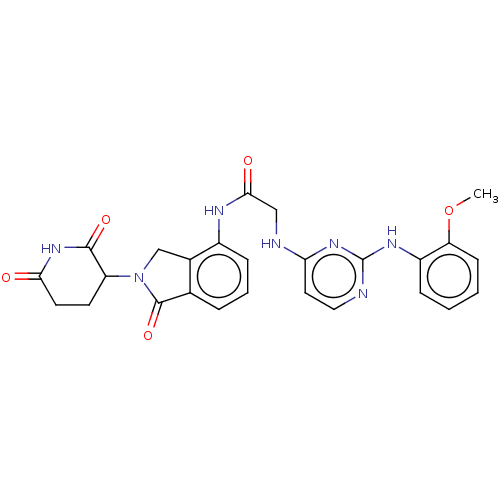
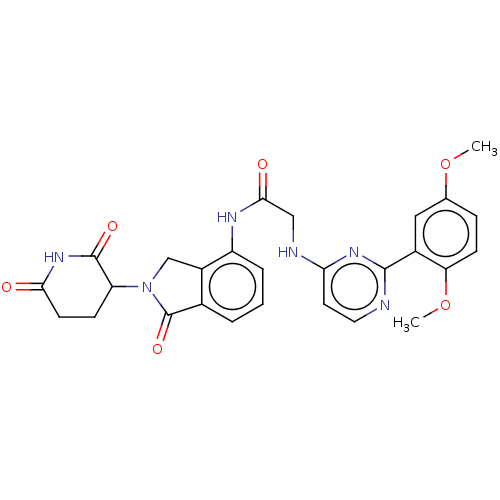
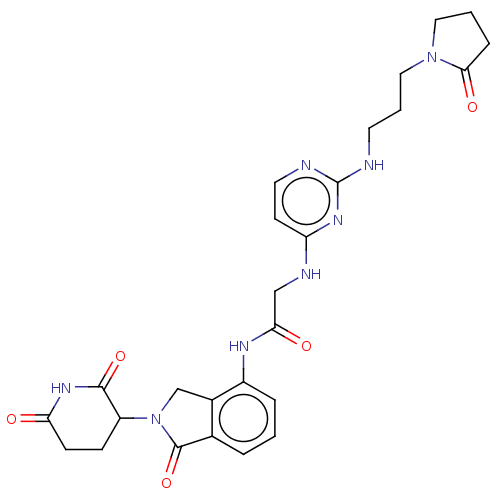
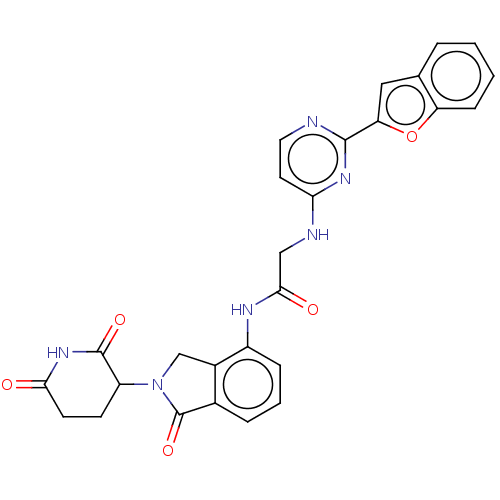
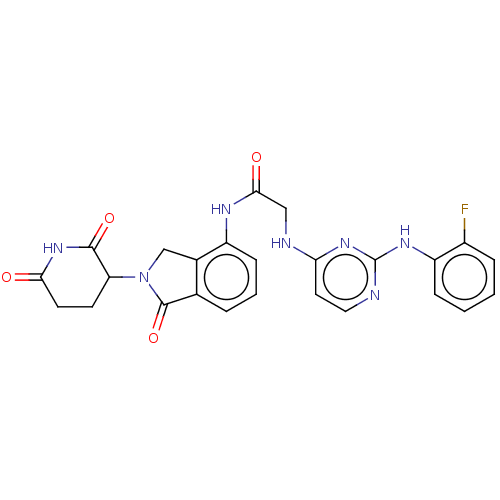
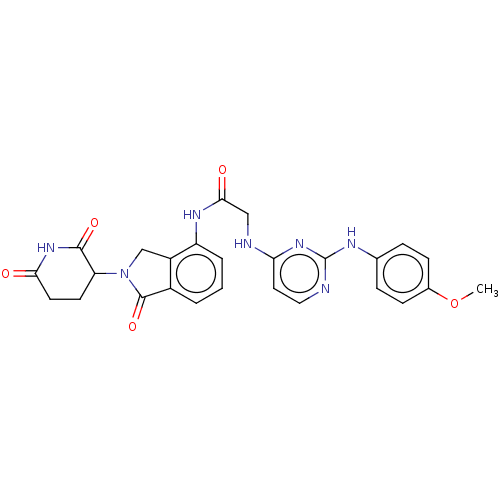
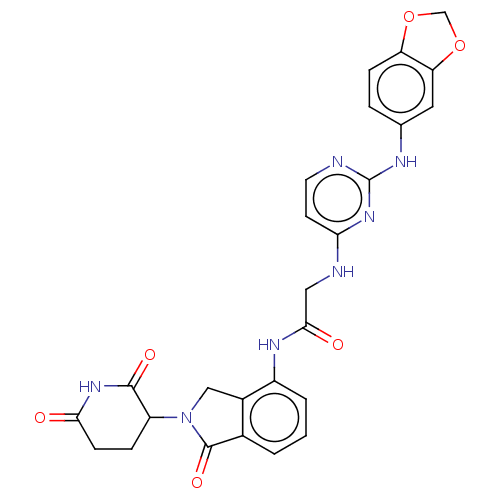
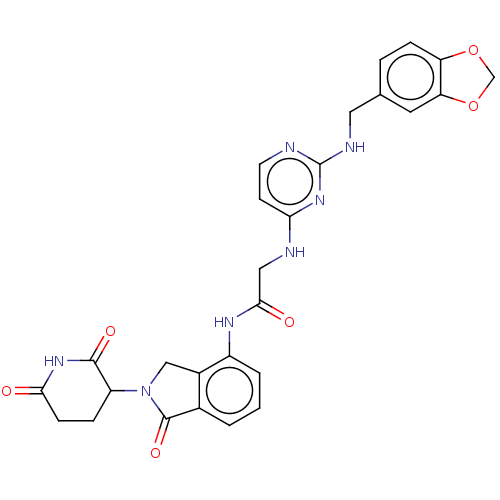
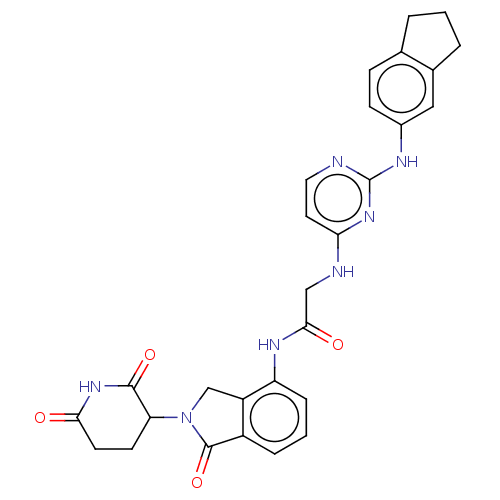
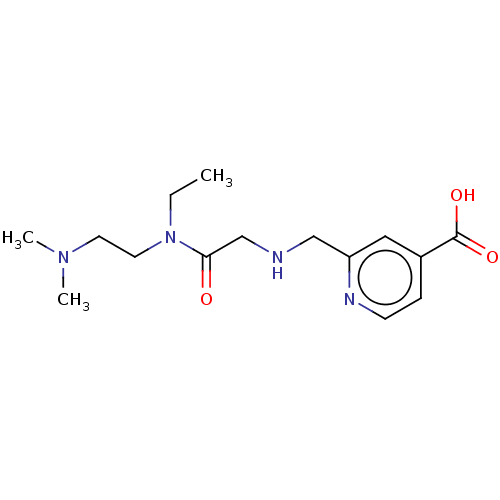
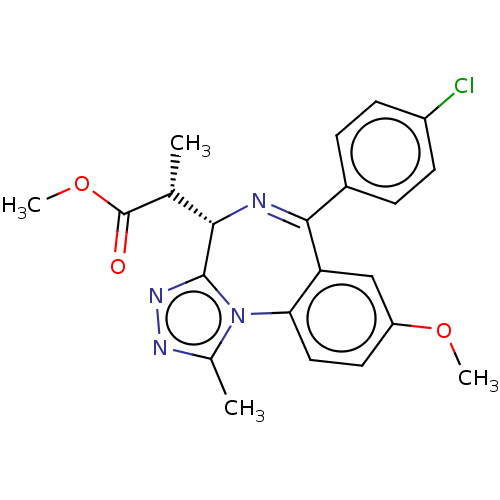
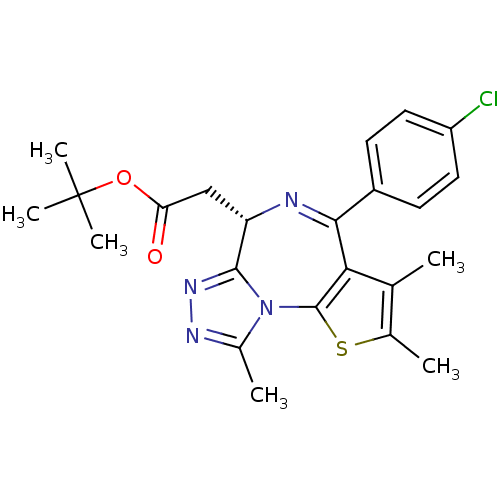
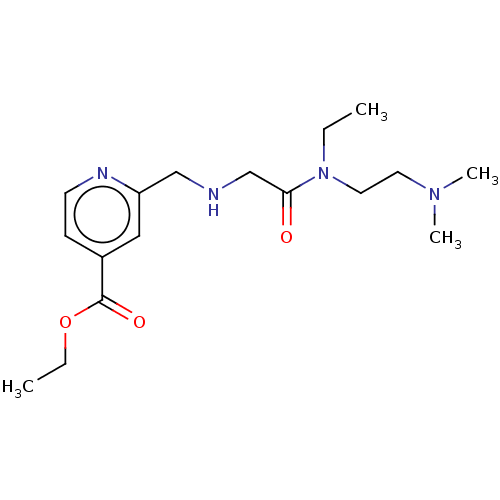
Affinity DataKi: 2.20nMAssay Description:Competitive inhibition of N-terminal His6-tagged KDM4C (1 to 352 residues) (unknown origin) expressed in Escherichia coli Rosetta 2(DE3)pLysS using A...More data for this Ligand-Target Pair
TargetPeptidyl-prolyl cis-trans isomerase A(Homo sapiens (Human))
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Affinity DataKi: 11nMAssay Description:Inhibition of CypA PPIase activity (unknown origin) using Glt-(Ala)n-Pro-Phe-4-nitroanilides as substrateMore data for this Ligand-Target Pair
TargetPeptidyl-prolyl cis-trans isomerase A(Homo sapiens (Human))
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Affinity DataKi: 12nMAssay Description:Inhibition of CypA PPIase activity (unknown origin) using Glt-(Ala)n-Pro-Phe-4-nitroanilides as substrateMore data for this Ligand-Target Pair
TargetPeptidyl-prolyl cis-trans isomerase A(Homo sapiens (Human))
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Affinity DataKi: 12nMAssay Description:Inhibition of CypA PPIase activity (unknown origin) using Glt-(Ala)n-Pro-Phe-4-nitroanilides as substrateMore data for this Ligand-Target Pair
TargetPeptidyl-prolyl cis-trans isomerase A(Homo sapiens (Human))
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Affinity DataKi: 12nMAssay Description:Inhibition of CypA PPIase activity (unknown origin) using Glt-(Ala)n-Pro-Phe-4-nitroanilides as substrateMore data for this Ligand-Target Pair
TargetPeptidyl-prolyl cis-trans isomerase A(Homo sapiens (Human))
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Affinity DataKi: 23nMAssay Description:Inhibition of CypA PPIase activity (unknown origin) using Glt-(Ala)n-Pro-Phe-4-nitroanilides as substrateMore data for this Ligand-Target Pair
TargetPeptidyl-prolyl cis-trans isomerase B(Homo sapiens (Human))
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Affinity DataKi: 35nMAssay Description:Inhibition of CypB PPIase activity (unknown origin)More data for this Ligand-Target Pair
TargetPeptidyl-prolyl cis-trans isomerase A(Homo sapiens (Human))
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Affinity DataKi: 37nMAssay Description:Inhibition of CypA PPIase activity (unknown origin) using Glt-(Ala)n-Pro-Phe-4-nitroanilides as substrateMore data for this Ligand-Target Pair
Affinity DataKi: 43nMAssay Description:Competitive inhibition of N-terminal His6-tagged KDM4C (1 to 352 residues) (unknown origin) expressed in Escherichia coli Rosetta 2(DE3)pLysS using A...More data for this Ligand-Target Pair
TargetPeptidyl-prolyl cis-trans isomerase A(Homo sapiens (Human))
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Affinity DataKi: 44nMAssay Description:Inhibition of CypA PPIase activity (unknown origin) using Glt-(Ala)n-Pro-Phe-4-nitroanilides as substrateMore data for this Ligand-Target Pair
TargetPeptidyl-prolyl cis-trans isomerase A(Homo sapiens (Human))
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Affinity DataKi: 58nMAssay Description:Inhibition of CypA PPIase activity (unknown origin) using Glt-(Ala)n-Pro-Phe-4-nitroanilides as substrateMore data for this Ligand-Target Pair
TargetPeptidyl-prolyl cis-trans isomerase A(Homo sapiens (Human))
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Affinity DataKi: 65nMAssay Description:Inhibition of CypA PPIase activity (unknown origin) using Glt-(Ala)n-Pro-Phe-4-nitroanilides as substrateMore data for this Ligand-Target Pair
TargetPeptidyl-prolyl cis-trans isomerase C(Homo sapiens (Human))
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Max Planck Research Unit For Enzymology Of Protein Folding
Curated by ChEMBL
Affinity DataKi: 118nMAssay Description:Inhibition of CypC PPIase activity (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 683nMAssay Description:Competitive inhibition of N-terminal His6-tagged KDM4C (1 to 352 residues) (unknown origin) expressed in Escherichia coli Rosetta 2(DE3)pLysS using A...More data for this Ligand-Target Pair
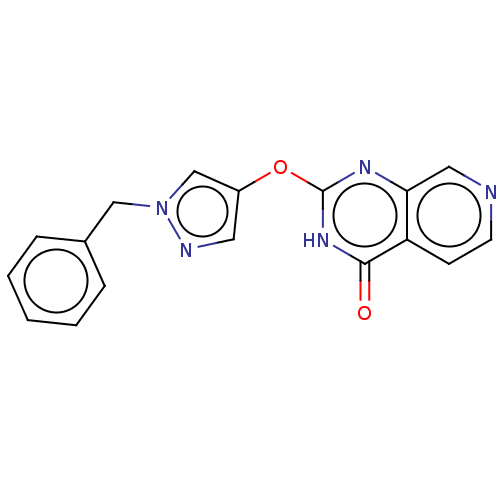
Affinity DataIC50: 4nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
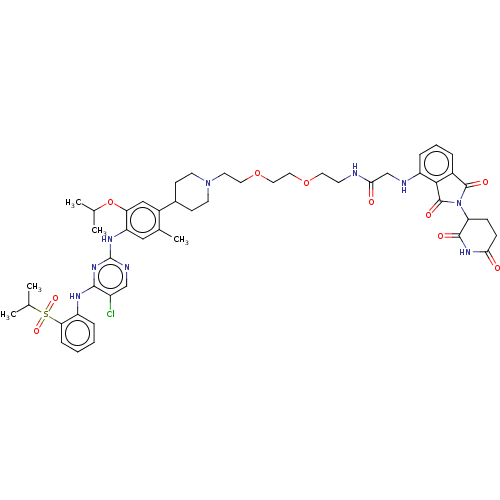
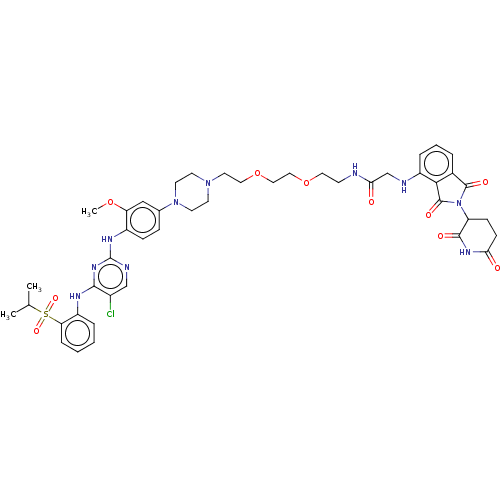
Affinity DataIC50: 10nMAssay Description:Induction of ALK degradation in human NCI-H3122 cells after 16 hrs by immunoblot methodMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Induction of ALK degradation in human NCI-H3122 cells after 16 hrs by immunoblot methodMore data for this Ligand-Target Pair
Affinity DataIC50: 13nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 24nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 26nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
TargetBromodomain-containing protein 4(Homo sapiens (Human))
Dana Farber Cancer Institute
Curated by ChEMBL
Dana Farber Cancer Institute
Curated by ChEMBL
Affinity DataIC50: 39nMAssay Description:Displacement of FITC-conjugated JQ1 from wildtype BRD4 BD1 (unknown origin) expressed in Escherichia coli BL21-DE3 rosetta cells incubated for 15 min...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:Induction of ALK degradation in human KARPAS299 cells after 16 hrs by immunoblot methodMore data for this Ligand-Target Pair
TargetBromodomain-containing protein 4(Homo sapiens (Human))
Dana Farber Cancer Institute
Curated by ChEMBL
Dana Farber Cancer Institute
Curated by ChEMBL
Affinity DataIC50: 49nMAssay Description:Displacement of FITC-conjugated JQ1 from wildtype BRD4 BD1 (unknown origin) expressed in Escherichia coli BL21-DE3 rosetta cells incubated for 15 min...More data for this Ligand-Target Pair
Affinity DataIC50: 49nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair

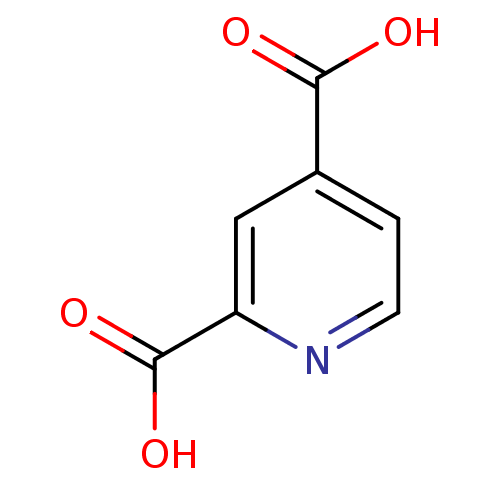
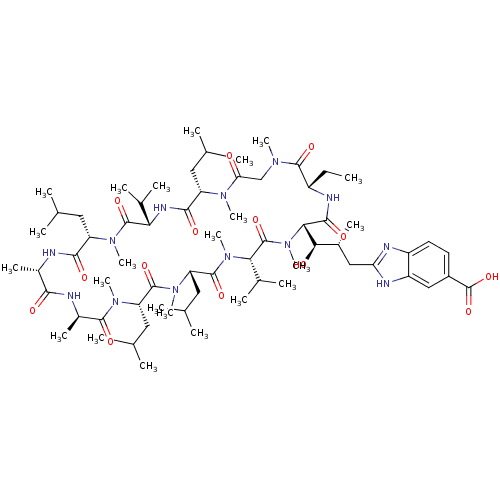
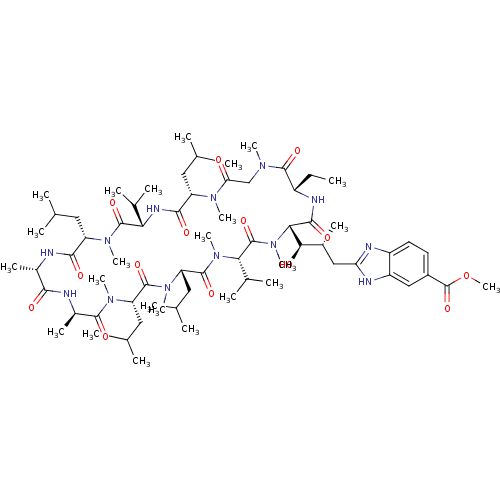
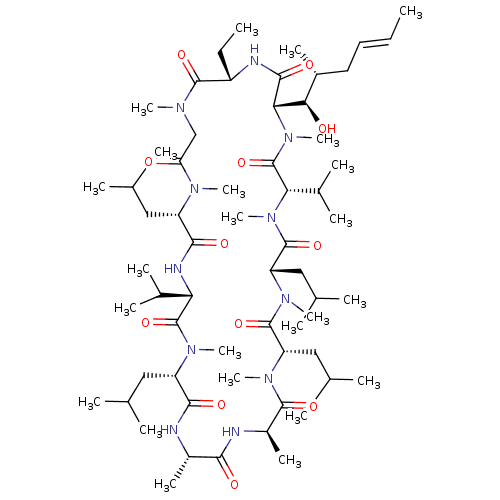
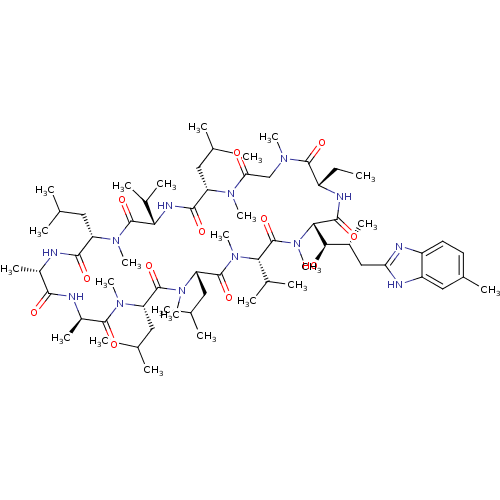
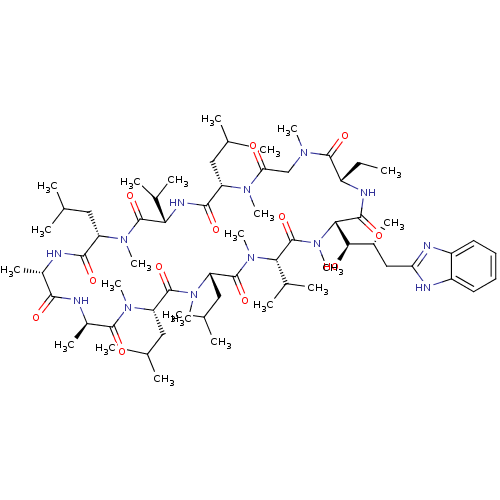
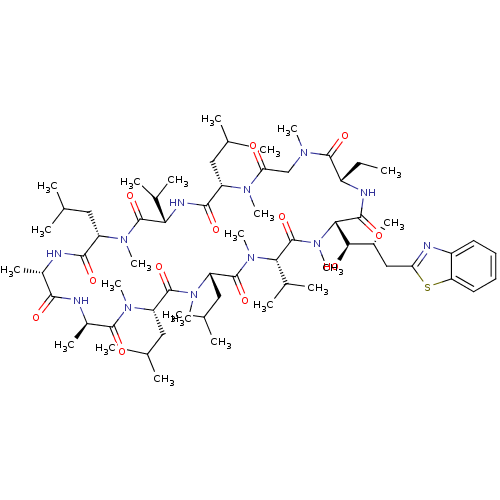
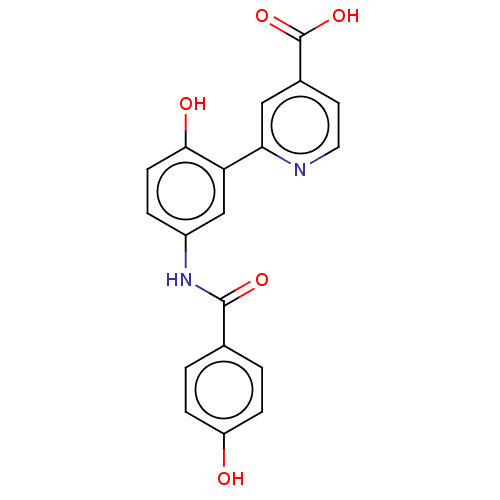


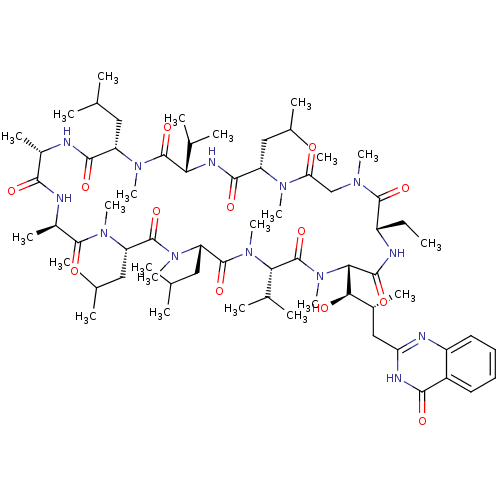

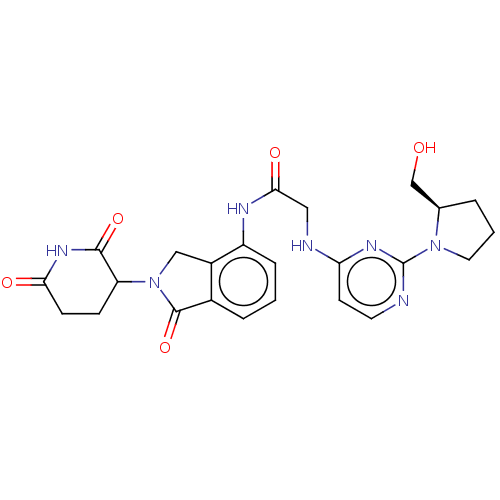
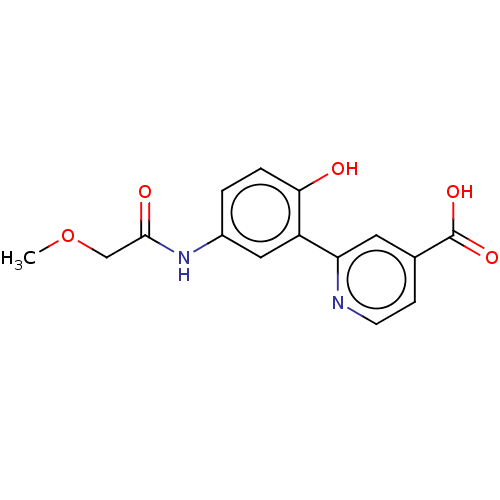

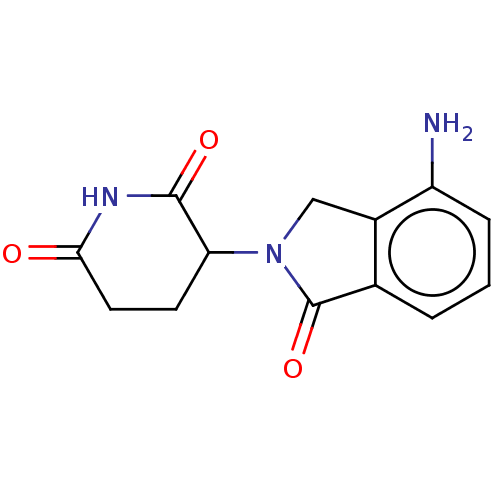
 3D Structure (crystal)
3D Structure (crystal)