Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 11 hits for monomerid = 152729,152730,152731,50405937,50438391,50438393,152729,152730,152731,50405937,50438391,50438393
Found 11 hits for monomerid = 152729,152730,152731,50405937,50438391,50438393,152729,152730,152731,50405937,50438391,50438393
TargetAcetylpolyamine amidohydrolase(Mycoplana ramosa (Gram-negative bacterium))
University of Pennsylvania
University of Pennsylvania
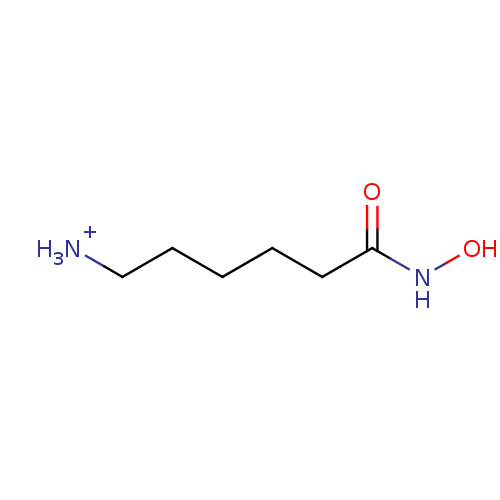
Affinity DataIC50: 68nMpH: 8.2Assay Description:Activity assays were conducted at 25 °C and contained 250 nM APAH (∼50% Zn2+ occupancy), 150 μM substrate, and 0−250 μM inhibit...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Inhibition of zebrafish HDAC10 (2 to 675 residues) expressed in Escherichia coli BL21(DE3) cells using N-acetylputrescine as substrate preincubated f...More data for this Ligand-Target Pair
TargetAcetylpolyamine amidohydrolase(Mycoplana ramosa (Gram-negative bacterium))
University of Pennsylvania
University of Pennsylvania
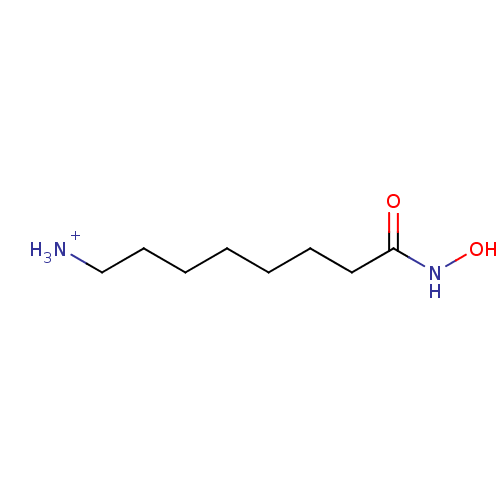
Affinity DataIC50: 130nMpH: 8.2Assay Description:Activity assays were conducted at 25 °C and contained 250 nM APAH (∼50% Zn2+ occupancy), 150 μM substrate, and 0−250 μM inhibit...More data for this Ligand-Target Pair
TargetAcetylpolyamine amidohydrolase(Mycoplana ramosa (Gram-negative bacterium))
University of Pennsylvania
University of Pennsylvania
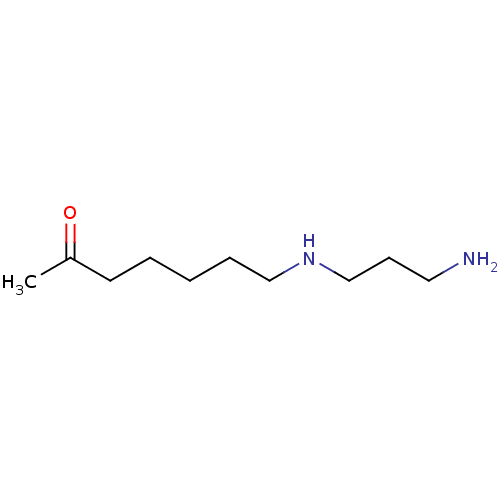
Affinity DataIC50: 150nMpH: 8.2Assay Description:Activity assays were conducted at 25 °C and contained 250 nM APAH (∼50% Zn2+ occupancy), 150 μM substrate, and 0−250 μM inhibit...More data for this Ligand-Target Pair
TargetPeroxisomal N(1)-acetyl-spermine/spermidine oxidase(Human)
University of The Pacific
Curated by ChEMBL
University of The Pacific
Curated by ChEMBL
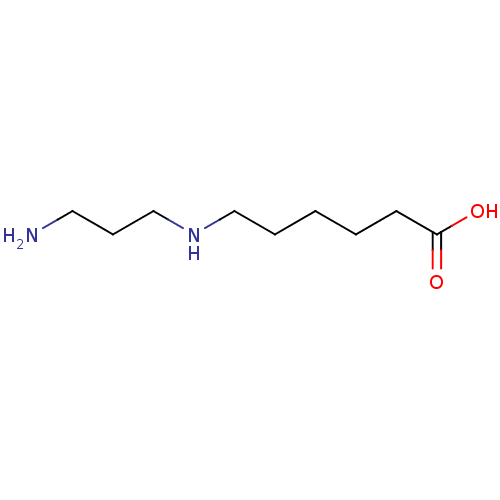
Affinity DataKi: 180nMAssay Description:Inhibition of deacetylation of [acetyl-3H]-N8-Acetylspermidine deacetylase in rat liverMore data for this Ligand-Target Pair
TargetAcetylpolyamine amidohydrolase(Mycoplana ramosa (Gram-negative bacterium))
University of Pennsylvania
University of Pennsylvania
Affinity DataIC50: 390nMAssay Description:Inhibition of Mycoplana ramosa APAH expressed in Escherichia coli BL21 (DE3) using BML-KI104 as substrate after 30 mins by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMpH: 8.2Assay Description:Activity assays were conducted at 25 °C and contained 500 nM HDAC8 enzyme, 150 μM Ac-Arg-His-Lys(Ac)-Lys(Ac)-aminomethylcoumarin substrate (BML-...More data for this Ligand-Target Pair
Affinity DataIC50: 5.30E+3nMpH: 8.2Assay Description:Activity assays were conducted at 25 °C and contained 500 nM HDAC8 enzyme, 150 μM Ac-Arg-His-Lys(Ac)-Lys(Ac)-aminomethylcoumarin substrate (BML-...More data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+4nMpH: 8.2Assay Description:Activity assays were conducted at 25 °C and contained 500 nM HDAC8 enzyme, 150 μM Ac-Arg-His-Lys(Ac)-Lys(Ac)-aminomethylcoumarin substrate (BML-...More data for this Ligand-Target Pair
TargetAcetylpolyamine amidohydrolase(Mycoplana ramosa (Gram-negative bacterium))
University of Pennsylvania
University of Pennsylvania
Affinity DataIC50: 1.60E+5nMAssay Description:Inhibition of Mycoplana ramosa APAH expressed in Escherichia coli BL21 (DE3) using BML-KI104 as substrate after 30 mins by fluorimetric assayMore data for this Ligand-Target Pair
TargetAcetylpolyamine amidohydrolase(Mycoplana ramosa (Gram-negative bacterium))
University of Pennsylvania
University of Pennsylvania
Affinity DataIC50: 1.80E+6nMAssay Description:Inhibition of Mycoplana ramosa APAH expressed in Escherichia coli BL21 (DE3) using BML-KI104 as substrate after 30 mins by fluorimetric assayMore data for this Ligand-Target Pair
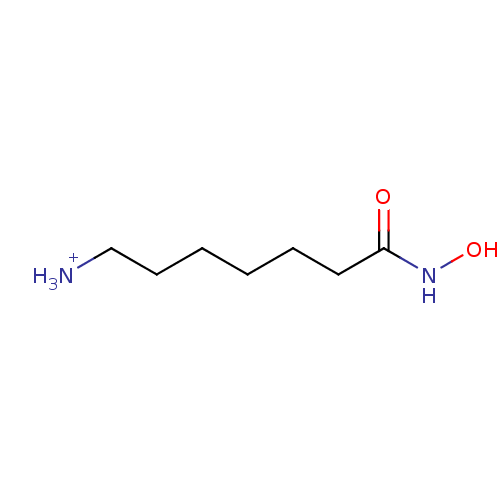
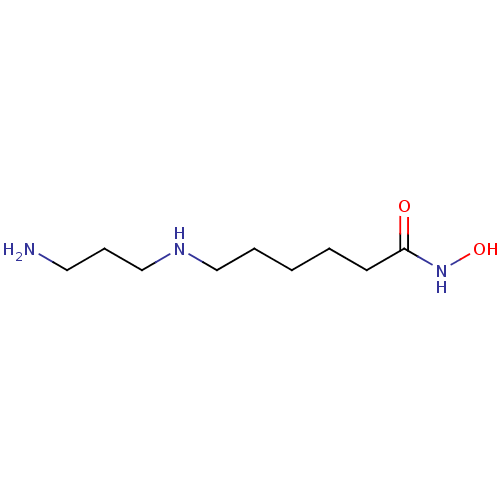
 3D Structure (crystal)
3D Structure (crystal)