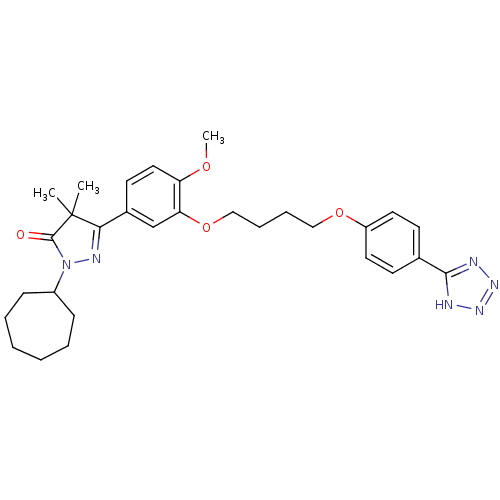
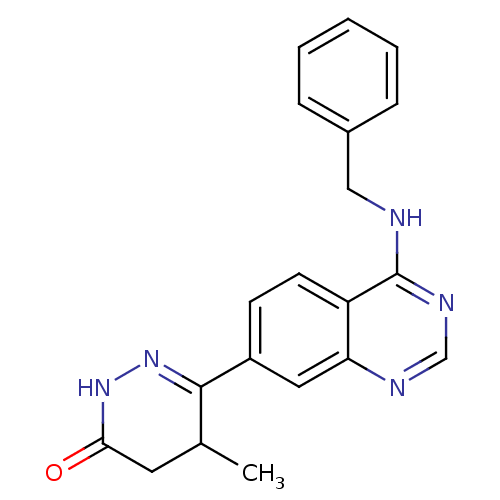
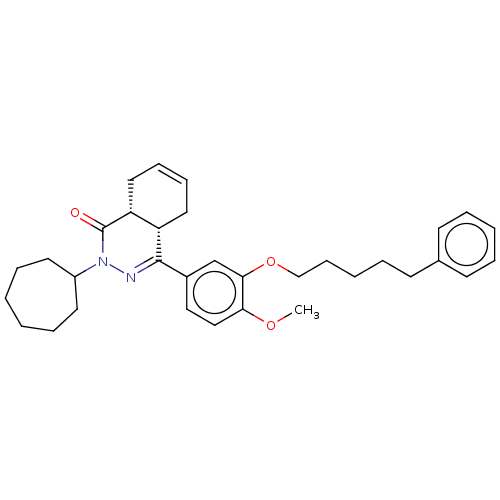
Found 570 of ic50 for PHOSPHODIESTERASE
Found 570 of ic50 for PHOSPHODIESTERASEhaving polymerids = 542,579,729,5115,6682,6810,7054,50000053,50000089,50000756,50001605,50002848,... and
complexids = 50000902,50006298
Affinity DataIC50: 0.00450nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.00680nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.00850nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0190nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0350nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0440nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0450nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0520nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0900nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0900nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.100nMAssay Description:Inhibition of rhesus monkey brain PDE2A3 using FAM-labeled cAMP as substrate after 60 mins by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.110nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.130nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.150nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.170nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.180nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.220nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.230nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.320nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.360nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.380nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.390nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.390nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.460nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.480nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.570nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.650nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.780nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.830nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 0.870nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 1.08nMpH: 8.0Assay Description:The PDE assay was performed according to a modified method referred to a report of Kotera et al. (Kotera et al., Biochem. Pharmacol., vol. 60, pp. 13...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20nMAssay Description:Inhibition of recombinant Trypanosoma brucei PDEB1 using [3H]-cAMP as substrate after 15 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of recombinant Trypanosoma brucei PDEB1 using [3H]-cAMP as substrate after 15 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of recombinant Trypanosoma brucei PDEB1 using [3H]-cAMP as substrate after 15 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of Trypanosoma brucei PDEB1More data for this Ligand-Target Pair
TargetPhosphodiesterase/cAMP-specific 3',5'-cyclic phosphodiesterase 4A/cAMP-specific 3',5'-cyclic phosphodiesterase 4C/cAMP-specific 3',5'-cyclic phosphodiesterase 4D(Mus musculus)TBA
Affinity DataIC50: 4nMAssay Description:Displacement of [3H]-Rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of Trypanosoma brucei TbrPDEB1More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of Trypanosoma brucei full length PDEB1 expressed in insect SF21 cells using cAMP as substrate by scintillation proximity assayMore data for this Ligand-Target Pair
TargetPhosphodiesterase/cAMP-specific 3',5'-cyclic phosphodiesterase 4A/cAMP-specific 3',5'-cyclic phosphodiesterase 4C/cAMP-specific 3',5'-cyclic phosphodiesterase 4D(Mus musculus)TBA
Affinity DataIC50: 4nMAssay Description:Inhibition of [3H]rolipram binding to Phosphodiesterase 4 (PDE IV)More data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of bovine arterial Phosphodiesterase 3More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of recombinant Trypanosoma brucei PDEB1 using [3H]-cAMP as substrate after 15 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of recombinant Trypanosoma brucei PDEB1 using [3H]-cAMP as substrate after 15 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Inhibition of recombinant Trypanosoma brucei PDEB1 using [3H]-cAMP as substrate after 15 mins by scintillation proximity assayMore data for this Ligand-Target Pair
TargetPhosphodiesterase/cAMP-specific 3',5'-cyclic phosphodiesterase 4A/cAMP-specific 3',5'-cyclic phosphodiesterase 4C/cAMP-specific 3',5'-cyclic phosphodiesterase 4D(Mus musculus)TBA
Affinity DataIC50: 32nMAssay Description:Displacement of [3H]-Rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 49nMAssay Description:Inhibition of Trypanosoma brucei recombinant TbrPDEB1 expressed in Sf21 insect cells assessed as reduction of [3H]-cAMP hydrolysis by scintillation p...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Inhibition of bovine arterial Phosphodiesterase 3More data for this Ligand-Target Pair
Affinity DataIC50: 55nMAssay Description:Inhibition of Trypanosoma brucei His-tagged catalytic domains of TbrPDEB1 expressed in Escherichia coli BL21 assessed as conversion of cAMP to AMP af...More data for this Ligand-Target Pair
Affinity DataIC50: 63nMAssay Description:Inhibition of recombinant Trypanosoma brucei PDEB1 using [3H]-cAMP as substrate after 15 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 63nMAssay Description:Inhibition of recombinant Trypanosoma brucei PDEB1 using [3H]-cAMP as substrate after 15 mins by scintillation proximity assayMore data for this Ligand-Target Pair

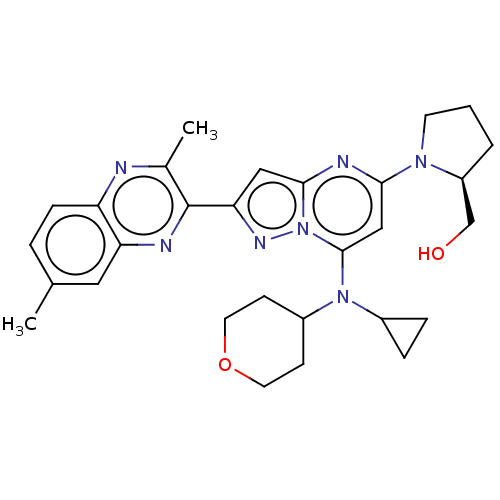
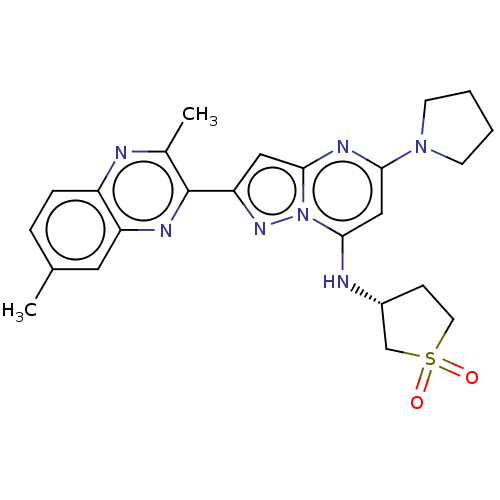
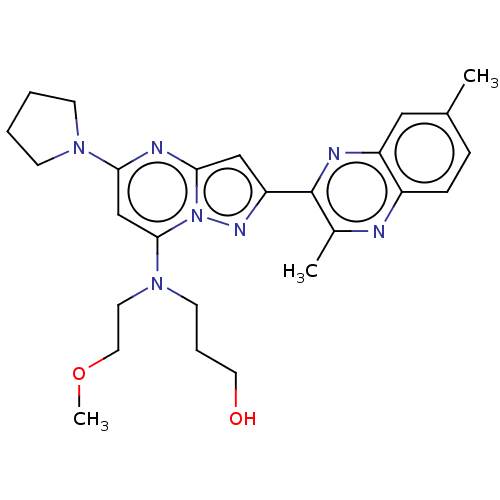
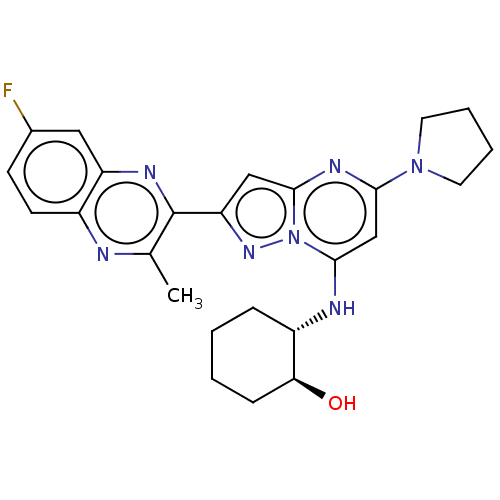
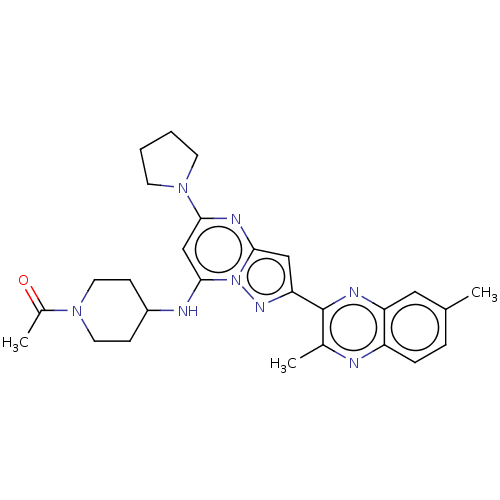
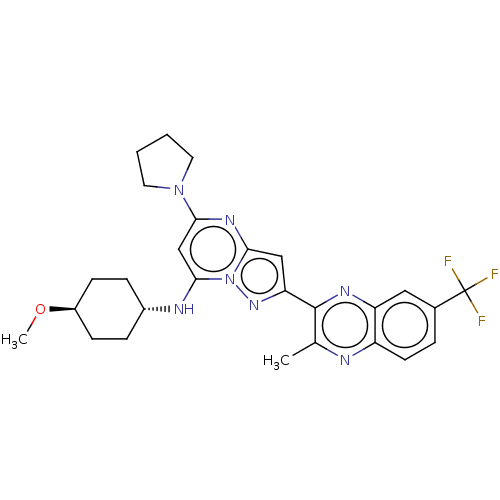
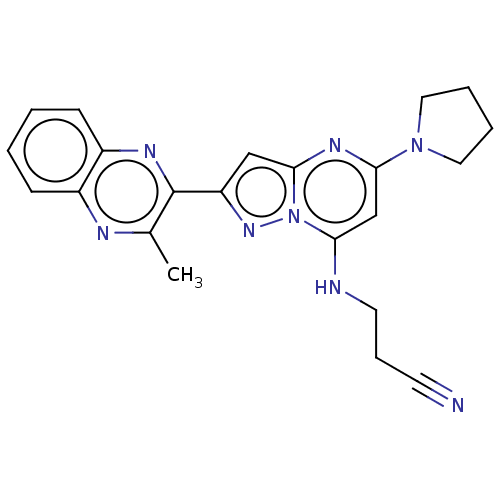

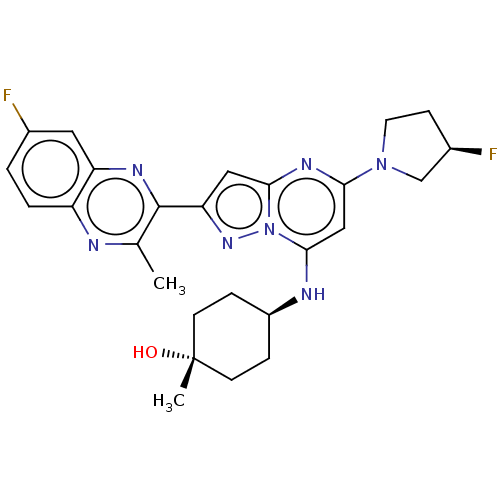

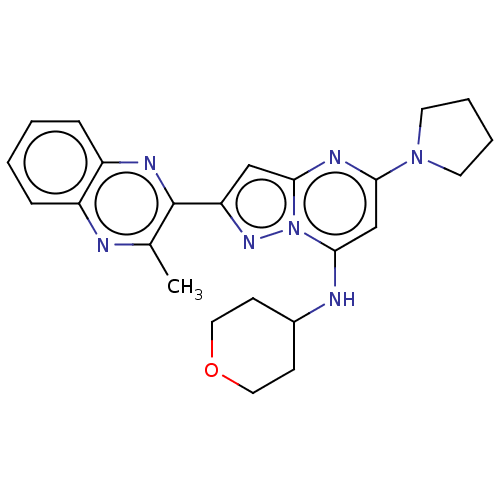
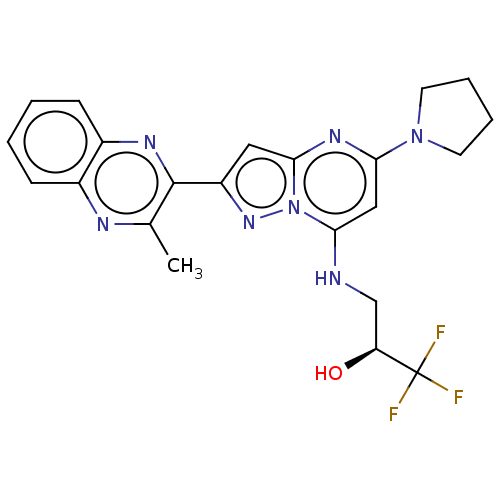

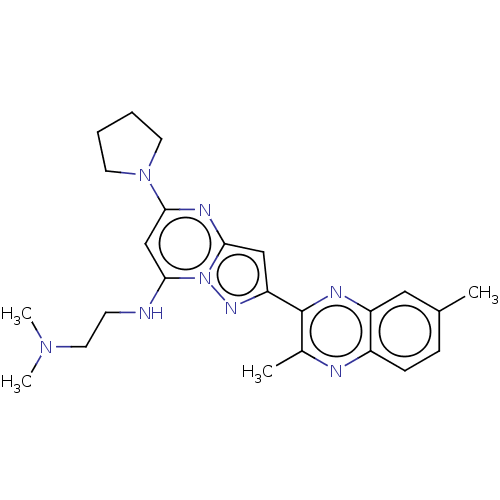
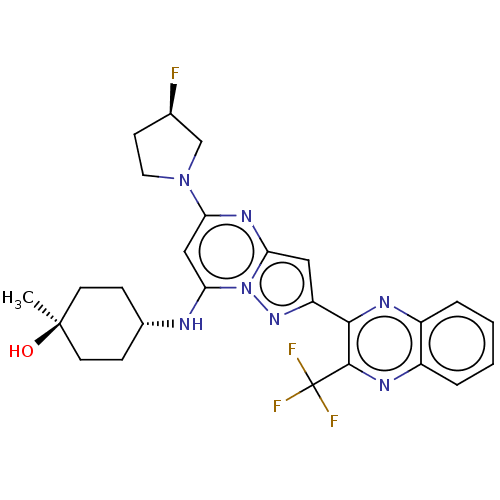
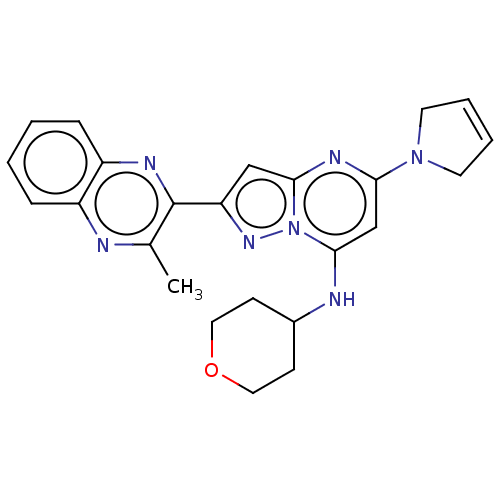
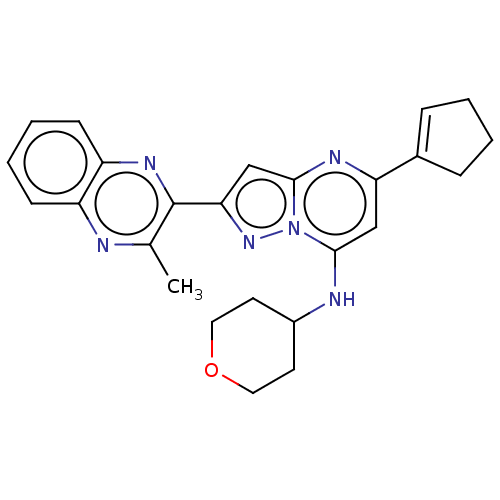
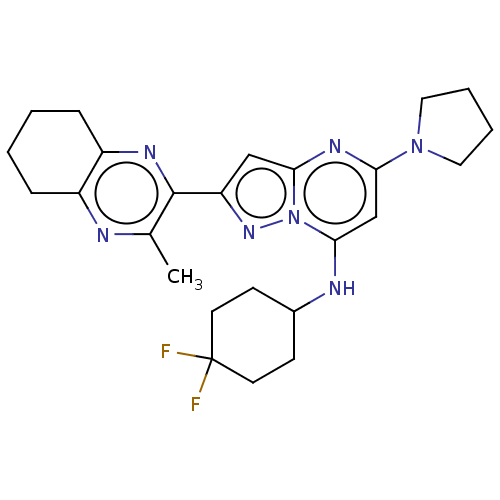
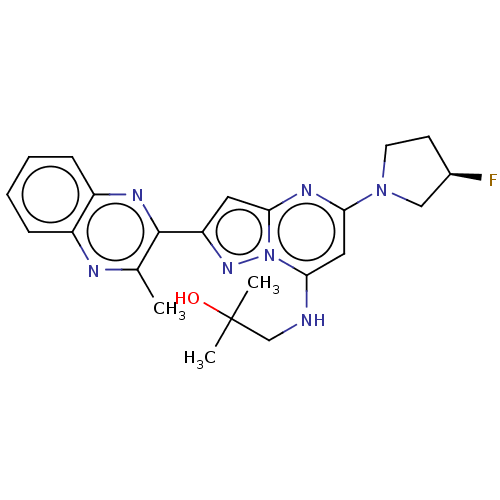
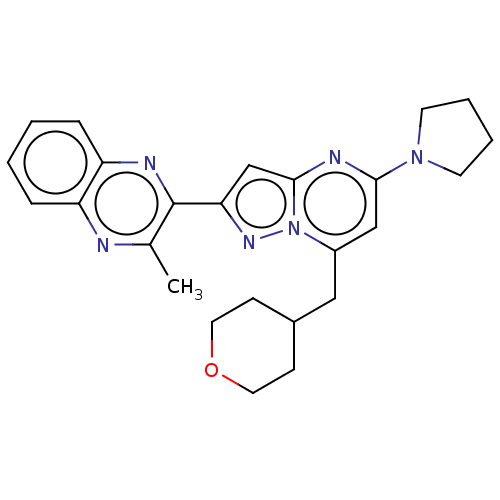
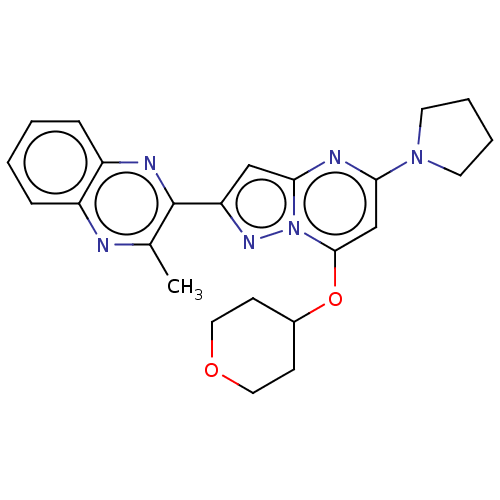

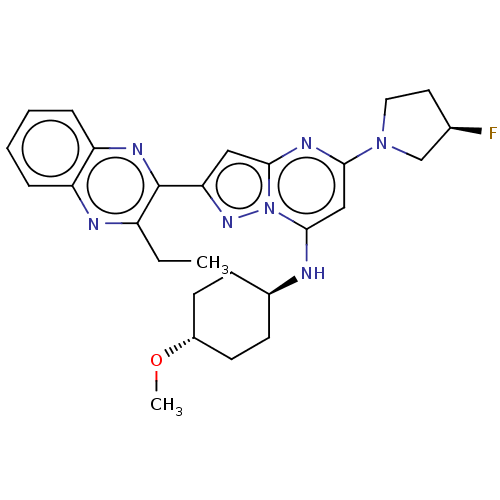
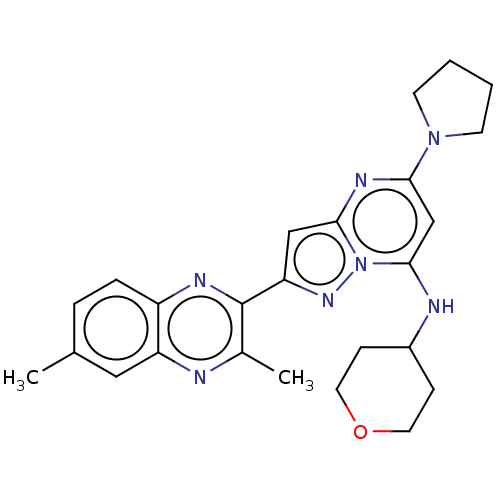
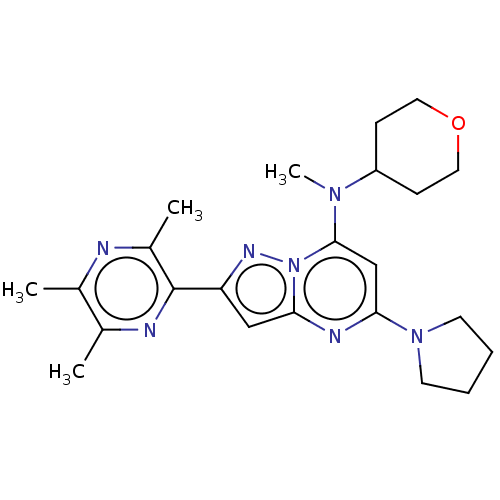
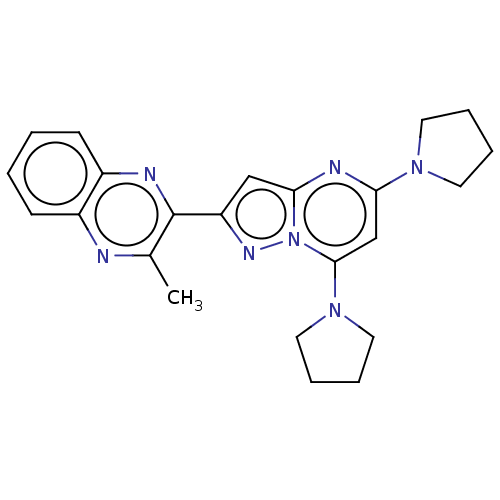
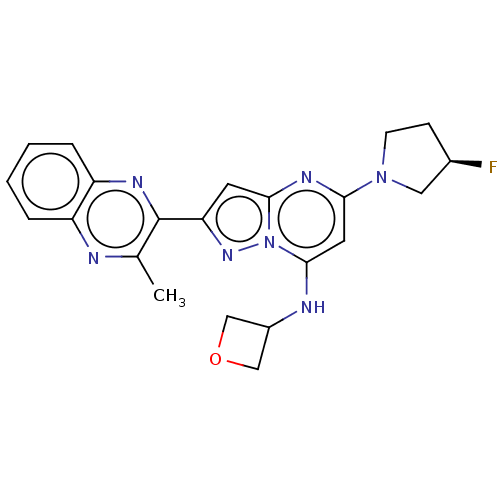

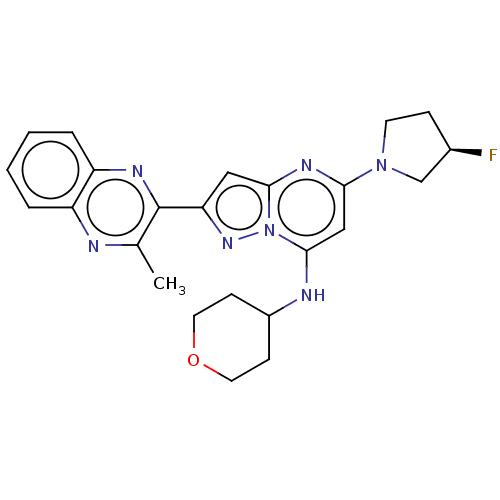
 3D Structure (crystal)
3D Structure (crystal)