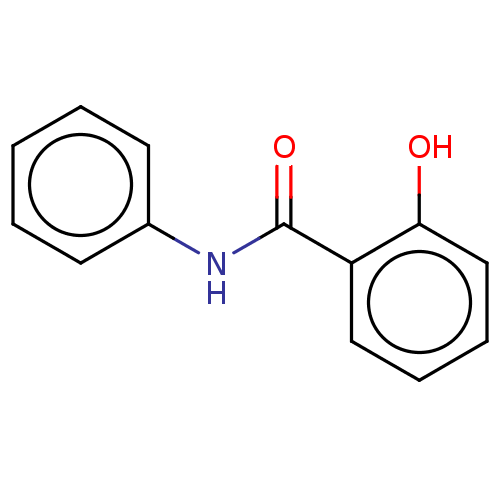
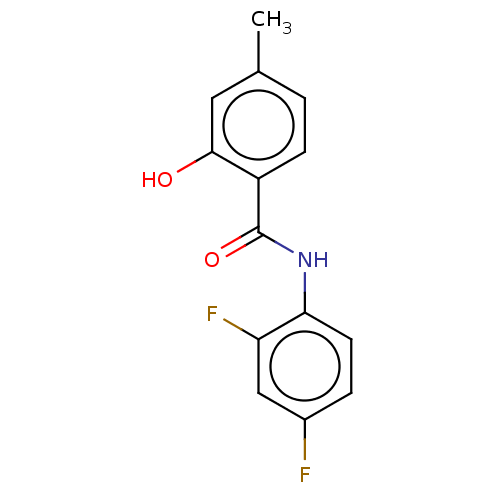
Report error Found 33 Enz. Inhib. hit(s) with all data for entry = 8127
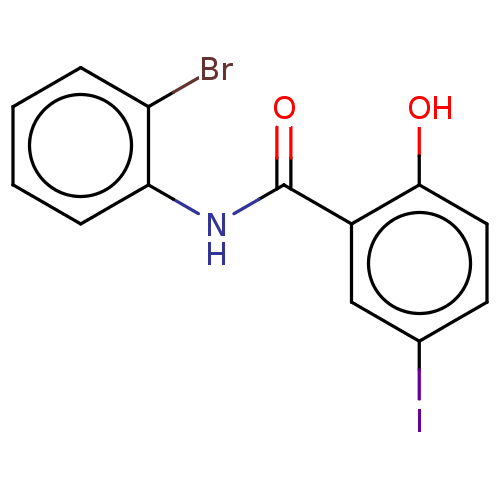
Affinity DataIC50: 320nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
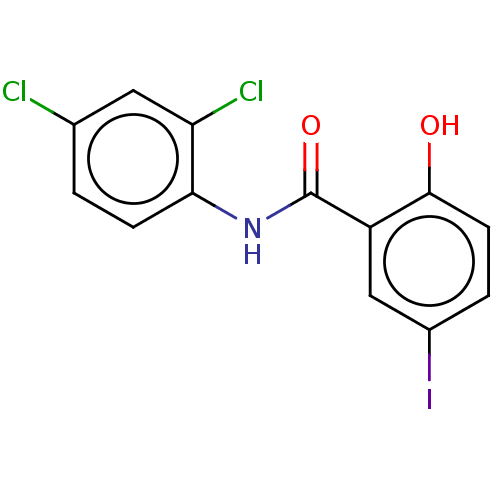
Affinity DataIC50: 630nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
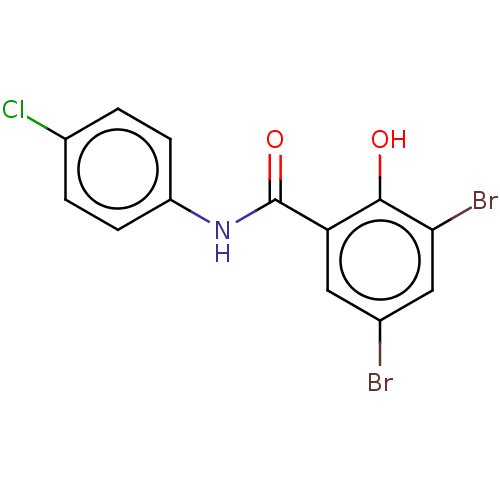
Affinity DataIC50: 920nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
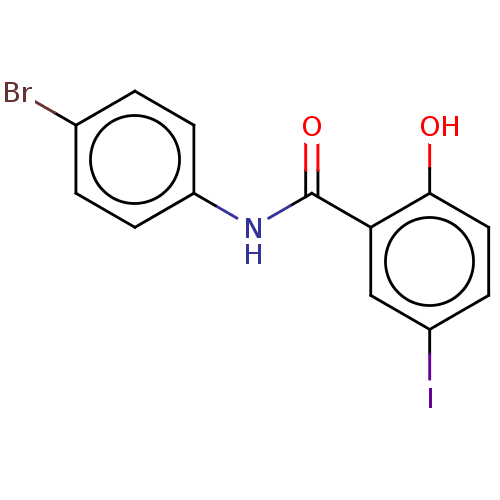
Affinity DataIC50: 2.68E+3nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 4.37E+3nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 9.66E+3nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 1.22E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 1.37E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 1.42E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 2.19E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 2.22E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 2.24E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 2.48E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 2.57E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 2.85E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 2.96E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 3.11E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 3.47E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 4.02E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 4.18E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 4.53E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 7.4 T: 2°CAssay Description:Compounds 1-32 were dissolved in 100% DMSO and diluted to the appropriate concentrations with 25 mM HEPES at pH 7.4. In each well, 10 μL of comp...More data for this Ligand-Target Pair