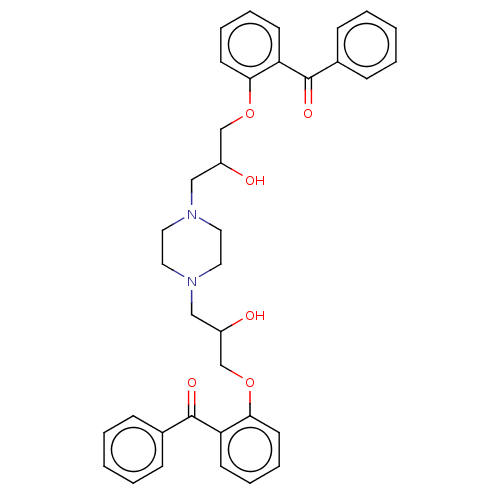
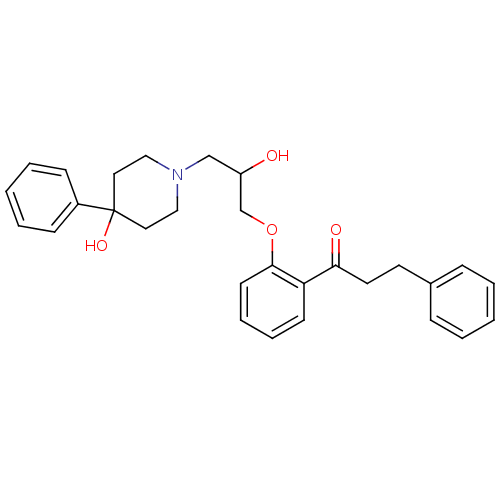
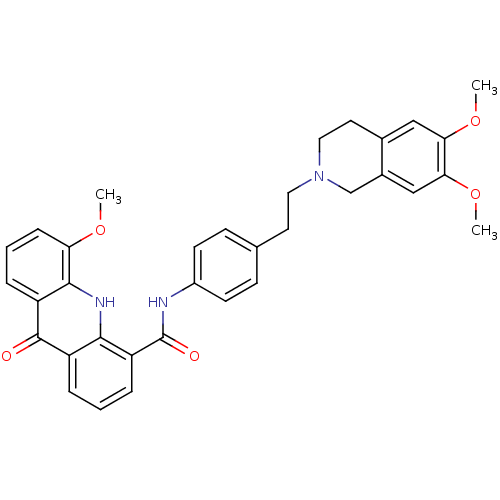
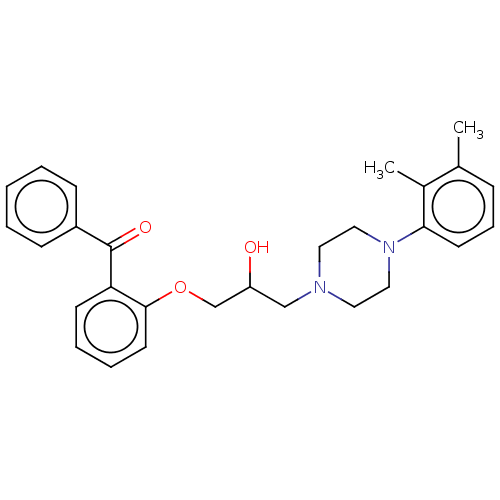
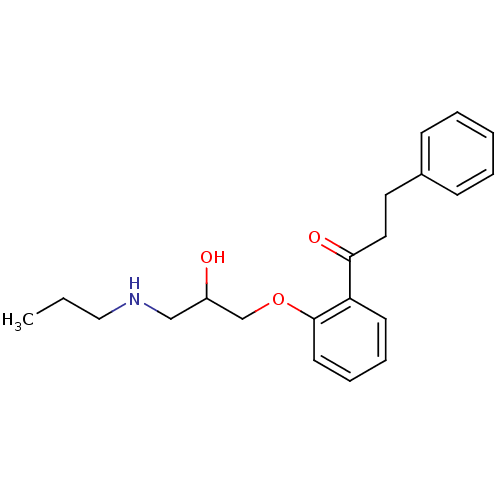
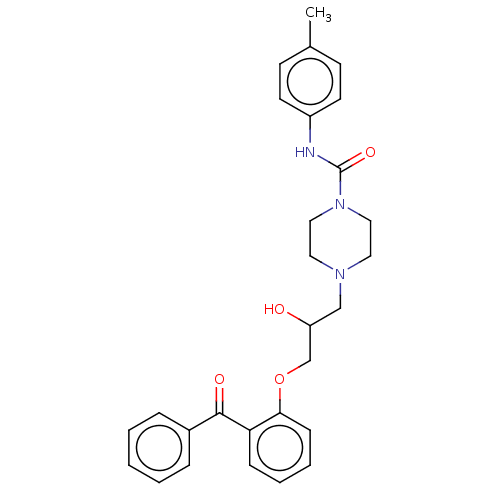
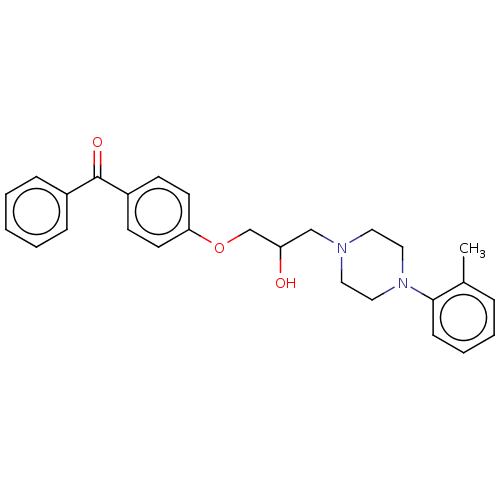
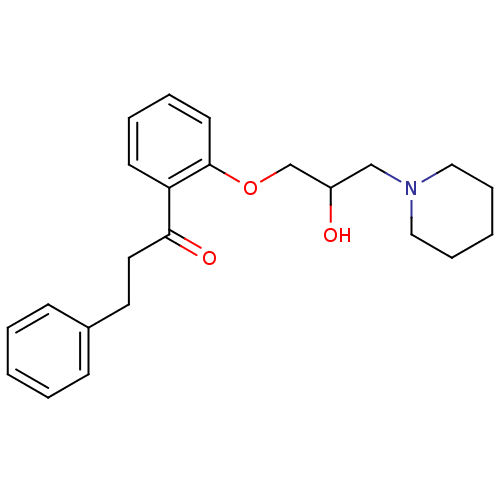
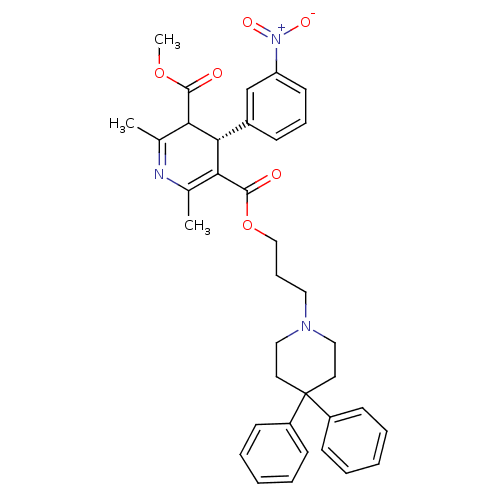
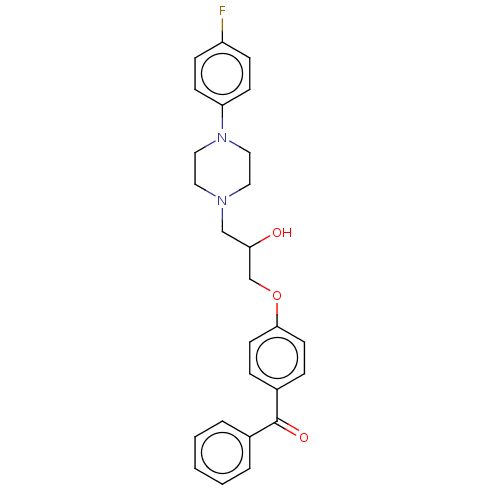
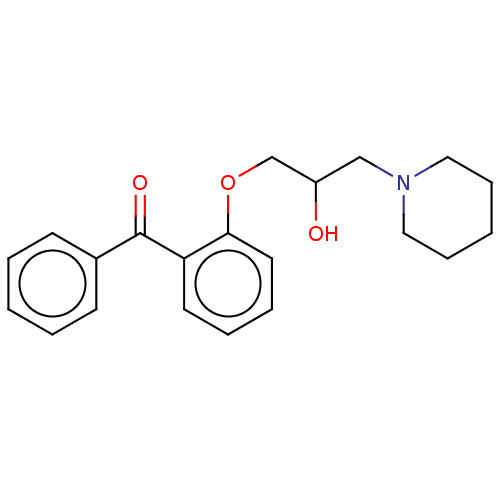
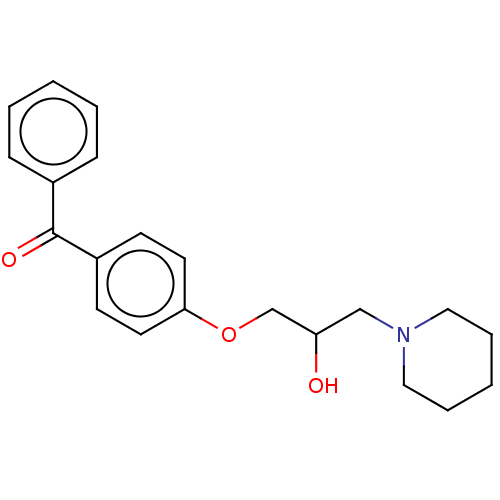
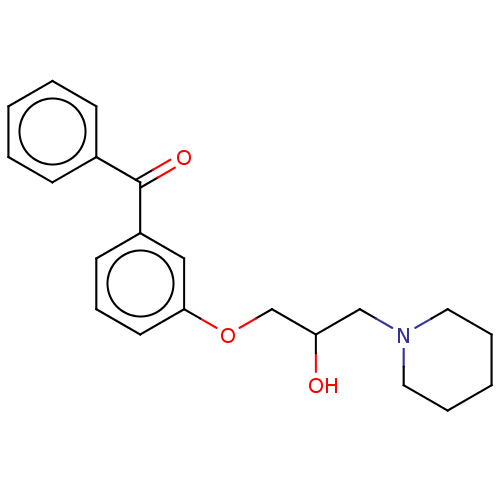
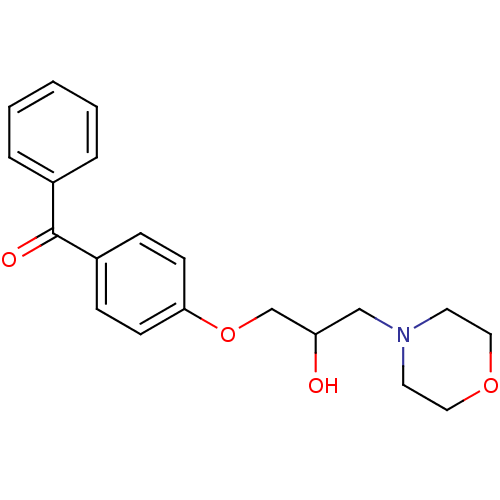
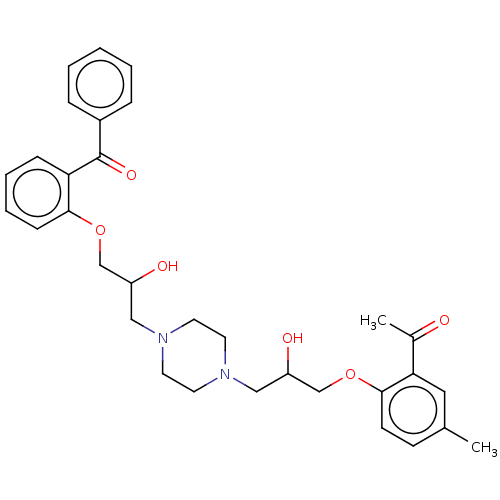
Report error Found 31 Enz. Inhib. hit(s) with all data for entry = 50004672
Affinity DataIC50: 5.60nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 32nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 33nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 58nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 59nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 72nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 80nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 102nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 310nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 331nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 380nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 480nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 500nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 501nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 575nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 580nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 603nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 650nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 708nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 970nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.21E+3nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.18E+3nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 3.55E+3nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 5.32E+3nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 9.48E+3nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.34E+4nMAssay Description:Inhibition of P-glycoprotein-mediated daunorubicin efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysisMore data for this Ligand-Target Pair
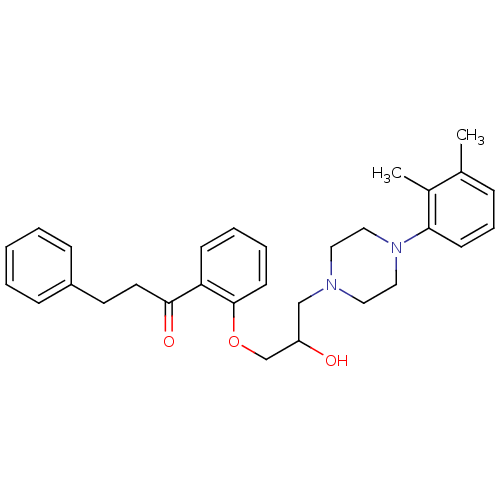

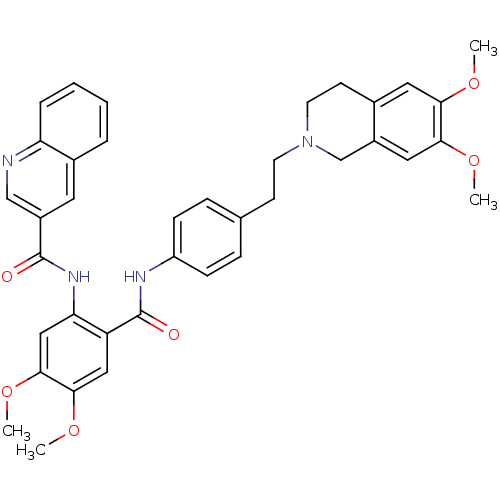
 3D Structure (crystal)
3D Structure (crystal)