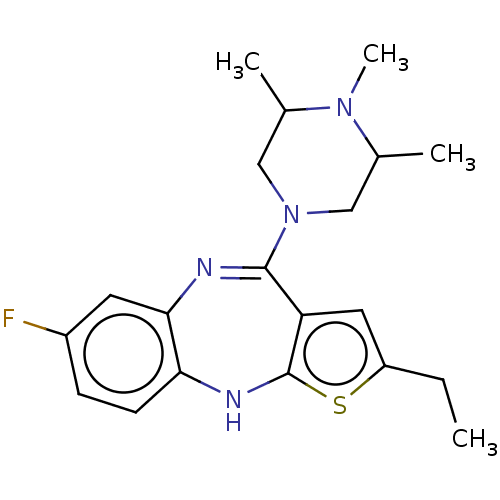
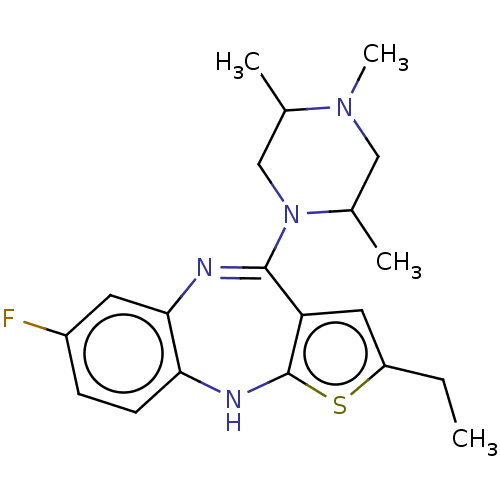
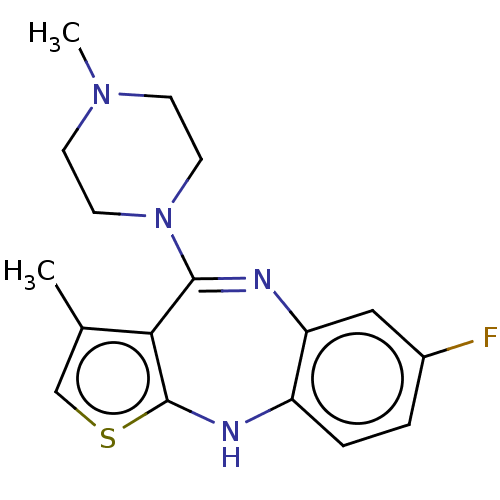
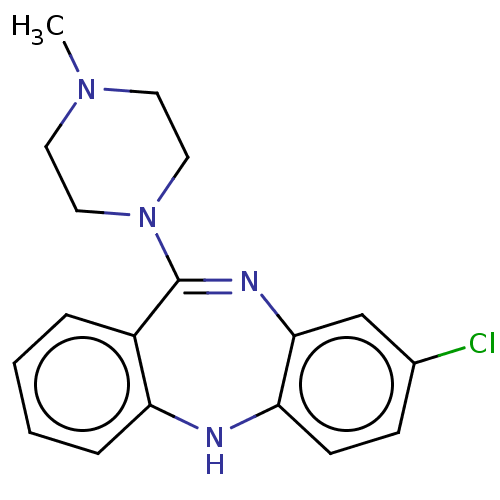
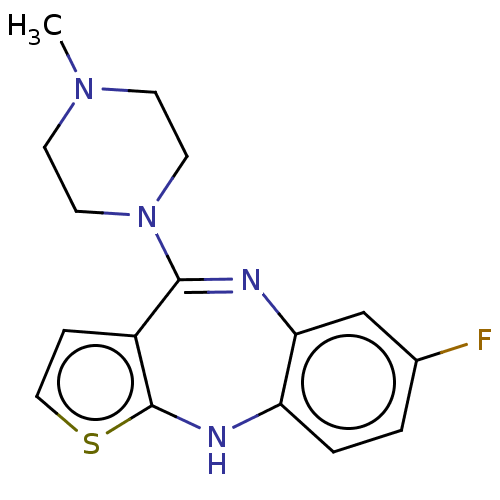
Report error Found 29 Enz. Inhib. hit(s) with all data for entry = 50048629
Affinity DataIC50: 9nMAssay Description:Displacement of [3H]QNB from Muscarinic acetylcholine receptor of rat brain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Displacement of [3H]QNB from Muscarinic acetylcholine receptor of rat brain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Displacement of [3H]spiroperidol from Dopamine receptor of rat striatum membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Displacement of [3H]spiroperidol from Dopamine receptor of rat striatum membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Displacement of [3H]spiroperidol from Dopamine receptor of rat striatum membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:Displacement of [3H]QNB from Muscarinic acetylcholine receptor of rat brain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:Displacement of [3H]QNB from Muscarinic acetylcholine receptor of rat brain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 31nMAssay Description:Displacement of [3H]spiroperidol from Dopamine receptor of rat striatum membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 34nMAssay Description:Displacement of [3H]QNB from Muscarinic acetylcholine receptor of rat brain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 64nMAssay Description:Displacement of [3H]QNB from Muscarinic acetylcholine receptor of rat brain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 74nMAssay Description:Displacement of [3H]QNB from Muscarinic acetylcholine receptor of rat brain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 75nMAssay Description:Displacement of [3H]spiroperidol from Dopamine receptor of rat striatum membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 77nMAssay Description:Displacement of [3H]spiroperidol from Dopamine receptor of rat striatum membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 91nMAssay Description:Displacement of [3H]spiroperidol from Dopamine receptor of rat striatum membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 93nMAssay Description:Displacement of [3H]spiroperidol from Dopamine receptor of rat striatum membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 97nMAssay Description:Displacement of [3H]QNB from Muscarinic acetylcholine receptor of rat brain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 101nMAssay Description:Displacement of [3H]QNB from Muscarinic acetylcholine receptor of rat brain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 116nMAssay Description:Displacement of [3H]QNB from Muscarinic acetylcholine receptor of rat brain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 172nMAssay Description:Displacement of [3H]spiroperidol from Dopamine receptor of rat striatum membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 179nMAssay Description:Displacement of [3H]QNB from Muscarinic acetylcholine receptor of rat brain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 180nMAssay Description:Displacement of [3H]spiroperidol from Dopamine receptor of rat striatum membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 199nMAssay Description:Displacement of [3H]spiroperidol from Dopamine receptor of rat striatum membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 223nMAssay Description:Displacement of [3H]spiroperidol from Dopamine receptor of rat striatum membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 247nMAssay Description:Displacement of [3H]spiroperidol from Dopamine receptor of rat striatum membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 247nMAssay Description:Displacement of [3H]QNB from Muscarinic acetylcholine receptor of rat brain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 258nMAssay Description:Displacement of [3H]QNB from Muscarinic acetylcholine receptor of rat brain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 574nMAssay Description:Displacement of [3H]QNB from Muscarinic acetylcholine receptor of rat brain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 624nMAssay Description:Displacement of [3H]spiroperidol from Dopamine receptor of rat striatum membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:The ability to displace [3H]QNB from Muscarinic acetylcholine receptor in rat brain membrane was determinedMore data for this Ligand-Target Pair