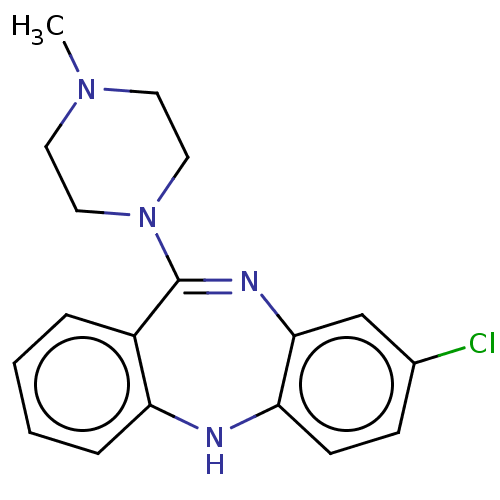
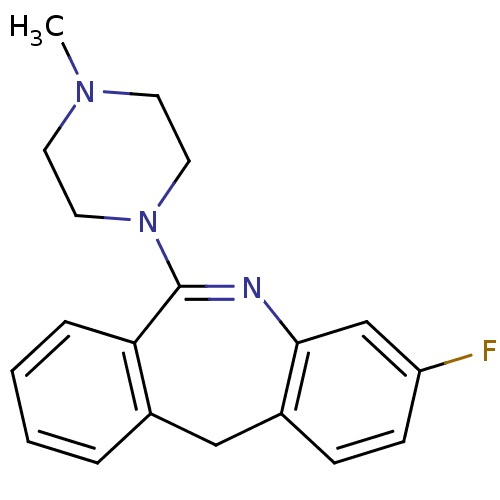
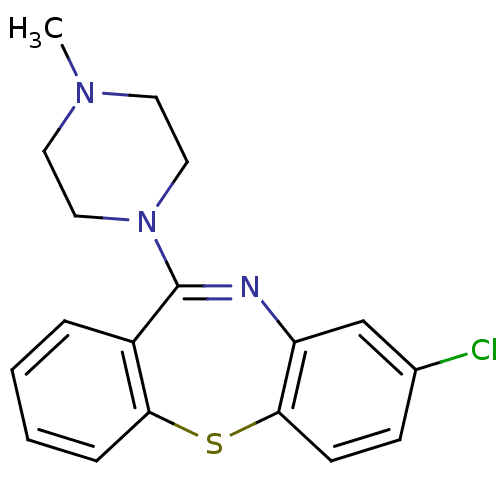
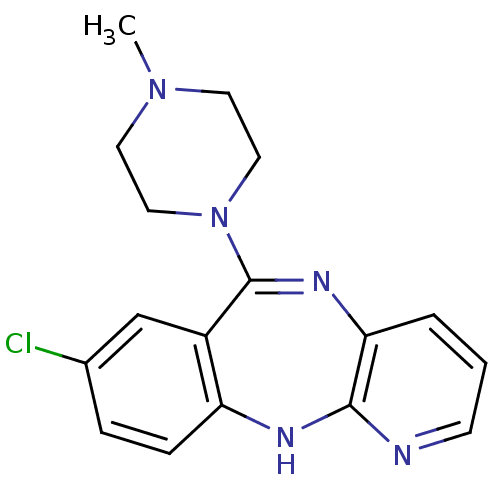
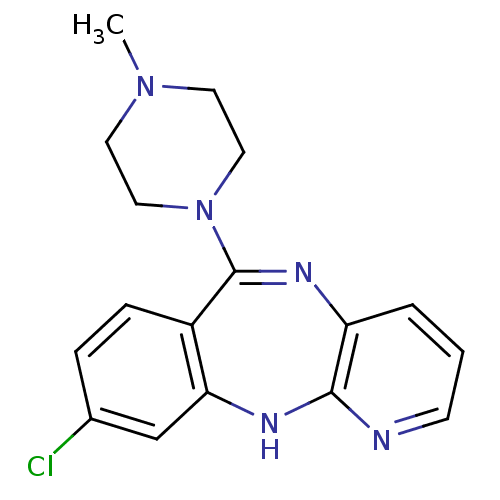
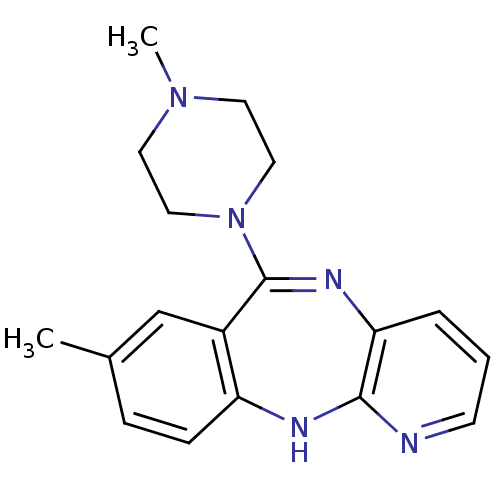
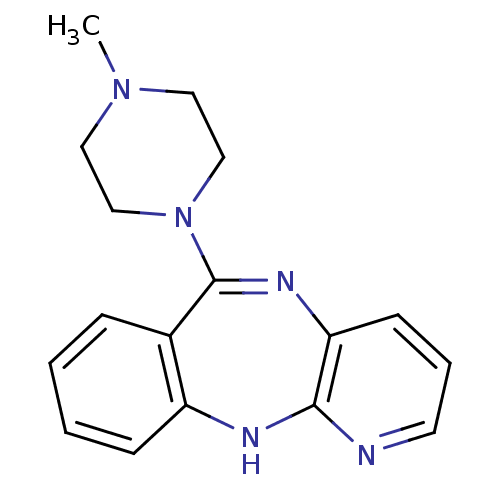
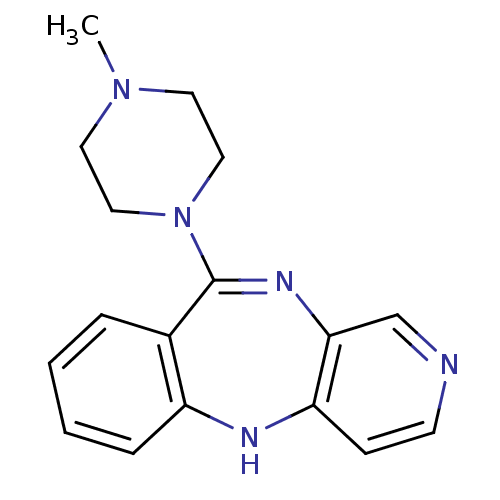
Report error Found 80 Enz. Inhib. hit(s) with all data for entry = 50005725
Affinity DataKi: 0.600nMAssay Description:In vitro binding affinity against rat 5-hydroxytryptamine 2 receptor.More data for this Ligand-Target Pair
Affinity DataKi: 0.880nMAssay Description:In vitro binding affinity against Dopamine D2 receptor in rat striatal tissue.More data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:In vitro binding affinity against rat 5-hydroxytryptamine 2 receptor.More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:In vitro binding affinity against Dopamine D2 receptor in rat striatal tissue.More data for this Ligand-Target Pair
Affinity DataKi: 3.30nMAssay Description:In vitro binding affinity against rat 5-hydroxytryptamine 2 receptor.More data for this Ligand-Target Pair
Affinity DataKi: 3.80nMAssay Description:In vitro binding affinity against rat 5-hydroxytryptamine 2 receptor.More data for this Ligand-Target Pair
Affinity DataKi: 3.80nMAssay Description:In vitro binding affinity against rat 5-hydroxytryptamine 2 receptor.More data for this Ligand-Target Pair
Affinity DataKi: 4.10nMAssay Description:In vitro binding affinity against rat 5-hydroxytryptamine 2 receptor.More data for this Ligand-Target Pair
Affinity DataKi: 4.40nMAssay Description:In vitro binding affinity against Dopamine D2 receptor in rat striatal tissue.More data for this Ligand-Target Pair
Affinity DataKi: 5.60nMAssay Description:In vitro binding affinity against Muscarinic acetylcholine receptors in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 6.20nMAssay Description:In vitro binding affinity against rat 5-hydroxytryptamine 2 receptor.More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:In vitro binding affinity against Dopamine D2 receptor in rat striatal tissue.More data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:In vitro binding affinity against Dopamine receptor D1 in rat striatal tissueMore data for this Ligand-Target Pair
Affinity DataKi: 21nMAssay Description:In vitro binding affinity against Muscarinic acetylcholine receptors in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:In vitro binding affinity against rat 5-hydroxytryptamine 2 receptor.More data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:In vitro binding affinity against Muscarinic acetylcholine receptors in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:In vitro binding affinity against Muscarinic acetylcholine receptors in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 28nMAssay Description:In vitro binding affinity against Dopamine receptor D1 in rat striatal tissueMore data for this Ligand-Target Pair
Affinity DataKi: 29nMAssay Description:In vitro binding affinity against rat 5-hydroxytryptamine 2 receptor.More data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:In vitro binding affinity against Dopamine D2 receptor in rat striatal tissue.More data for this Ligand-Target Pair
Affinity DataKi: 35nMAssay Description:In vitro binding affinity against Dopamine receptor D1 in rat striatal tissueMore data for this Ligand-Target Pair
Affinity DataKi: 45nMAssay Description:In vitro binding affinity against Dopamine D2 receptor in rat striatal tissue.More data for this Ligand-Target Pair
Affinity DataKi: 48nMAssay Description:In vitro binding affinity against rat 5-hydroxytryptamine 2 receptor.More data for this Ligand-Target Pair
Affinity DataKi: 48nMAssay Description:In vitro binding affinity against Muscarinic acetylcholine receptors in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 61nMAssay Description:In vitro binding affinity against Muscarinic acetylcholine receptors in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 65nMAssay Description:In vitro binding affinity against Muscarinic acetylcholine receptors in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 76nMAssay Description:In vitro binding affinity against Dopamine receptor D1 in rat striatal tissueMore data for this Ligand-Target Pair
Affinity DataKi: 94nMAssay Description:In vitro binding affinity against rat 5-hydroxytryptamine 2 receptor.More data for this Ligand-Target Pair
Affinity DataKi: 103nMAssay Description:In vitro binding affinity against rat 5-hydroxytryptamine 2 receptor.More data for this Ligand-Target Pair
Affinity DataKi: 103nMAssay Description:In vitro binding affinity against rat 5-hydroxytryptamine 2 receptor.More data for this Ligand-Target Pair
Affinity DataKi: 115nMAssay Description:In vitro binding affinity against Dopamine receptor D1 in rat striatal tissueMore data for this Ligand-Target Pair
Affinity DataKi: 118nMAssay Description:In vitro binding affinity against Dopamine D2 receptor in rat striatal tissue.More data for this Ligand-Target Pair
Affinity DataKi: 124nMAssay Description:In vitro binding affinity against rat 5-hydroxytryptamine 2 receptor.More data for this Ligand-Target Pair
Affinity DataKi: 124nMAssay Description:In vitro binding affinity against rat 5-hydroxytryptamine 2 receptor.More data for this Ligand-Target Pair
Affinity DataKi: 133nMAssay Description:In vitro binding affinity against Dopamine D2 receptor in rat striatal tissue.More data for this Ligand-Target Pair
Affinity DataKi: 136nMAssay Description:In vitro binding affinity against Muscarinic acetylcholine receptors in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 141nMAssay Description:In vitro binding affinity against Dopamine receptor D1 in rat striatal tissueMore data for this Ligand-Target Pair
Affinity DataKi: 175nMAssay Description:In vitro binding affinity against Muscarinic acetylcholine receptors in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 214nMAssay Description:In vitro binding affinity against Dopamine receptor D1 in rat striatal tissueMore data for this Ligand-Target Pair
Affinity DataKi: 240nMAssay Description:In vitro binding affinity against Dopamine receptor D1 in rat striatal tissueMore data for this Ligand-Target Pair
Affinity DataKi: 297nMAssay Description:In vitro binding affinity against Dopamine D2 receptor in rat striatal tissue.More data for this Ligand-Target Pair
Affinity DataKi: 302nMAssay Description:In vitro binding affinity against Dopamine receptor D1 in rat striatal tissueMore data for this Ligand-Target Pair
Affinity DataKi: 303nMAssay Description:In vitro binding affinity against Muscarinic acetylcholine receptors in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 311nMAssay Description:In vitro binding affinity against Muscarinic acetylcholine receptors in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 366nMAssay Description:In vitro binding affinity against Muscarinic acetylcholine receptors in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 398nMAssay Description:In vitro binding affinity against Dopamine receptor D1 in rat striatal tissueMore data for this Ligand-Target Pair
Affinity DataKi: 415nMAssay Description:In vitro binding affinity against Muscarinic acetylcholine receptors in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 430nMAssay Description:In vitro binding affinity against Muscarinic acetylcholine receptors in rat brain.More data for this Ligand-Target Pair
Affinity DataKi: 442nMAssay Description:In vitro binding affinity against Dopamine D2 receptor in rat striatal tissue.More data for this Ligand-Target Pair
Affinity DataKi: 523nMAssay Description:In vitro binding affinity against Muscarinic acetylcholine receptors in rat brain.More data for this Ligand-Target Pair