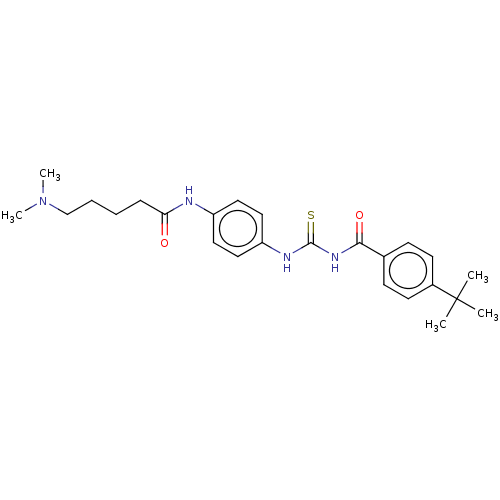
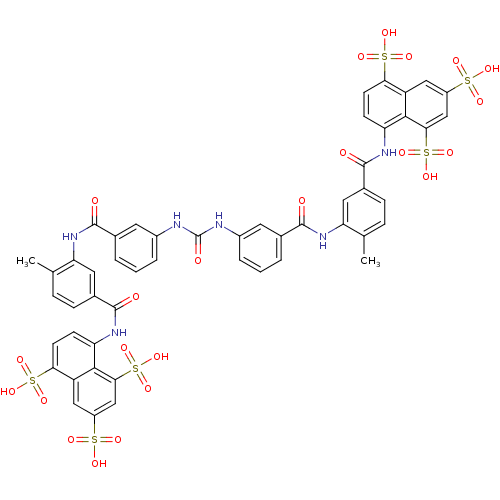
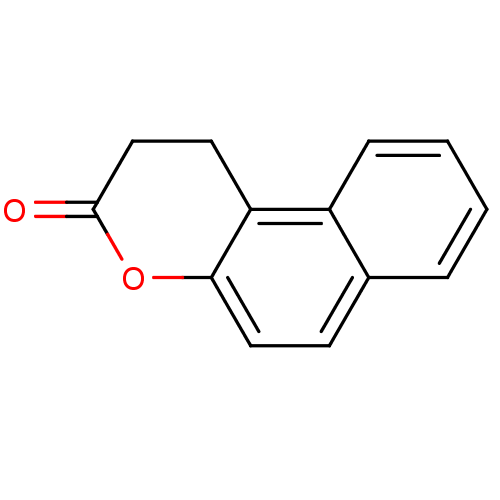
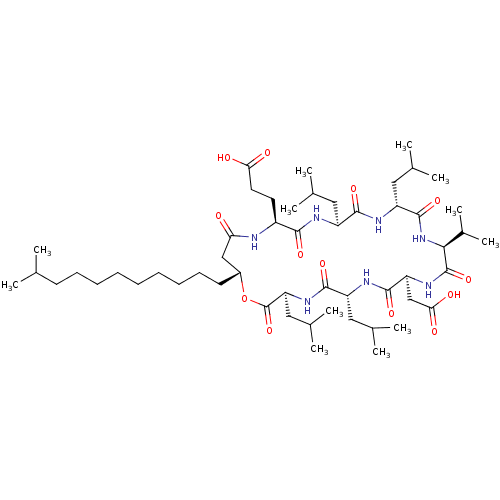
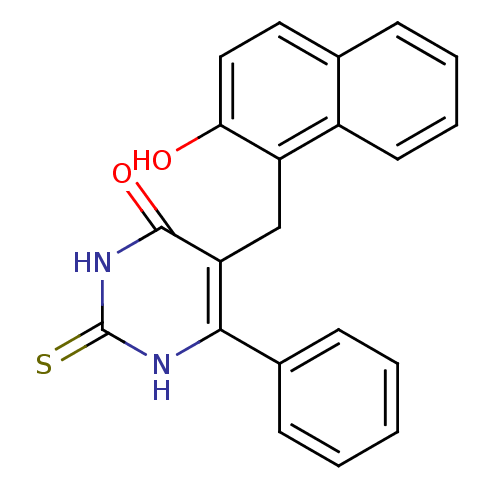
Report error Found 35 Enz. Inhib. hit(s) with Target = 'NAD-dependent protein deacetylase HST2'
Affinity DataKi: 1.20E+3nMAssay Description:Noncompetitive inhibition of yeast Hst2 using NAD+ substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:Inhibition of yeast Hst2 by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.50E+3nMAssay Description:Noncompetitive inhibition of yeast Hst2 using NAD+ substrateMore data for this Ligand-Target Pair
Affinity DataKi: 2.50E+3nMAssay Description:Mixed inhibition of yeast Hst2 using Acetyl-lysine substrateMore data for this Ligand-Target Pair
Affinity DataKd: 3.30E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
Affinity DataKd: 3.30E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
Affinity DataKd: 3.30E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
Affinity DataKd: 3.30E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
Affinity DataKd: 3.30E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
Affinity DataKd: 3.30E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
Affinity DataKd: 3.30E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
Affinity DataKd: 3.30E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
Affinity DataKi: 6.30E+3nMAssay Description:Noncompetitive inhibition of yeast Hst2 using Acetyl-lysine substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 6.50E+3nMAssay Description:Inhibition of yeast Hst2 by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataKd: 8.60E+3nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
Affinity DataIC50: 1.25E+4nMAssay Description:Inhibition of yeast Hst2 by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.45E+4nMAssay Description:Inhibition of yeast Hst2 by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataKd: 1.60E+4nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
Affinity DataIC50: 1.99E+4nMAssay Description:Inhibition of yeast Hst2 by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.99E+4nMAssay Description:Inhibition of yeast Hst2 by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of yeast Hst2 by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataKd: 2.10E+4nMAssay Description:Binding affinity to Saccharomyces cerevisiae Hst2 by isothermal titration calorimetryMore data for this Ligand-Target Pair
Affinity DataKi: 3.00E+4nMAssay Description:Noncompetitive inhibition of yeast Hst2 using NAD+ substrateMore data for this Ligand-Target Pair
Affinity DataKi: 3.90E+4nMAssay Description:Noncompetitive inhibition of yeast Hst2 using Acetyl-lysine substrateMore data for this Ligand-Target Pair
Affinity DataKi: 4.20E+4nMAssay Description:Mixed inhibition of yeast Hst2 using NAD+ substrateMore data for this Ligand-Target Pair
Affinity DataKi: 4.30E+4nMAssay Description:Noncompetitive inhibition of yeast Hst2 using Acetyl-lysine substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 4.80E+4nMAssay Description:Inhibition of yeast Hst2 by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 9.10E+4nMAssay Description:Inhibition of yeast Hst2 by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of yeast Hst2 by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+5nMAssay Description:Inhibition of yeast Hst2 by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+5nMAssay Description:Inhibition of yeast Hst2 by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.60E+5nMAssay Description:Inhibition of yeast Hst2 by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+5nMAssay Description:Inhibition of yeast Hst2 by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7.00E+5nMAssay Description:Inhibition of yeast Hst2 by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of yeast Hst2 by fluorimetric assayMore data for this Ligand-Target Pair
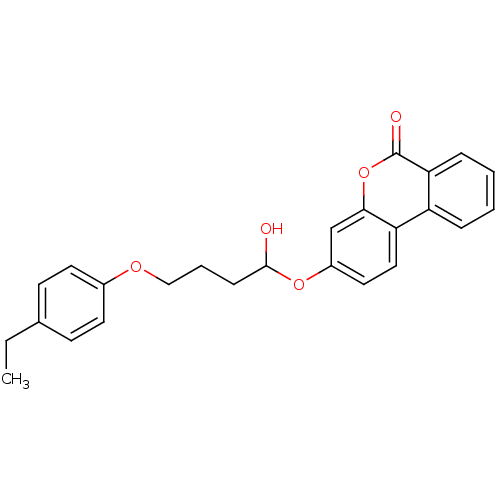
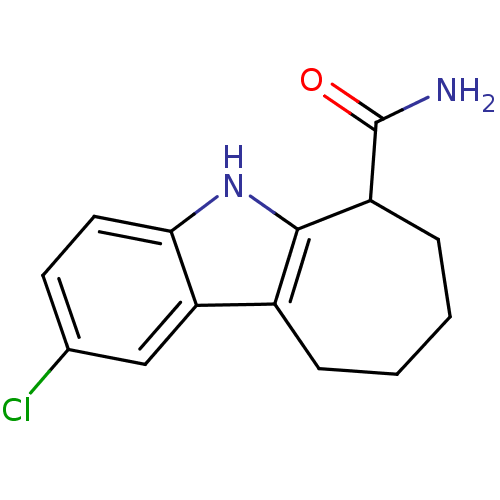
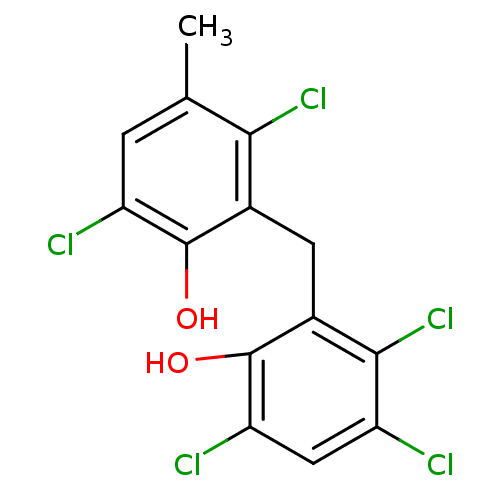





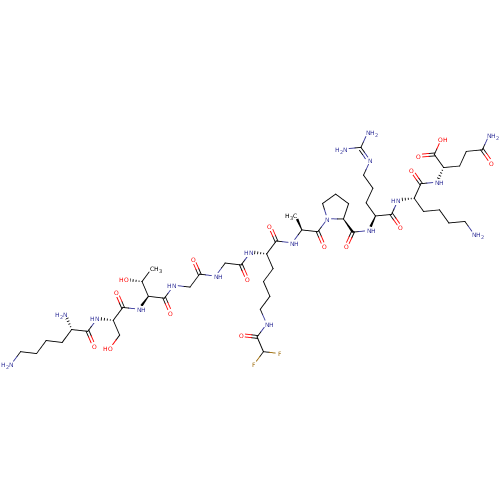


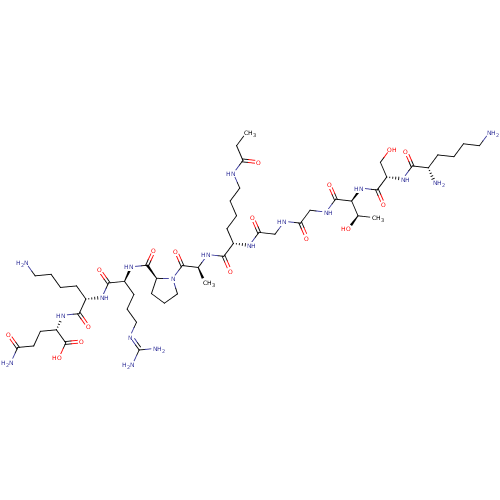


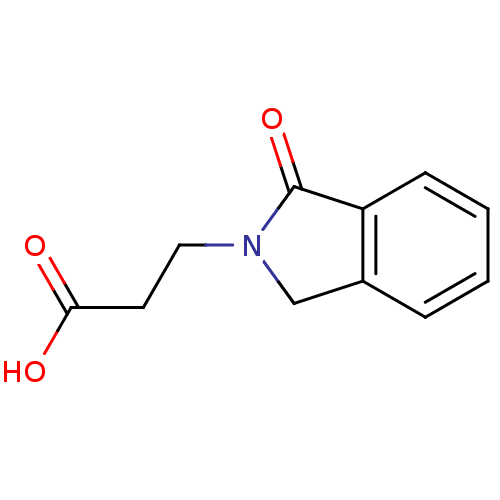
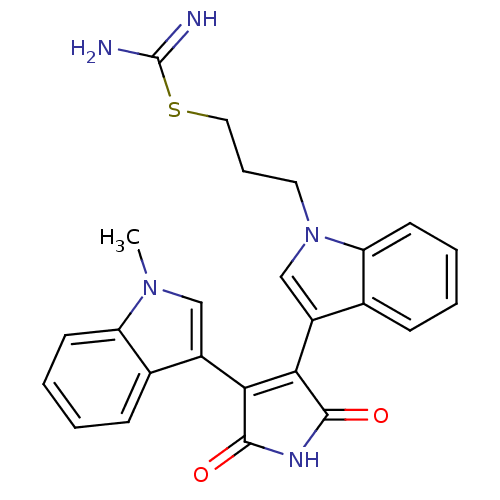




 3D Structure (crystal)
3D Structure (crystal)