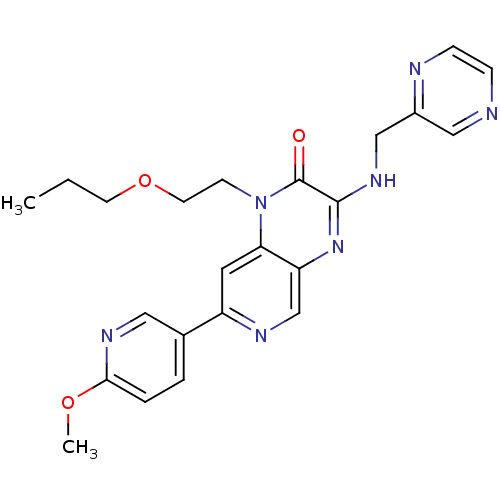
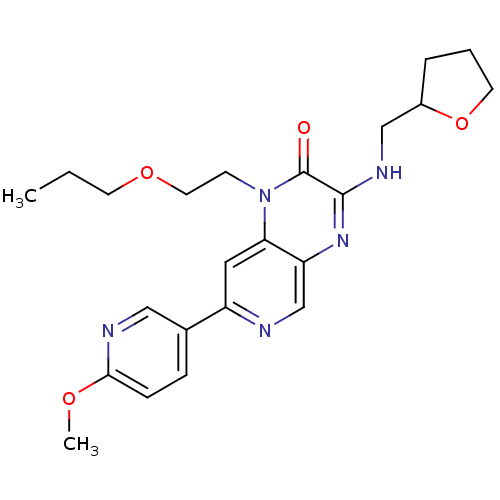
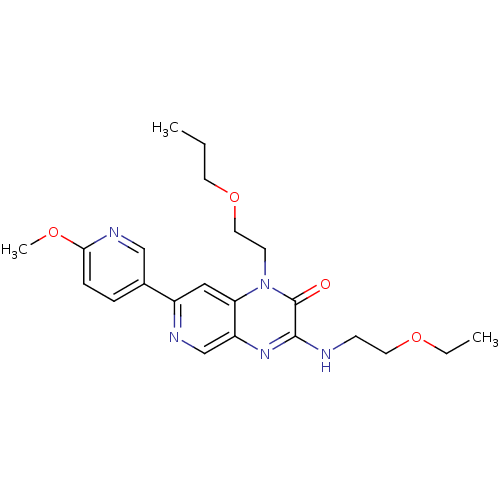
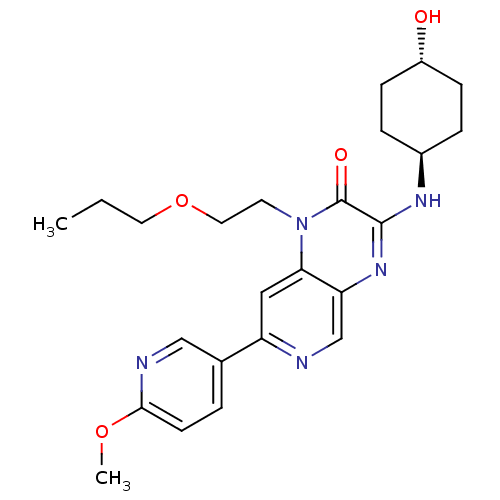
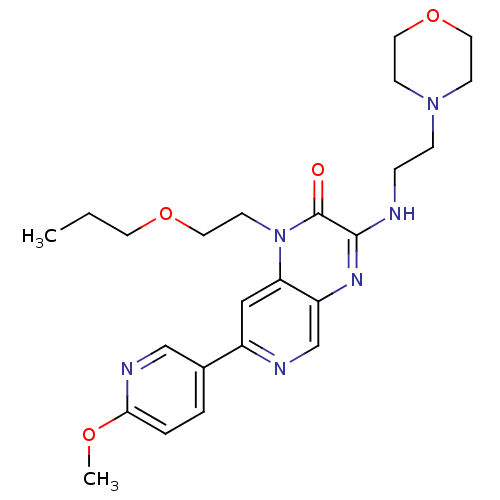
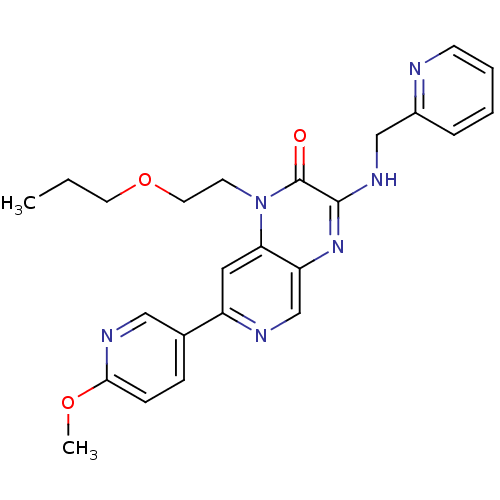
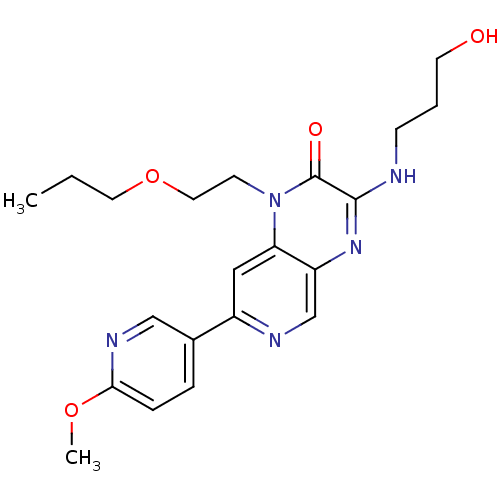
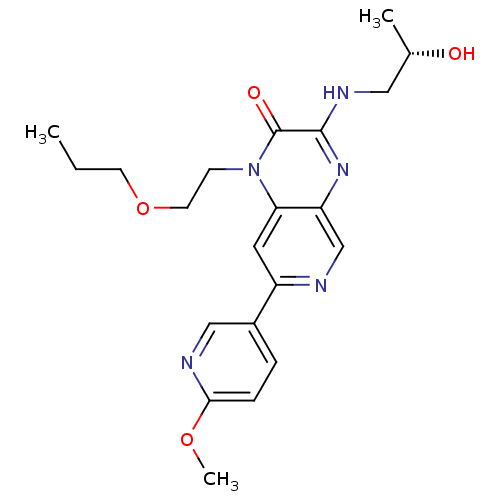
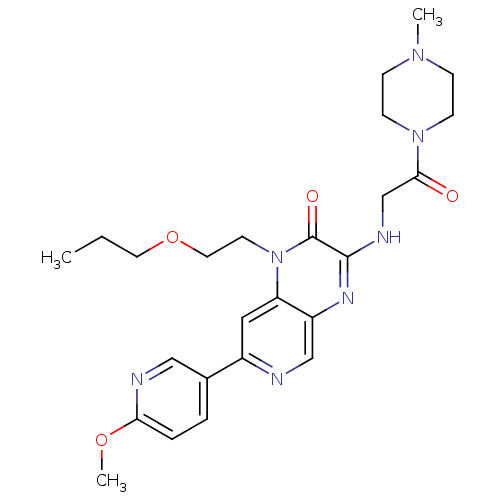
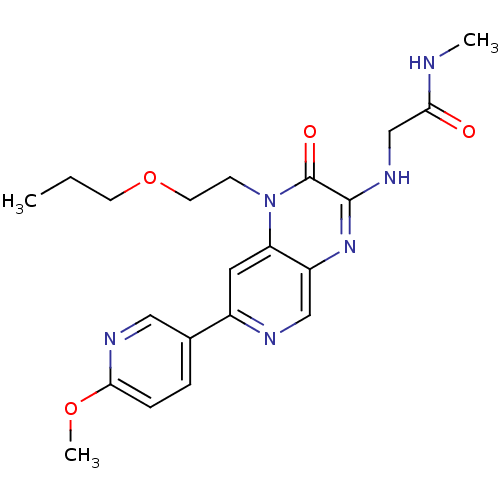
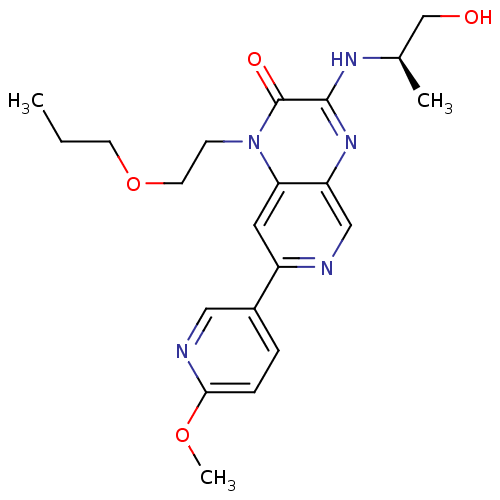
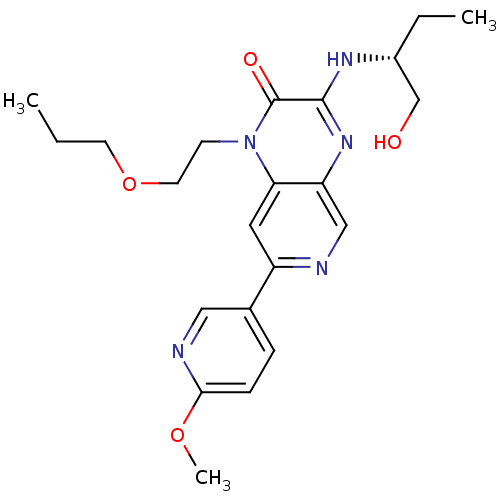
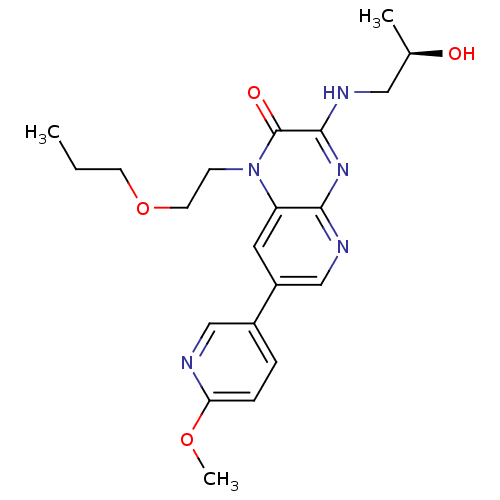
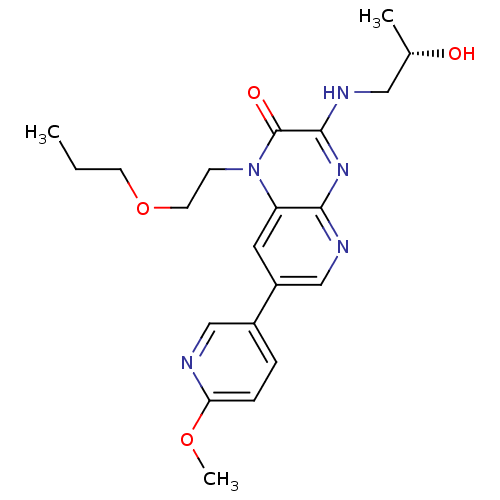
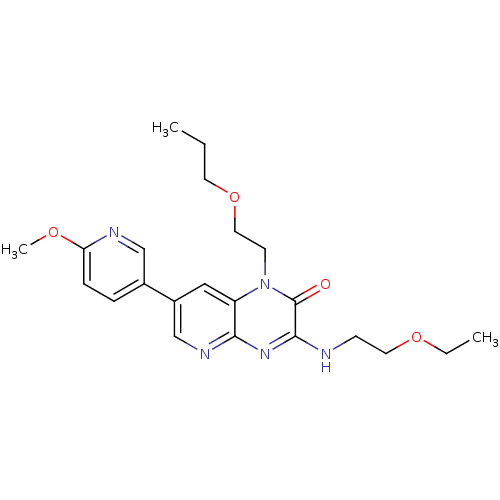
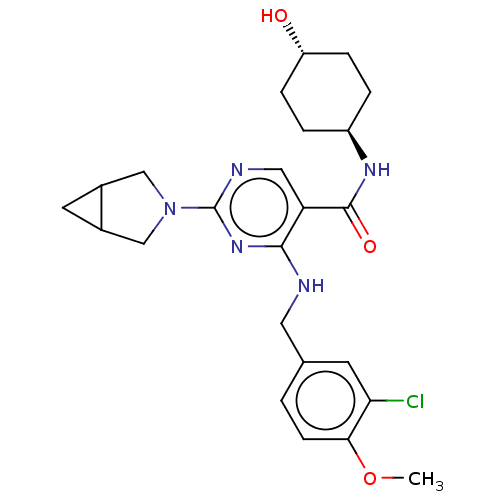
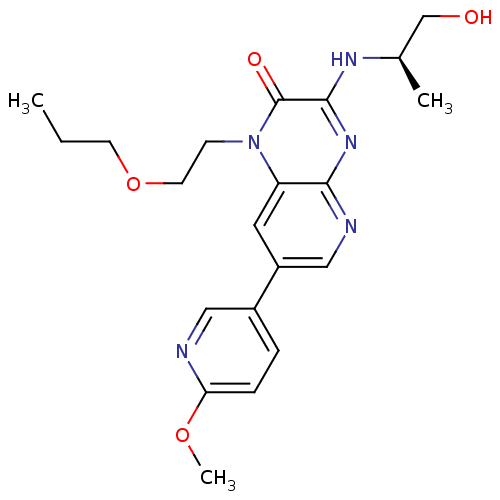
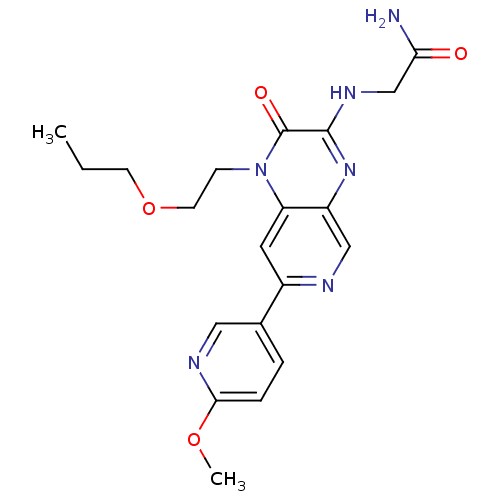
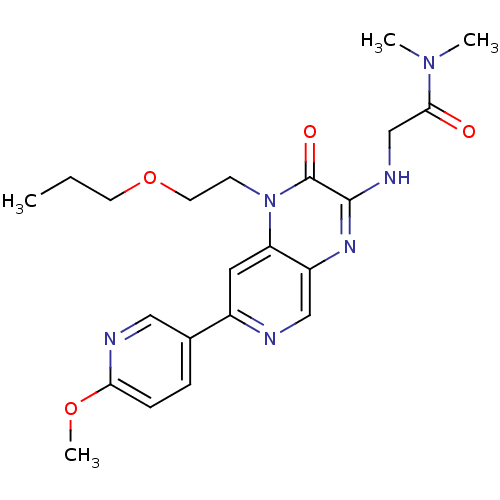
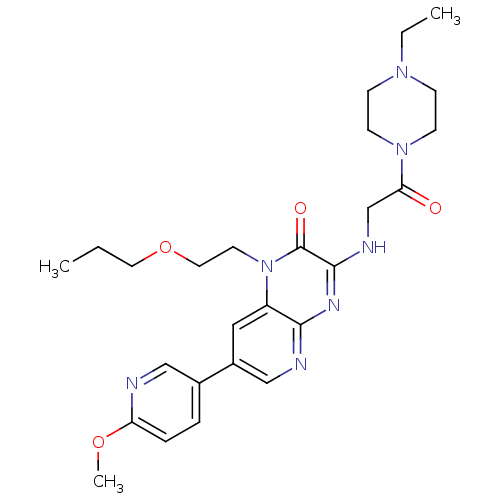
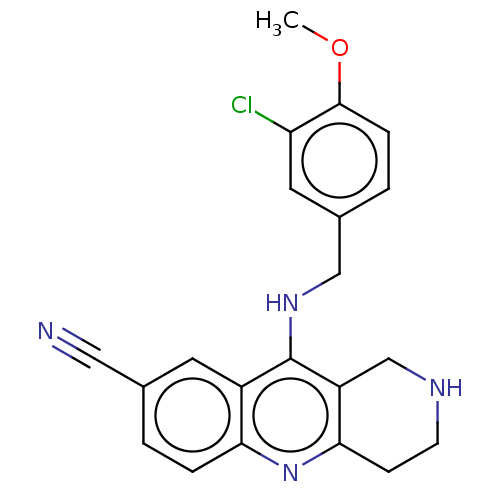
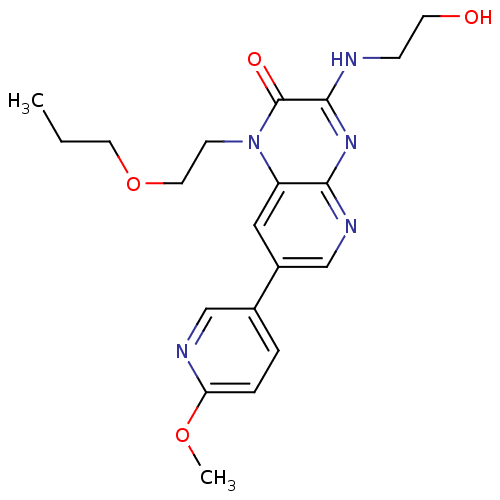
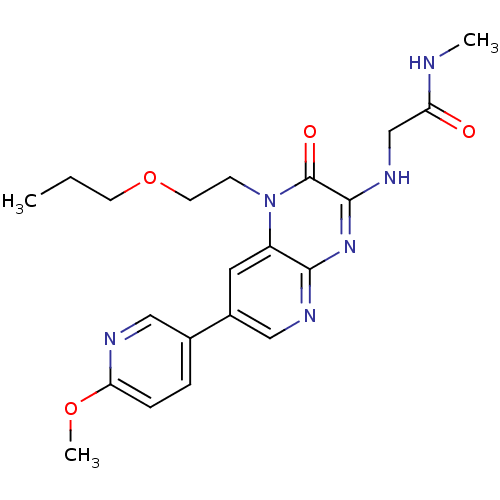
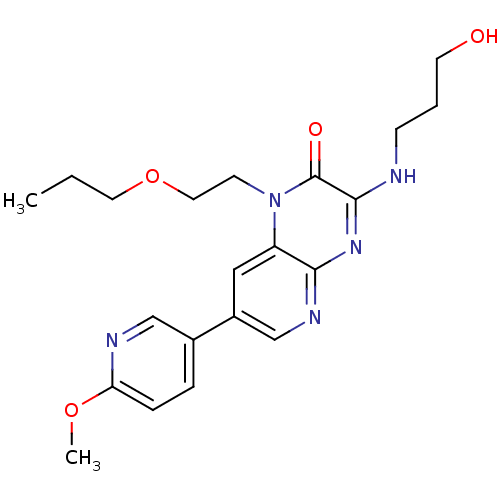
Report error Found 143 Enz. Inhib. hit(s) with Target = 'Rod cGMP-specific 3',5'-cyclic phosphodiesterase subunit alpha'
Affinity DataIC50: 8.89nMpH: 7.5Assay Description:Method: The test compound or solvent was incubated at 25° C. for 15 min with enzyme solution (0.2 μg/mL) in Tris-HCl buffer of pH 7.5. 100 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:A series of dilutions of the test compounds were prepared with 10% DMSO in assay buffer and 5 μl of the dilution was added to a 50 μl react...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Materials and Methods: PDE5 inhibition was assayed at BPS Bioscience (San Diego, Calif.) using BPS PDE assay kits (BPS Catalog number 60300, enzyme l...More data for this Ligand-Target Pair
Affinity DataIC50: 30.1nMAssay Description:Materials and Methods: PDE5 inhibition was assayed at BPS Bioscience (San Diego, Calif.) using BPS PDE assay kits (BPS Catalog number 60300, enzyme l...More data for this Ligand-Target Pair
Affinity DataIC50: 30.1nMAssay Description:A series of dilutions of the test compounds were prepared with 10% DMSO in assay buffer and 5 μl of the dilution was added to a 50 μl react...More data for this Ligand-Target Pair
Affinity DataIC50: 58nMAssay Description:Inhibition of PDE6A (484 to 817 residues) (unknown origin) expressed in Escherichia coli BL21(DE3) using [3H]cGMP or [3H]cAMP as substrate after 15 m...More data for this Ligand-Target Pair
Affinity DataIC50: 58nMpH: 7.5Assay Description:Method: The test compound or solvent was incubated at 25° C. for 15 min with enzyme solution (0.2 μg/mL) in Tris-HCl buffer of pH 7.5. 100 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 61nMpH: 7.5Assay Description:Method: The test compound or solvent was incubated at 25° C. for 15 min with enzyme solution (0.2 μg/mL) in Tris-HCl buffer of pH 7.5. 100 μ...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Materials and Methods: PDE5 inhibition was assayed at BPS Bioscience (San Diego, Calif.) using BPS PDE assay kits (BPS Catalog number 60300, enzyme l...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:A series of dilutions of the test compounds were prepared with 10% DMSO in assay buffer and 5 μl of the dilution was added to a 50 μl react...More data for this Ligand-Target Pair
Affinity DataIC50: 140nMAssay Description:Inhibition of bovine retina PDE6More data for this Ligand-Target Pair