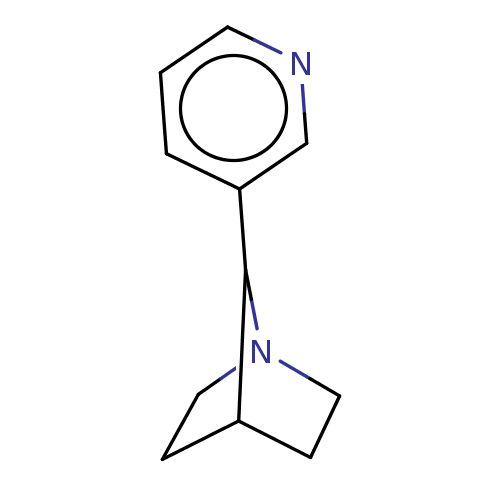
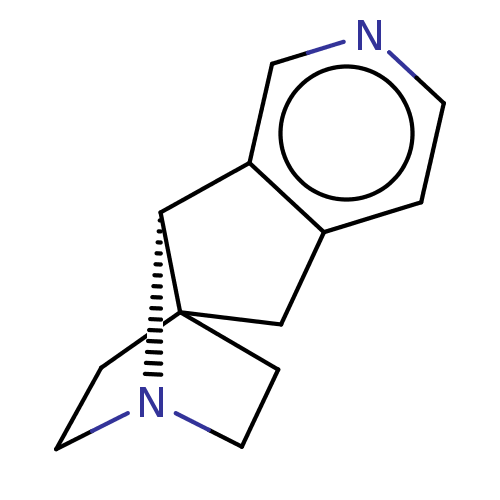
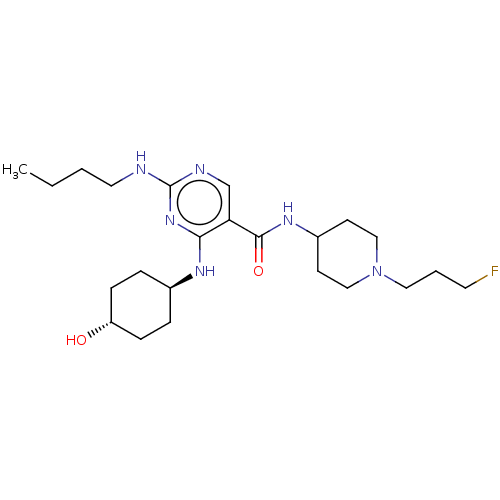
TargetNeuronal acetylcholine receptor subunit alpha-4/beta-2(Rattus norvegicus (Rat))
Vienna University of Technology
Curated by ChEMBL
Vienna University of Technology
Curated by ChEMBL
Affinity DataKi: 0.940nMAssay Description:Displacement of [3H]cytisine from Nicotinic acetylcholine receptor alpha4-beta2 in rat forebrainMore data for this Ligand-Target Pair
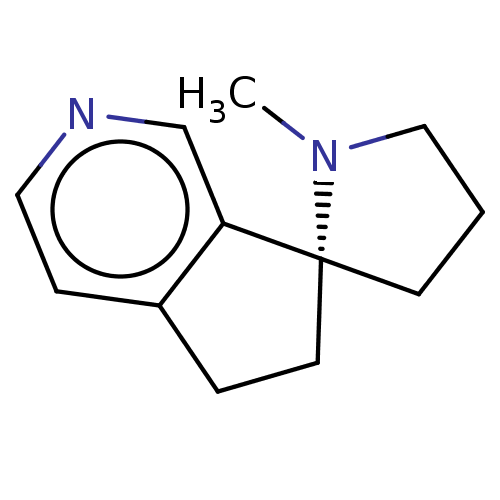
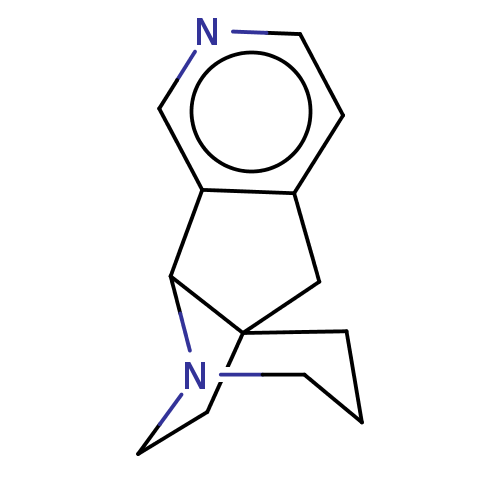
TargetNeuronal acetylcholine receptor subunit alpha-4/beta-2(Rattus norvegicus (Rat))
Vienna University of Technology
Curated by ChEMBL
Vienna University of Technology
Curated by ChEMBL
Affinity DataKi: 1.89nMAssay Description:Displacement of [3H]cytisine from Nicotinic acetylcholine receptor alpha4-beta2 in rat forebrainMore data for this Ligand-Target Pair
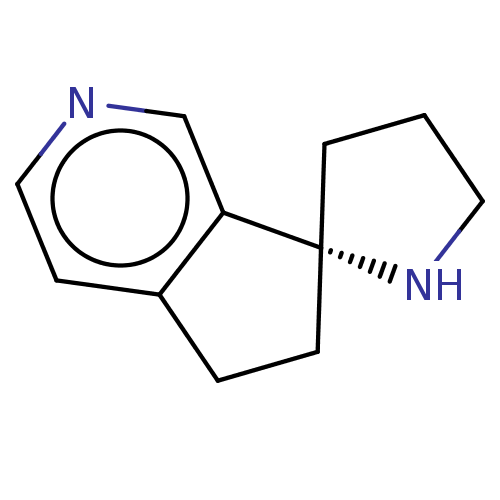
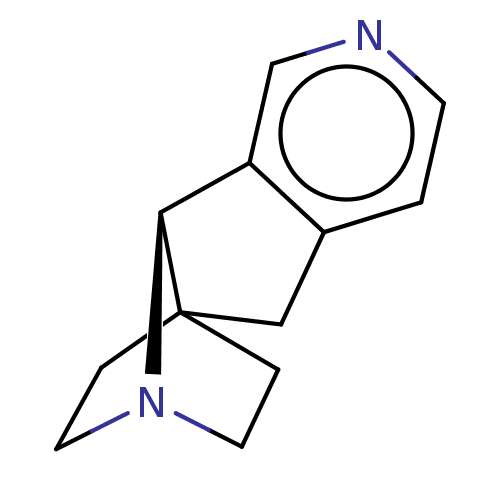
TargetNeuronal acetylcholine receptor subunit alpha-4/beta-2(Rattus norvegicus (Rat))
Vienna University of Technology
Curated by ChEMBL
Vienna University of Technology
Curated by ChEMBL
Affinity DataKi: 1.90nMAssay Description:Displacement of [3H]cytisine from Nicotinic acetylcholine receptor alpha4-beta2 in rat forebrainMore data for this Ligand-Target Pair
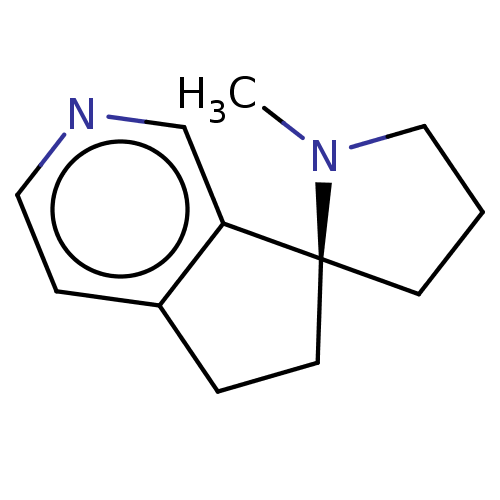
TargetNeuronal acetylcholine receptor subunit alpha-4/beta-2(Rattus norvegicus (Rat))
Vienna University of Technology
Curated by ChEMBL
Vienna University of Technology
Curated by ChEMBL
Affinity DataKi: 4.80nMAssay Description:Displacement of [3H]cytisine from Nicotinic acetylcholine receptor alpha4-beta2 in rat forebrainMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-4/beta-2(Rattus norvegicus (Rat))
Vienna University of Technology
Curated by ChEMBL
Vienna University of Technology
Curated by ChEMBL
Affinity DataKi: 53nMAssay Description:Displacement of [3H]cytisine from Nicotinic acetylcholine receptor alpha4-beta2 in rat forebrainMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-4/beta-2(Rattus norvegicus (Rat))
Vienna University of Technology
Curated by ChEMBL
Vienna University of Technology
Curated by ChEMBL
Affinity DataKi: 148nMAssay Description:Displacement of [3H]cytisine from Nicotinic acetylcholine receptor alpha4-beta2 in rat forebrainMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-4/beta-2(Rattus norvegicus (Rat))
Vienna University of Technology
Curated by ChEMBL
Vienna University of Technology
Curated by ChEMBL
Affinity DataKi: 533nMAssay Description:Displacement of [3H]cytisine from Nicotinic acetylcholine receptor alpha4-beta2 in rat forebrainMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-4/beta-2(Rattus norvegicus (Rat))
Vienna University of Technology
Curated by ChEMBL
Vienna University of Technology
Curated by ChEMBL
Affinity DataKi: 962nMAssay Description:Displacement of [3H]cytisine from Nicotinic acetylcholine receptor alpha4-beta2 in rat forebrainMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-4/beta-2(Rattus norvegicus (Rat))
Vienna University of Technology
Curated by ChEMBL
Vienna University of Technology
Curated by ChEMBL
Affinity DataKi: 1.04E+3nMAssay Description:Displacement of [3H]cytisine from Nicotinic acetylcholine receptor alpha4-beta2 in rat forebrainMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-4/beta-2(Rattus norvegicus (Rat))
Vienna University of Technology
Curated by ChEMBL
Vienna University of Technology
Curated by ChEMBL
Affinity DataKi: 1.04E+3nMAssay Description:Displacement of [3H]cytisine from Nicotinic acetylcholine receptor alpha4-beta2 in rat forebrainMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-4/beta-2(Rattus norvegicus (Rat))
Vienna University of Technology
Curated by ChEMBL
Vienna University of Technology
Curated by ChEMBL
Affinity DataKi: 1.36E+3nMAssay Description:Displacement of [3H]cytisine from Nicotinic acetylcholine receptor alpha4-beta2 in rat forebrainMore data for this Ligand-Target Pair
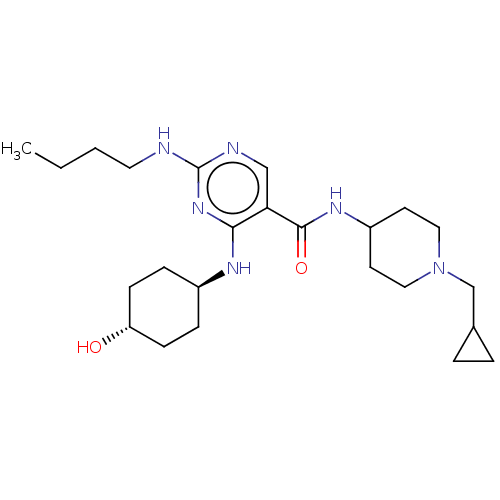
Affinity DataIC50: 3.60nMAssay Description:Inhibition of MERTK (unknown origin) using 5-FAM-EFPIYDFLPAKKK-CONH2 as substrate incubated for 180 mins in presence of ATP by microfluidic capillary...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of MERTK (unknown origin) using 5-FAM-EFPIYDFLPAKKK-CONH2 as substrate incubated for 180 mins in presence of ATP by microfluidic capillary...More data for this Ligand-Target Pair
Affinity DataIC50: 4.60nMAssay Description:Inhibition of MERTK (unknown origin) using 5-FAM-EFPIYDFLPAKKK-CONH2 as substrate incubated for 180 mins in presence of ATP by microfluidic capillary...More data for this Ligand-Target Pair
TargetReceptor-type tyrosine-protein kinase FLT3(Homo sapiens (Human))
Monash University
Curated by ChEMBL
Monash University
Curated by ChEMBL
Affinity DataIC50: 49nMAssay Description:Inhibition of FLT3 (unknown origin) using 5-FAM-KKKKEEIYFFF-CONH2 as substrate incubated for 180 mins in presence of ATP by microfluidic capillary el...More data for this Ligand-Target Pair
TargetReceptor-type tyrosine-protein kinase FLT3(Homo sapiens (Human))
Monash University
Curated by ChEMBL
Monash University
Curated by ChEMBL
Affinity DataIC50: 110nMAssay Description:Inhibition of FLT3 (unknown origin) using 5-FAM-KKKKEEIYFFF-CONH2 as substrate incubated for 180 mins in presence of ATP by microfluidic capillary el...More data for this Ligand-Target Pair
TargetReceptor-type tyrosine-protein kinase FLT3(Homo sapiens (Human))
Monash University
Curated by ChEMBL
Monash University
Curated by ChEMBL
Affinity DataIC50: 120nMAssay Description:Inhibition of FLT3 (unknown origin) using 5-FAM-KKKKEEIYFFF-CONH2 as substrate incubated for 180 mins in presence of ATP by microfluidic capillary el...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase receptor TYRO3(Homo sapiens (Human))
Monash University
Curated by ChEMBL
Monash University
Curated by ChEMBL
Affinity DataIC50: 150nMAssay Description:Inhibition of Tyro3 (unknown origin) using 5-FAM-EFPIYDFLPAKKK-CONH2 as substrate incubated for 180 mins in presence of ATP by microfluidic capillary...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Inhibition of Axl (unknown origin) using 5-FAM-KKKKEEIYFFF-CONH2 as substrate incubated for 180 mins in presence of ATP by microfluidic capillary ele...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:Inhibition of Axl (unknown origin) using 5-FAM-KKKKEEIYFFF-CONH2 as substrate incubated for 180 mins in presence of ATP by microfluidic capillary ele...More data for this Ligand-Target Pair
Affinity DataIC50: 218nMAssay Description:Inhibition of Axl (unknown origin) using 5-FAM-KKKKEEIYFFF-CONH2 as substrate incubated for 180 mins in presence of ATP by microfluidic capillary ele...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase receptor TYRO3(Homo sapiens (Human))
Monash University
Curated by ChEMBL
Monash University
Curated by ChEMBL
Affinity DataIC50: 390nMAssay Description:Inhibition of Tyro3 (unknown origin) using 5-FAM-EFPIYDFLPAKKK-CONH2 as substrate incubated for 180 mins in presence of ATP by microfluidic capillary...More data for this Ligand-Target Pair
TargetTyrosine-protein kinase receptor TYRO3(Homo sapiens (Human))
Monash University
Curated by ChEMBL
Monash University
Curated by ChEMBL
Affinity DataIC50: 400nMAssay Description:Inhibition of Tyro3 (unknown origin) using 5-FAM-EFPIYDFLPAKKK-CONH2 as substrate incubated for 180 mins in presence of ATP by microfluidic capillary...More data for this Ligand-Target Pair
Affinity DataKd: 13nMAssay Description:Binding affinity to MERTK in cuprizone-challenged C57BL/6 mouse whole brain assessed as dissociation constant at 18.1 nM by autoradiographyMore data for this Ligand-Target Pair
Affinity DataKd: 20nMAssay Description:Binding affinity to MERTK in cuprizone-challenged C57BL/6 mouse brain CC/HC region assessed as dissociation constant at 18.1 nM by autoradiographyMore data for this Ligand-Target Pair

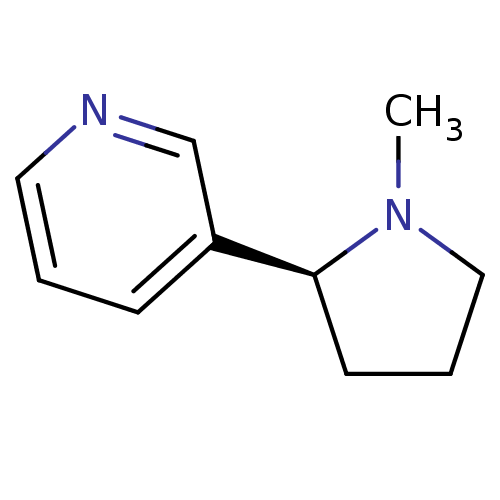
 3D Structure (crystal)
3D Structure (crystal)