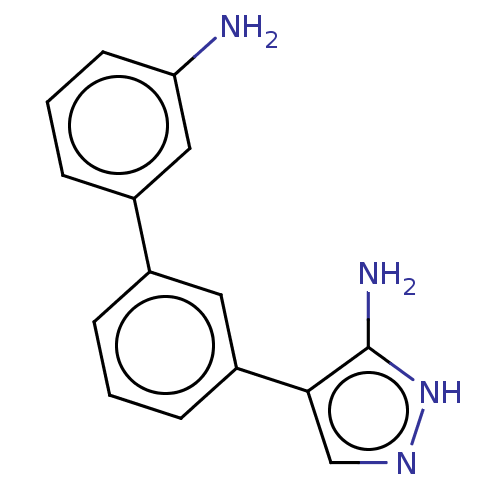
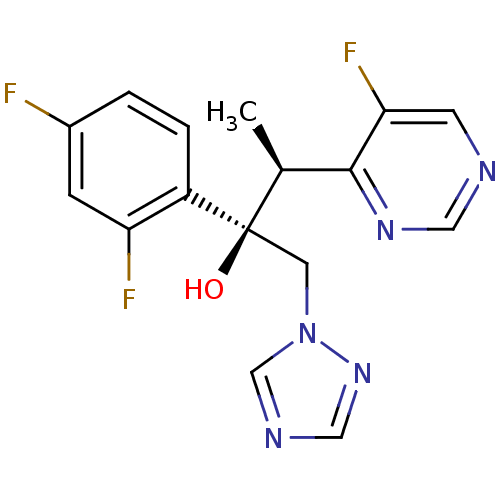
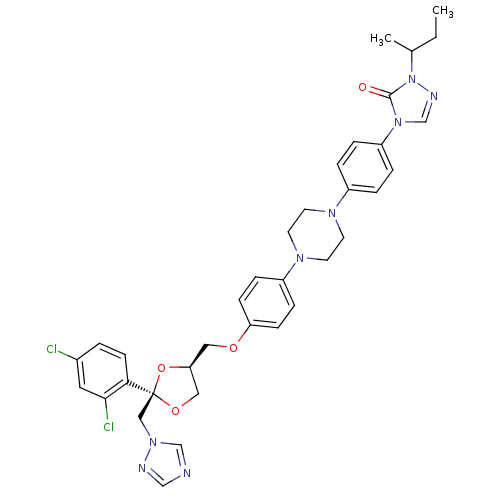
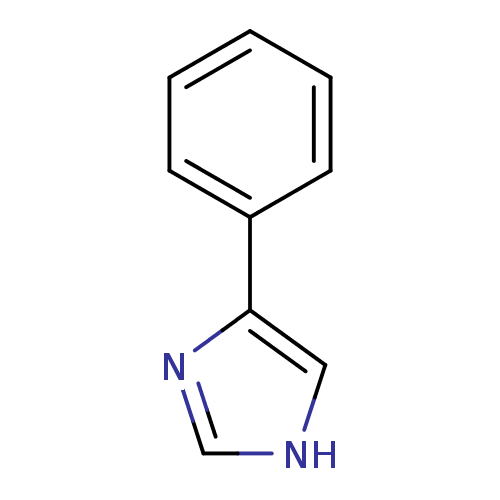
TargetCytochrome P450 3A5(Homo sapiens (Human))
St. Jude Children'S Research Hospital
Curated by ChEMBL
St. Jude Children'S Research Hospital
Curated by ChEMBL
Affinity DataIC50: 21nMAssay Description:Inhibition of CYP3A5 in doxycycline-induced CYP3A5 overexpressing wild type human AsPC1 cells assessed as decrease in 1-hydroxymidazolam formation us...More data for this Ligand-Target Pair
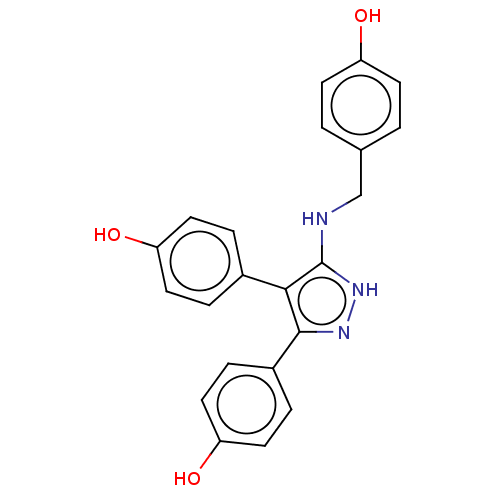
TargetCytochrome P450 3A5(Homo sapiens (Human))
St. Jude Children'S Research Hospital
Curated by ChEMBL
St. Jude Children'S Research Hospital
Curated by ChEMBL
Affinity DataIC50: 44nMAssay Description:Inhibition of CYP3A5 in wild type human AsPC1 cells assessed as decrease in 1-hydroxymidazolam formation using midazolam as substrate after 24 hrs by...More data for this Ligand-Target Pair
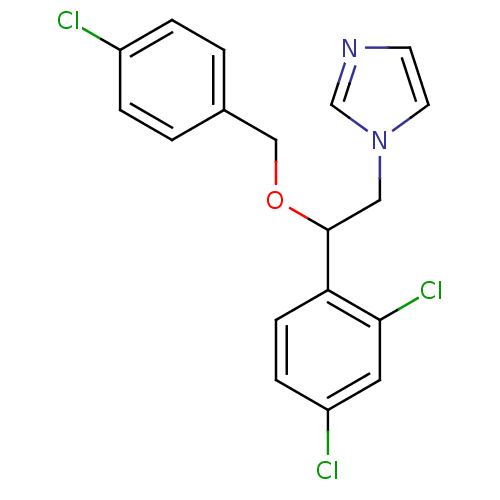
TargetCytochrome P450 3A5(Homo sapiens (Human))
St. Jude Children'S Research Hospital
Curated by ChEMBL
St. Jude Children'S Research Hospital
Curated by ChEMBL
Affinity DataIC50: 78nMAssay Description:Inhibition of CYP3A5 in lentiviral pLVX-TRE3G-ZsGreen1-CYP3A5 transduced wild type human AsPC1 cells overexpressing CYP3A5 assessed as decrease in 1-...More data for this Ligand-Target Pair
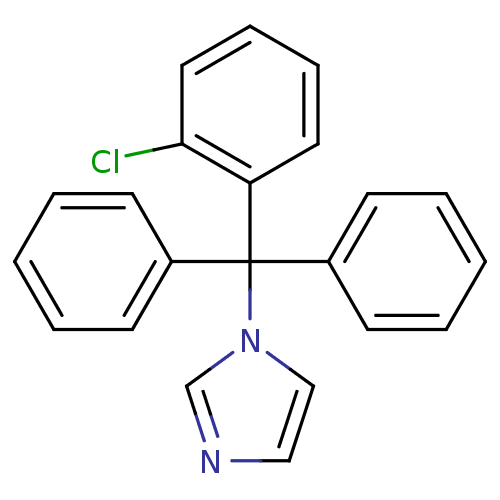
TargetCytochrome P450 3A5(Homo sapiens (Human))
St. Jude Children'S Research Hospital
Curated by ChEMBL
St. Jude Children'S Research Hospital
Curated by ChEMBL
Affinity DataIC50: 103nMAssay Description:Inhibition of CYP3A5 in CRISPR/Cas9-mediated CYP3A5 knock-out and doxycycline-induced CYP3A5 overexpressing human AsPC1 cells assessed as decrease in...More data for this Ligand-Target Pair
TargetCytochrome P450 3A4(Homo sapiens (Human))
St. Jude Children'S Research Hospital
Curated by ChEMBL
St. Jude Children'S Research Hospital
Curated by ChEMBL
Affinity DataIC50: 117nMAssay Description:Inhibition of CYP3A4 in CRISPR/Cas9-mediated CYP3A5 knock-out and doxycycline-induced CYP3A4 overexpressing human AsPC1 cells assessed as decrease in...More data for this Ligand-Target Pair
TargetCytochrome P450 3A5(Homo sapiens (Human))
St. Jude Children'S Research Hospital
Curated by ChEMBL
St. Jude Children'S Research Hospital
Curated by ChEMBL
Affinity DataIC50: 141nMAssay Description:Inhibition of CYP3A5 in doxycycline-induced CYP3A5 overexpressing wild type human AsPC1 cells assessed as decrease in 1-hydroxymidazolam formation us...More data for this Ligand-Target Pair
TargetCytochrome P450 3A5(Homo sapiens (Human))
St. Jude Children'S Research Hospital
Curated by ChEMBL
St. Jude Children'S Research Hospital
Curated by ChEMBL
Affinity DataIC50: 162nMAssay Description:Inhibition of CYP3A5 in CRISPR/Cas9-mediated CYP3A5 knock-out and doxycycline-induced CYP3A5 overexpressing human AsPC1 cells assessed as decrease in...More data for this Ligand-Target Pair
TargetCytochrome P450 3A4(Homo sapiens (Human))
St. Jude Children'S Research Hospital
Curated by ChEMBL
St. Jude Children'S Research Hospital
Curated by ChEMBL
Affinity DataIC50: 206nMAssay Description:Inhibition of human recombinant CYP3A4 expressed in supersomes assessed as decrease in formation of D-luciferin using luciferin-IPA as substrate incu...More data for this Ligand-Target Pair
TargetCytochrome P450 3A5(Homo sapiens (Human))
St. Jude Children'S Research Hospital
Curated by ChEMBL
St. Jude Children'S Research Hospital
Curated by ChEMBL
Affinity DataIC50: 439nMAssay Description:Inhibition of CYP3A5 in lentiviral pLVX-TRE3G-ZsGreen1-CYP3A5 transduced wild type human AsPC1 cells overexpressing CYP3A5 assessed as decrease in 1-...More data for this Ligand-Target Pair
TargetCytochrome P450 3A5(Homo sapiens (Human))
St. Jude Children'S Research Hospital
Curated by ChEMBL
St. Jude Children'S Research Hospital
Curated by ChEMBL
Affinity DataIC50: 513nMAssay Description:Inhibition of CYP3A5 in wild type human AsPC1 cells assessed as decrease in 1-hydroxymidazolam formation using midazolam as substrate after 24 hrs by...More data for this Ligand-Target Pair
TargetCytochrome P450 3A4(Homo sapiens (Human))
St. Jude Children'S Research Hospital
Curated by ChEMBL
St. Jude Children'S Research Hospital
Curated by ChEMBL
Affinity DataIC50: 1.13E+4nMAssay Description:Inhibition of CYP3A4 in human liver microsomes using midazolam as substrate by LC-MS/MS analysisMore data for this Ligand-Target Pair
TargetCytochrome P450 3A5(Homo sapiens (Human))
St. Jude Children'S Research Hospital
Curated by ChEMBL
St. Jude Children'S Research Hospital
Curated by ChEMBL
Affinity DataIC50: 1.56E+4nMAssay Description:Inhibition of human recombinant CYP3A5 expressed in supersomes assessed as decrease in formation of D-luciferin using luciferin-IPA as substrate incu...More data for this Ligand-Target Pair
Affinity DataIC50: >2.50E+4nMAssay Description:Inhibition of CYP2C9 in human liver microsomes using tolbutamide as substrate by LC-MS/MS analysisMore data for this Ligand-Target Pair
Affinity DataIC50: >2.50E+4nMAssay Description:Inhibition of CYP2C9 in human liver microsomes using tolbutamide as substrate by LC-MS/MS analysisMore data for this Ligand-Target Pair
Affinity DataIC50: >2.50E+4nMAssay Description:Inhibition of CYP2C9 in human liver microsomes using tolbutamide as substrate by LC-MS/MS analysisMore data for this Ligand-Target Pair
Affinity DataIC50: >2.50E+4nMAssay Description:Inhibition of CYP2C19 in human liver microsomes using mephenytoin as substrate by LC-MS/MS analysisMore data for this Ligand-Target Pair
Affinity DataIC50: >2.50E+4nMAssay Description:Inhibition of CYP2C19 in human liver microsomes using mephenytoin as substrate by LC-MS/MS analysisMore data for this Ligand-Target Pair
Affinity DataIC50: >2.50E+4nMAssay Description:Inhibition of CYP2C19 in human liver microsomes using mephenytoin as substrate by LC-MS/MS analysisMore data for this Ligand-Target Pair
Affinity DataIC50: >2.50E+4nMAssay Description:Inhibition of CYP2D6 in human liver microsomes using dextromethorphan as substrate by LC-MS/MS analysisMore data for this Ligand-Target Pair
Affinity DataIC50: >2.50E+4nMAssay Description:Inhibition of CYP2D6 in human liver microsomes using dextromethorphan as substrate by LC-MS/MS analysisMore data for this Ligand-Target Pair
TargetCytochrome P450 3A4(Homo sapiens (Human))
St. Jude Children'S Research Hospital
Curated by ChEMBL
St. Jude Children'S Research Hospital
Curated by ChEMBL
Affinity DataIC50: >2.50E+4nMAssay Description:Inhibition of CYP3A4 in human liver microsomes using midazolam as substrate by LC-MS/MS analysisMore data for this Ligand-Target Pair
Affinity DataIC50: >2.50E+4nMAssay Description:Inhibition of CYP2D6 in human liver microsomes using dextromethorphan as substrate by LC-MS/MS analysisMore data for this Ligand-Target Pair
TargetCytochrome P450 3A4(Homo sapiens (Human))
St. Jude Children'S Research Hospital
Curated by ChEMBL
St. Jude Children'S Research Hospital
Curated by ChEMBL
Affinity DataIC50: >2.50E+4nMAssay Description:Inhibition of CYP3A4 in human liver microsomes using midazolam as substrate by LC-MS/MS analysisMore data for this Ligand-Target Pair
Affinity DataKd: 780nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
TargetCytochrome P450 3A4(Homo sapiens (Human))
St. Jude Children'S Research Hospital
Curated by ChEMBL
St. Jude Children'S Research Hospital
Curated by ChEMBL
Affinity DataKd: 900nMAssay Description:Binding affinity to His-tagged CYP3A4 (unknown origin) expressed in Escherichia coli DH5alpha assessed as type 2 spectral shift by measuring dissocia...More data for this Ligand-Target Pair
Affinity DataKd: 4.60E+3nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 370nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 5.30E+3nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 3.80E+3nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 980nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 4.60E+3nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 4.00E+3nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 1.34E+5nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 2.71E+4nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 2.10E+4nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 4.31E+4nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 8.60E+5nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 1.74E+5nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 3.02E+4nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 2.80E+5nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 2.16E+5nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 1.20E+4nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 1.20E+3nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 360nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 110nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
Affinity DataKd: 340nMAssay Description:Substrate and ligand binding assay using UV- visible absorbance analysis of CYP142 was done on a Cary UV-50 UV-visible scanning spectrophotometer (Va...More data for this Ligand-Target Pair
TargetBifunctional cytochrome P450/NADPH--P450 reductase(Bacillus megaterium)
University of Manchester
University of Manchester
Affinity DataKd: 82nMpH: 7.0Assay Description:Dissociation constants (Kd values) for binding of the substrates N-palmitoylglycine (NPG) and omeprazole (OMP) to WT and mutant BM3 heme domains were...More data for this Ligand-Target Pair
TargetBifunctional cytochrome P450/NADPH--P450 reductase [A82F](Bacillus megaterium)
University of Manchester
University of Manchester
Affinity DataKd: 297nMpH: 7.0Assay Description:Dissociation constants (Kd values) for binding of the substrates N-palmitoylglycine (NPG) and omeprazole (OMP) to WT and mutant BM3 heme domains were...More data for this Ligand-Target Pair
TargetBifunctional cytochrome P450/NADPH--P450 reductase [F87V](Bacillus megaterium)
University of Manchester
University of Manchester
Affinity DataKd: 204nMpH: 7.0Assay Description:Dissociation constants (Kd values) for binding of the substrates N-palmitoylglycine (NPG) and omeprazole (OMP) to WT and mutant BM3 heme domains were...More data for this Ligand-Target Pair
TargetBifunctional cytochrome P450/NADPH--P450 reductase [A82F,F87V](Bacillus megaterium)
University of Manchester
University of Manchester
Affinity DataKd: 4nMpH: 7.0Assay Description:Dissociation constants (Kd values) for binding of the substrates N-palmitoylglycine (NPG) and omeprazole (OMP) to WT and mutant BM3 heme domains were...More data for this Ligand-Target Pair

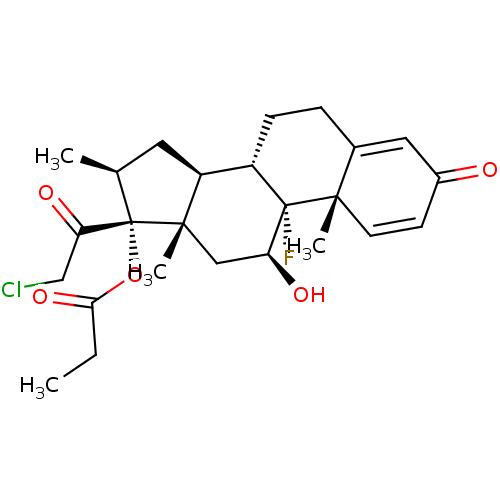
 3D Structure (crystal)
3D Structure (crystal)