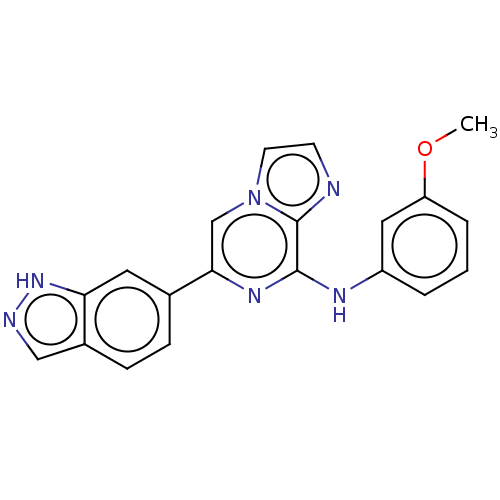
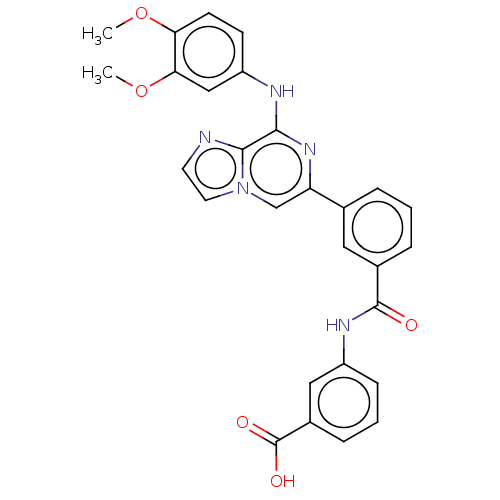
Affinity DataKi: 0.190nMAssay Description:Compound was evaluated for binding affinity towards dopamine receptor D2 in striatal membranes, using [3H]- spiperone as radioligand in the presence ...More data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Compound was evaluated for binding affinity towards dopamine receptor D2 in striatal membranes, using [3H]- spiperone as radioligand in the presence ...More data for this Ligand-Target Pair
Affinity DataKi: 0.25nMAssay Description:Compound was evaluated for binding affinity towards dopamine receptor D2 in striatal membranes, using [3H]- spiperone as radioligand in the presence ...More data for this Ligand-Target Pair
Affinity DataKi: 0.270nMAssay Description:Compound was evaluated for binding affinity towards Dopamine receptor D2 in striatal membranes, using [3H]- spiperone as radioligand in the absence o...More data for this Ligand-Target Pair
Affinity DataKi: 0.660nMAssay Description:Displacement of [3H]- spiperone from dopamine receptor D2 of striatal membranes without sodium chlorideMore data for this Ligand-Target Pair
Affinity DataKi: 0.760nMAssay Description:Displacement of [3H]- spiperone from dopamine receptor D2 of striatal membranes without sodium chlorideMore data for this Ligand-Target Pair
Affinity DataKi: 0.770nMAssay Description:Compound was evaluated for binding affinity towards dopamine receptor D2 in striatal membranes, using [3H]- spiperone as radioligand in the presence ...More data for this Ligand-Target Pair
Affinity DataKi: 0.770nMAssay Description:Displacement of [3H]- spiperone from dopamine receptor D2 of striatal membranes without sodium chlorideMore data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:Displacement of [3H]- spiperone from dopamine receptor D2 of striatal membranes without sodium chlorideMore data for this Ligand-Target Pair
Affinity DataKi: 0.990nMAssay Description:Displacement of [3H]- spiperone from dopamine receptor D2 of striatal membranes without sodium chlorideMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Compound was evaluated for binding affinity towards dopamine receptor D2 in striatal membranes, using [3H]- spiperone as radioligand in the presence ...More data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Displacement of [3H]- spiperone from dopamine receptor D2 of striatal membranes without sodium chlorideMore data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:Compound was evaluated for binding affinity towards dopamine receptor D2 in striatal membranes, using [3H]- spiperone as radioligand in the presence ...More data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(RAT)
University of California
Curated by ChEMBL
University of California
Curated by ChEMBL
Affinity DataKi: 1.40nMAssay Description:Compound was evaluated for binding affinity towards dopamine receptor D2 in striatal membranes, using [3H]- spiperone as radioligand in the presence ...More data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Compound was evaluated for binding affinity towards Dopamine receptor D2 in striatal membranes, using [3H]- spiperone as radioligand in the absence o...More data for this Ligand-Target Pair
Affinity DataKi: 2.20nMAssay Description:Compound was evaluated for binding affinity towards dopamine receptor D2 in striatal membranes, using [3H]- spiperone as radioligand in the presence ...More data for this Ligand-Target Pair
Affinity DataKi: 4.40nMAssay Description:Compound was evaluated for binding affinity towards dopamine receptor D2 in striatal membranes, using [3H]- spiperone as radioligand in the presence ...More data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(RAT)
University of California
Curated by ChEMBL
University of California
Curated by ChEMBL
Affinity DataKi: 5nMAssay Description:Compound was evaluated for binding affinity towards dopamine receptor D2 in striatal membranes, using [3H]- spiperone as radioligand in the presence ...More data for this Ligand-Target Pair
Affinity DataKi: 5.30nMAssay Description:Displacement of [3H]- spiperone from dopamine receptor D2 of striatal membranes without sodium chlorideMore data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Displacement of [3H]- spiperone from dopamine receptor D2 of striatal membranes without sodium chlorideMore data for this Ligand-Target Pair
Affinity DataKi: 8.30nMAssay Description:Displacement of [3H]- spiperone from dopamine receptor D2 of striatal membranes without sodium chlorideMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(RAT)
University of California
Curated by ChEMBL
University of California
Curated by ChEMBL
Affinity DataKi: 9.5nMAssay Description:binding affinity towards alpha-2 adrenergic receptor, using [3H]- atipamezole as radioligand from rat frontal cortex membranesMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(RAT)
University of California
Curated by ChEMBL
University of California
Curated by ChEMBL
Affinity DataKi: 11nMAssay Description:Compound was evaluated for binding affinity towards dopamine receptor D2 in striatal membranes, using [3H]- spiperone as radioligand in the presence ...More data for this Ligand-Target Pair
Affinity DataKi: 33nMAssay Description:Displacement of [3H]- spiperone from dopamine receptor D2 of striatal membranes without sodium chlorideMore data for this Ligand-Target Pair
Affinity DataKi: 66nMAssay Description:Displacement of [3H]- spiperone from dopamine receptor D2 of striatal membranes without sodium chlorideMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]- spiperone from dopamine receptor D2 of striatal membranes without sodium chlorideMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]- spiperone from dopamine receptor D2 of striatal membranes without sodium chlorideMore data for this Ligand-Target Pair
Affinity DataIC50: 0.440nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 0.840nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.5nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.90nM Kd: 1.5nMpH: 7.5 T: 2°CAssay Description:Biochemical assay using Lanthascreen (human, full-lenght, C-terminal v5-His6 expressed in Sf9 cell) assay from Invitrogen.More data for this Ligand-Target Pair
Affinity DataIC50: 2.10nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 2.30nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 2.60nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 2.70nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 3.80nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 3.80nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 3.90nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 4.30nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 4.70nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 6.30nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 7.60nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 7.70nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 8nM Kd: 25nMAssay Description:Biochemical binding assays (DiscoveRx) and in vitro enzyme inhibition assays (LanthaScreen, Life Technologies) were conducted to determine the bindin...More data for this Ligand-Target Pair
Affinity DataIC50: 8.5nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 8.60nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 8.90nMAssay Description:Inhibition of full length Syk (unknown origin) using biotinylated peptide substrateMore data for this Ligand-Target Pair

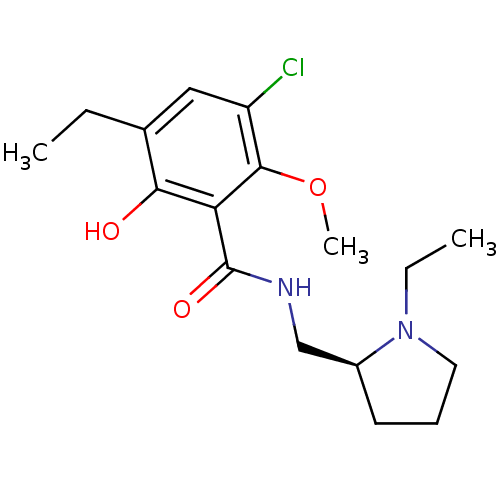
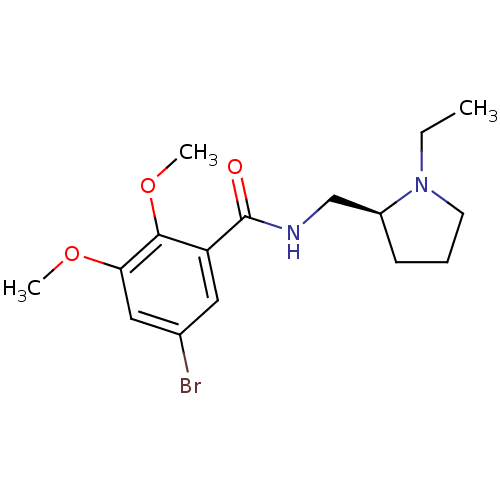

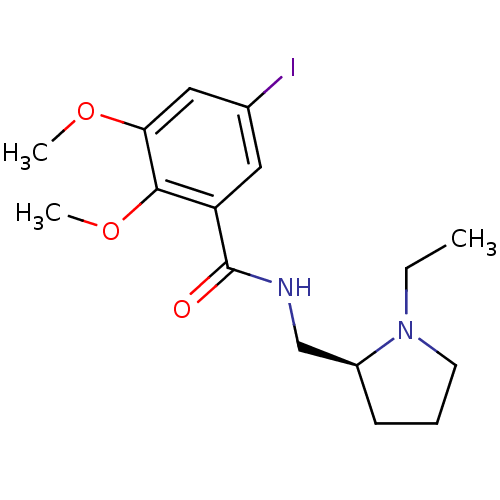
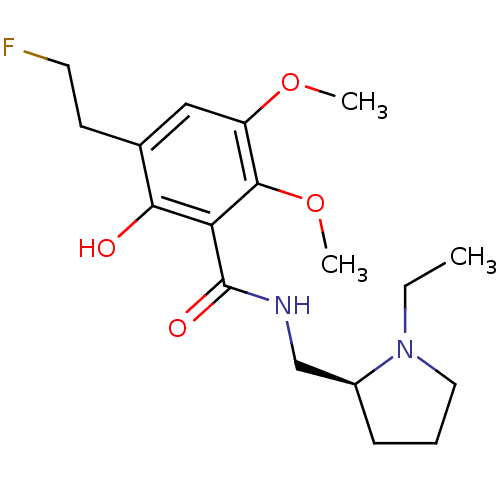
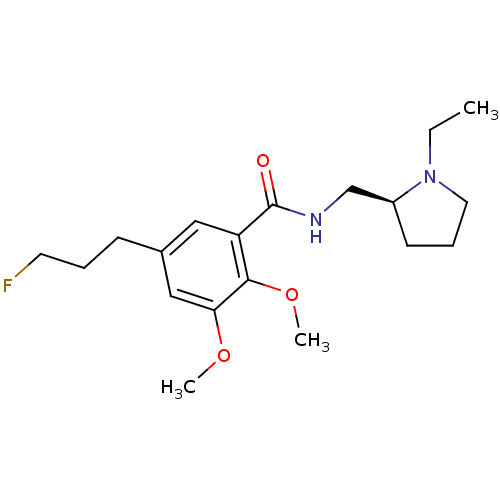
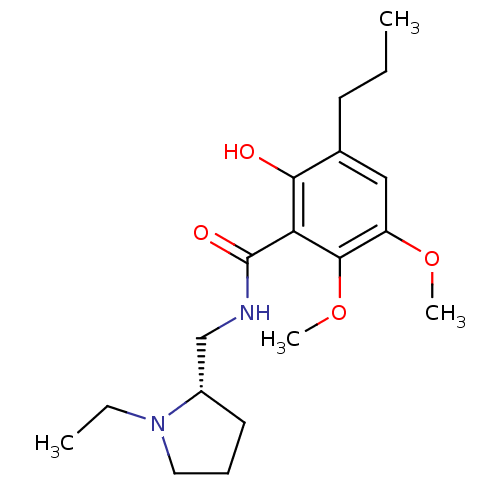

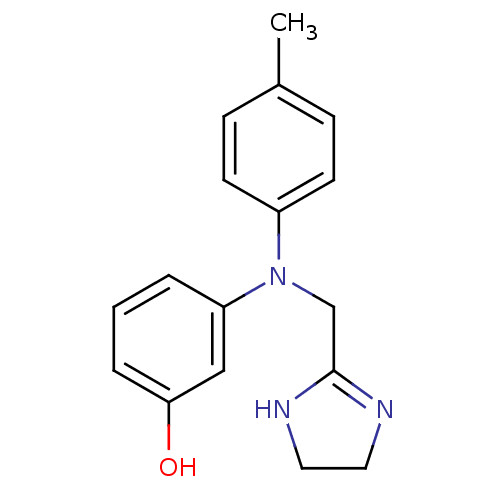
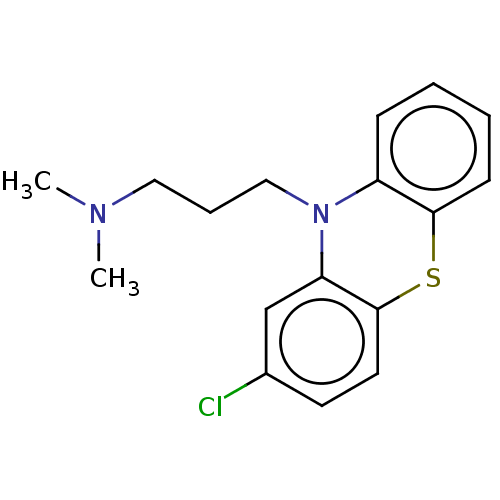
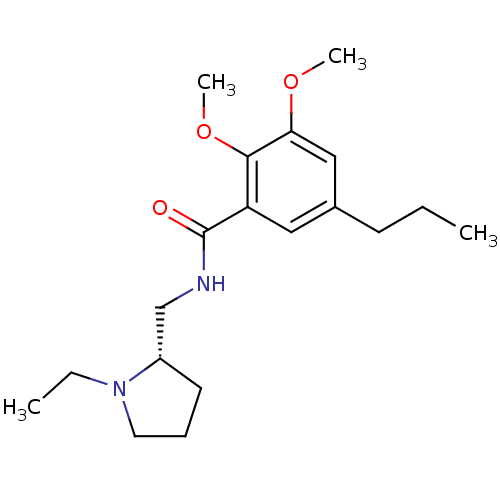
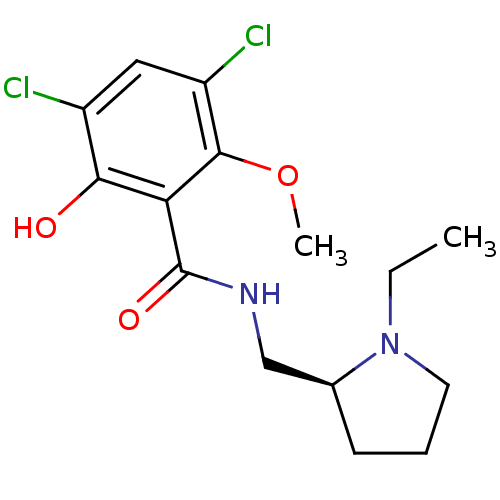
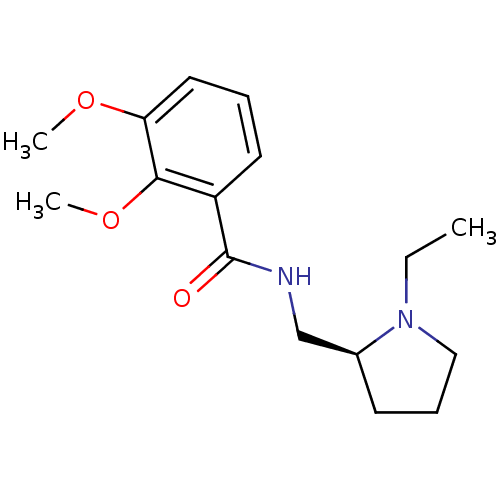

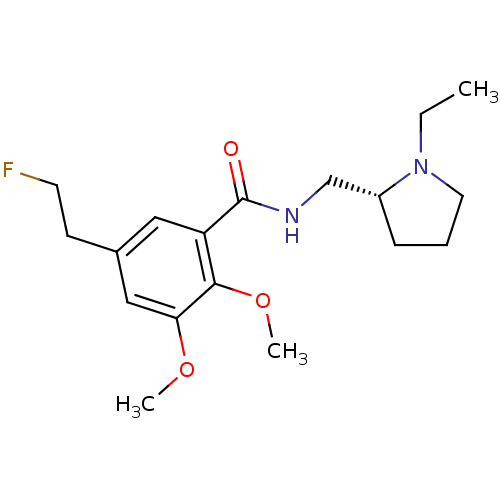
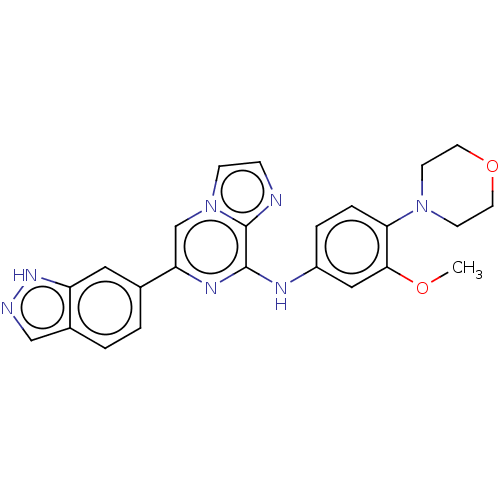
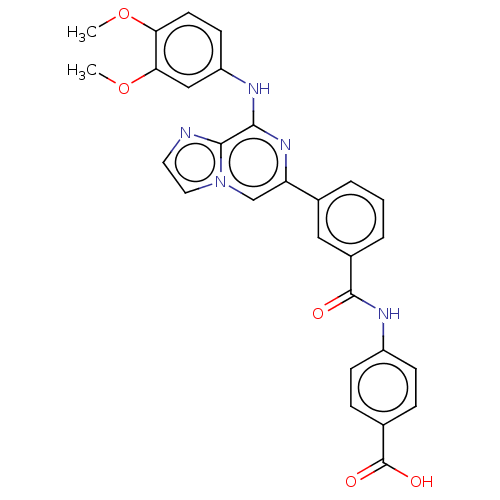
 3D Structure (crystal)
3D Structure (crystal)