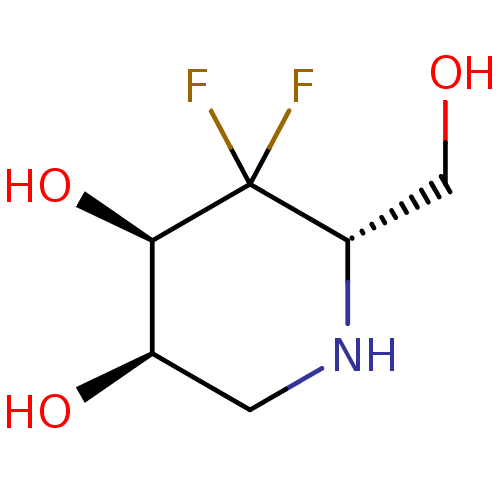
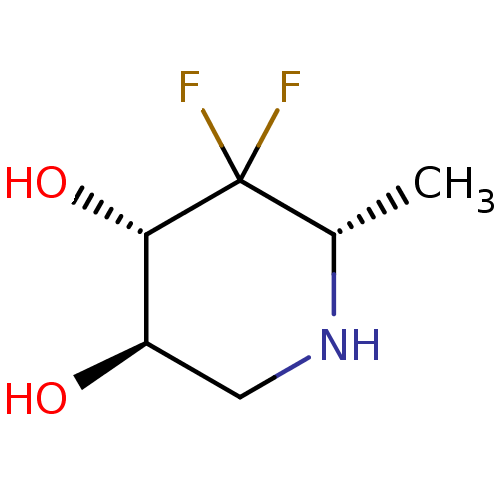
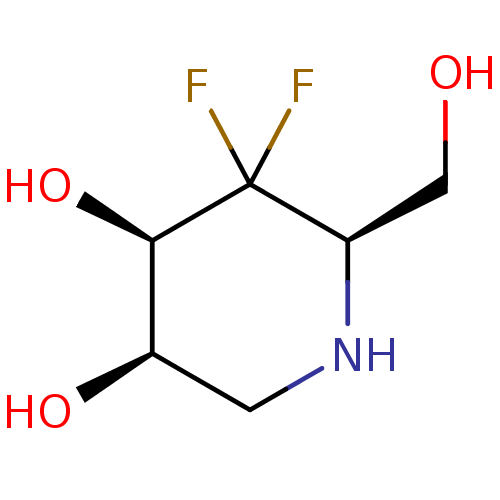
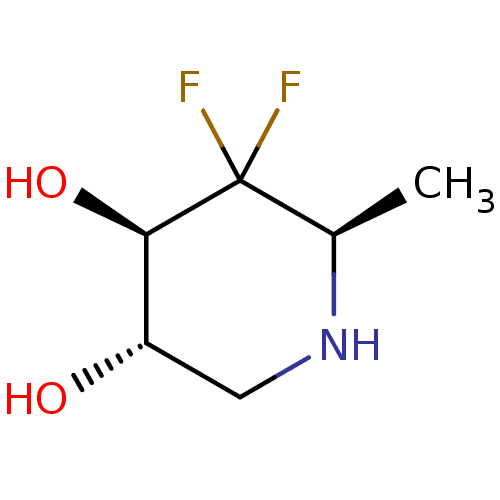
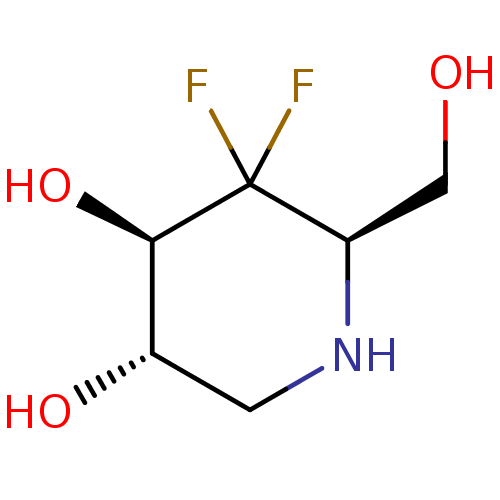
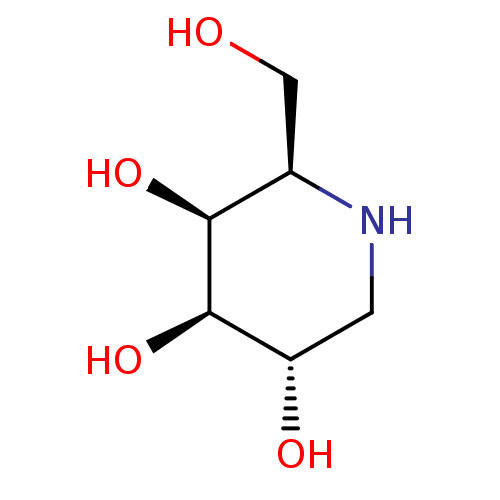
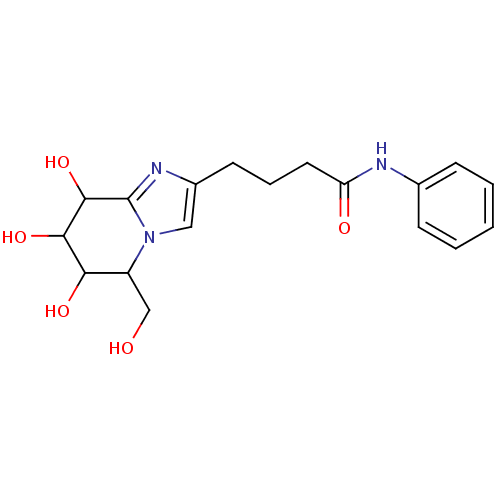
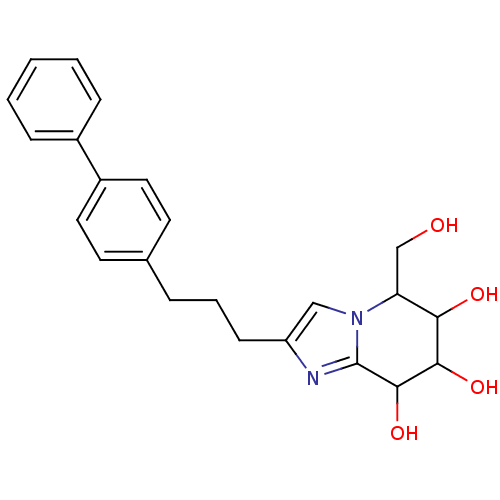
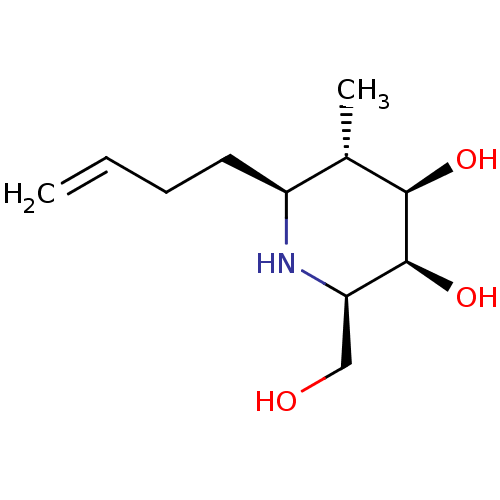
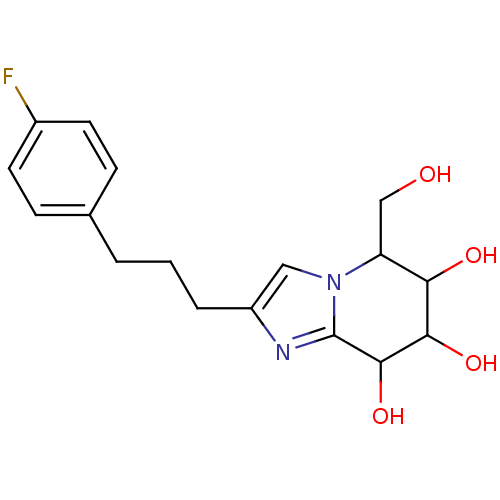
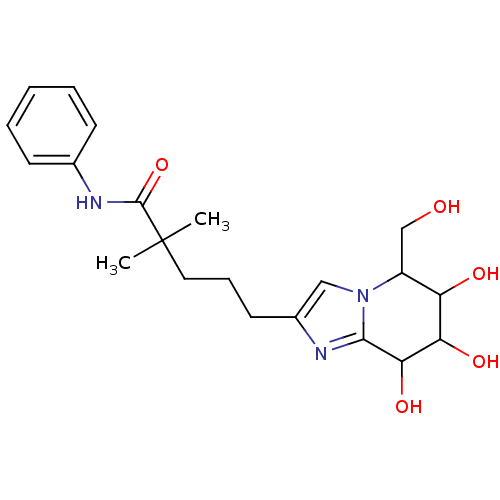
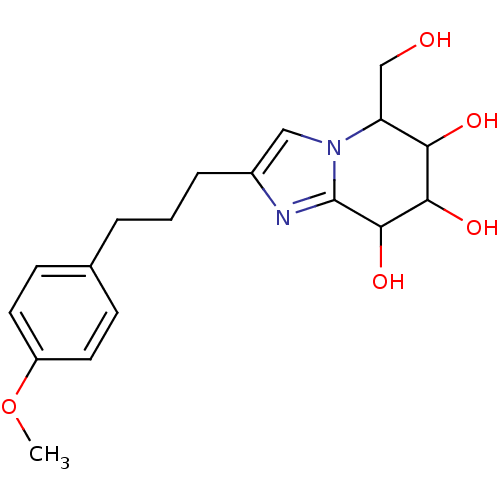
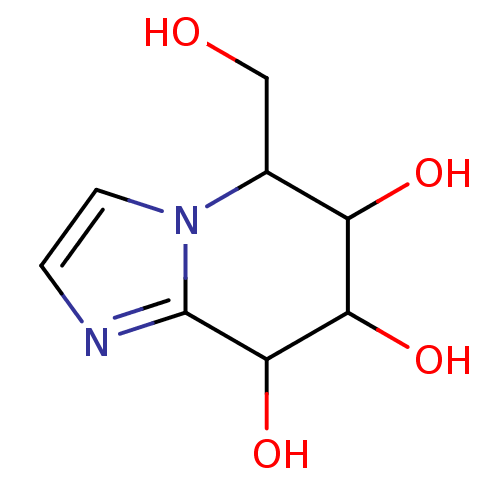
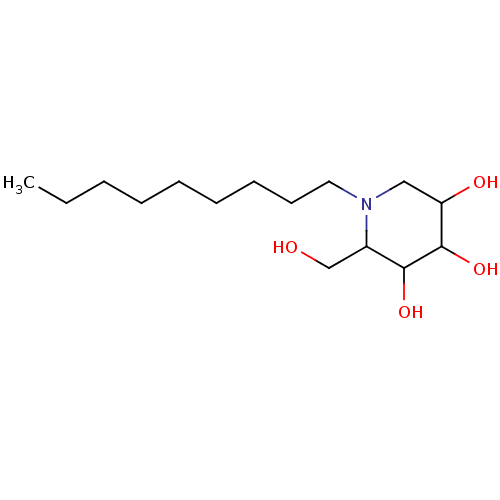
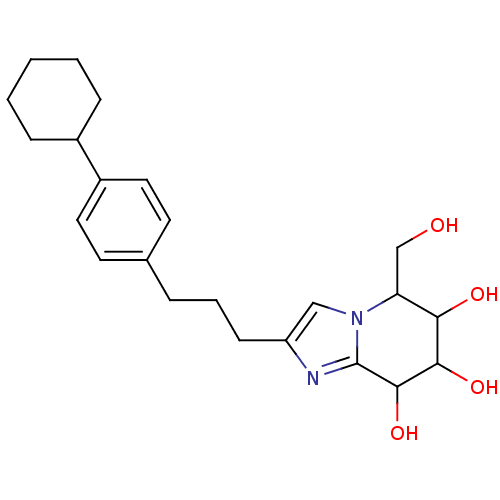
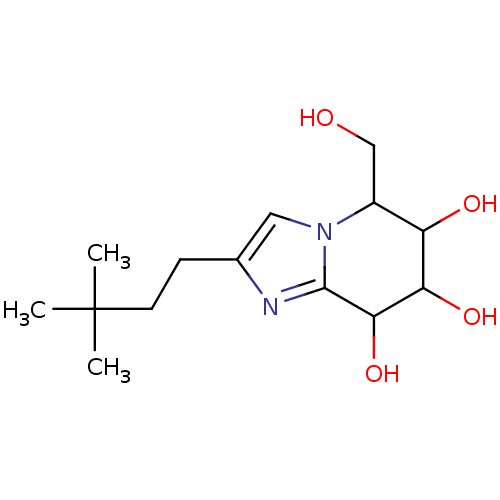
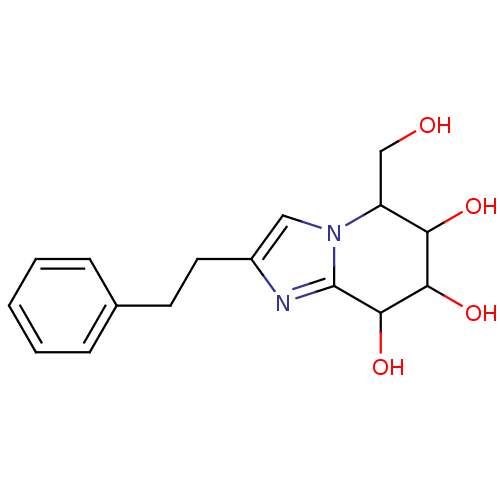
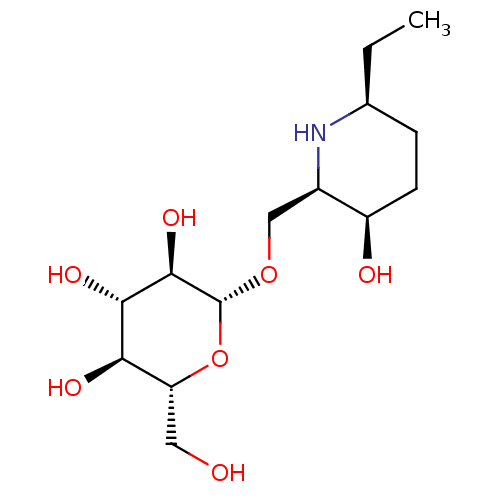
Affinity DataKi: >1.00E+6nMpH: 6.8Assay Description:Inhibition of green coffee beans alpha galactosidase at pH 6.8More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+6nMpH: 6.8Assay Description:Inhibition of green coffee beans alpha galactosidase at pH 6.8More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+6nMpH: 6.8Assay Description:Inhibition of green coffee beans alpha galactosidase at pH 6.8More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+6nMpH: 6.8Assay Description:Inhibition of green coffee beans alpha galactosidase at pH 6.8More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+6nMpH: 6.8Assay Description:Inhibition of green coffee beans alpha galactosidase at pH 6.8More data for this Ligand-Target Pair
Affinity DataIC50: 3nMpH: 6.5Assay Description:Inhibition of coffee bean alpha-galactosidase assessed as p-nitrophenol release at pH 6.5 by spectrometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 780nMpH: 6.5Assay Description:Inhibition of coffee bean alpha-galactosidase assessed as p-nitrophenol release at pH 6.5 by spectrometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMpH: 6.5Assay Description:Inhibition of coffee bean alpha-galactosidase assessed as p-nitrophenol release at pH 6.5 by spectrometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.48E+3nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 1.66E+3nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMpH: 6.5Assay Description:Inhibition of coffee bean alpha-galactosidase assessed as p-nitrophenol release at pH 6.5 by spectrometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 3.75E+3nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 4.48E+3nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 4.71E+3nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 4.90E+3nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 7.29E+3nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 9.49E+3nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 1.14E+4nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 1.25E+4nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 1.28E+4nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 1.61E+4nMpH: 10.6 T: 2°CAssay Description:In a 96-well plates, 10µL of commercial enzyme solutions without (control) or with 20µL of inhibitor were incubated at 37°C for 5 min....More data for this Ligand-Target Pair
Affinity DataIC50: 3.18E+5nMpH: 6.5Assay Description:Inhibition of coffee bean alpha-galactosidase assessed as p-nitrophenol release at pH 6.5 by spectrometric analysisMore data for this Ligand-Target Pair