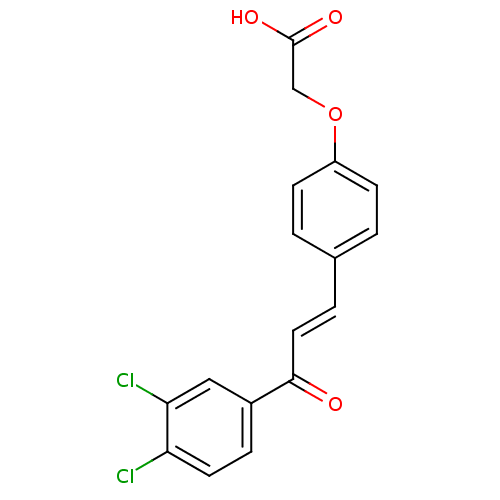
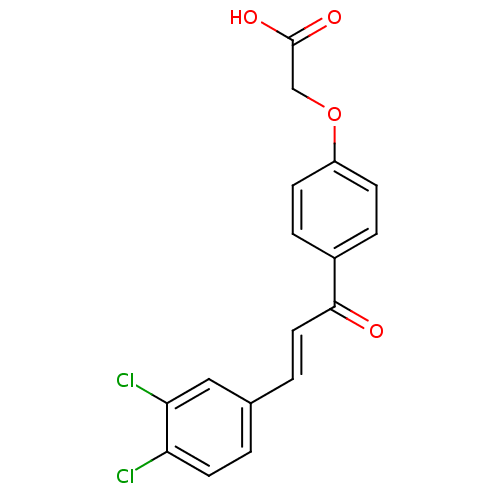
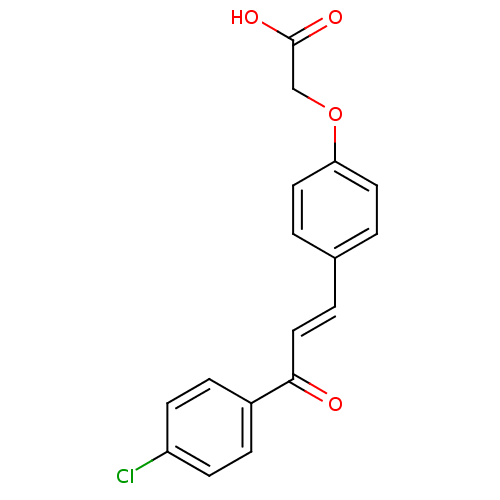
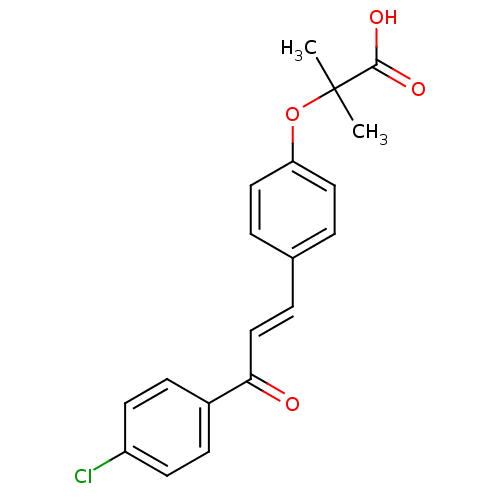
Report error Found 15 Enz. Inhib. hit(s) with all data for entry = 3315
Affinity DataIC50: 4.90E+4nMpH: 7.4 T: 2°CAssay Description:Interference of the p53-MDM2 binding by test compounds was measured in a 96-well polypropylene round-bottom microtiter plate (Costar, Serocluster). H...More data for this Ligand-Target Pair
Affinity DataKd: 9.00E+4nMpH: 7.4 T: 2°CAssay Description:NMR measurements consisted of monitoring changes in chemical shifts and line widths of the backbone amide resonances of uniformly 15N-enriched MDM2 s...More data for this Ligand-Target Pair
Affinity DataIC50: 1.17E+5nMpH: 7.4 T: 2°CAssay Description:Interference of the p53-MDM2 binding by test compounds was measured in a 96-well polypropylene round-bottom microtiter plate (Costar, Serocluster). H...More data for this Ligand-Target Pair
Affinity DataKd: 1.50E+5nMpH: 7.4 T: 2°CAssay Description:NMR measurements consisted of monitoring changes in chemical shifts and line widths of the backbone amide resonances of uniformly 15N-enriched MDM2 s...More data for this Ligand-Target Pair
Affinity DataIC50: 2.06E+5nMpH: 7.4 T: 2°CAssay Description:Interference of the p53-MDM2 binding by test compounds was measured in a 96-well polypropylene round-bottom microtiter plate (Costar, Serocluster). H...More data for this Ligand-Target Pair
Affinity DataKd: 2.20E+5nMpH: 7.4 T: 2°CAssay Description:NMR measurements consisted of monitoring changes in chemical shifts and line widths of the backbone amide resonances of uniformly 15N-enriched MDM2 s...More data for this Ligand-Target Pair
Affinity DataKd: 2.44E+5nMpH: 7.4 T: 2°CAssay Description:NMR measurements consisted of monitoring changes in chemical shifts and line widths of the backbone amide resonances of uniformly 15N-enriched MDM2 s...More data for this Ligand-Target Pair
Affinity DataKd: 2.50E+5nMpH: 7.4 T: 2°CAssay Description:NMR measurements consisted of monitoring changes in chemical shifts and line widths of the backbone amide resonances of uniformly 15N-enriched MDM2 s...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMpH: 7.4 T: 2°CAssay Description:Interference of the p53-MDM2 binding by test compounds was measured in a 96-well polypropylene round-bottom microtiter plate (Costar, Serocluster). H...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMpH: 7.4 T: 2°CAssay Description:Interference of the p53-MDM2 binding by test compounds was measured in a 96-well polypropylene round-bottom microtiter plate (Costar, Serocluster). H...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMpH: 7.4 T: 2°CAssay Description:Interference of the p53-MDM2 binding by test compounds was measured in a 96-well polypropylene round-bottom microtiter plate (Costar, Serocluster). H...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMpH: 7.4 T: 2°CAssay Description:Interference of the p53-MDM2 binding by test compounds was measured in a 96-well polypropylene round-bottom microtiter plate (Costar, Serocluster). H...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMpH: 7.4 T: 2°CAssay Description:Interference of the p53-MDM2 binding by test compounds was measured in a 96-well polypropylene round-bottom microtiter plate (Costar, Serocluster). H...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMpH: 7.4 T: 2°CAssay Description:Interference of the p53-MDM2 binding by test compounds was measured in a 96-well polypropylene round-bottom microtiter plate (Costar, Serocluster). H...More data for this Ligand-Target Pair
Affinity DataKd: 2.70E+5nMpH: 7.4 T: 2°CAssay Description:NMR measurements consisted of monitoring changes in chemical shifts and line widths of the backbone amide resonances of uniformly 15N-enriched MDM2 s...More data for this Ligand-Target Pair