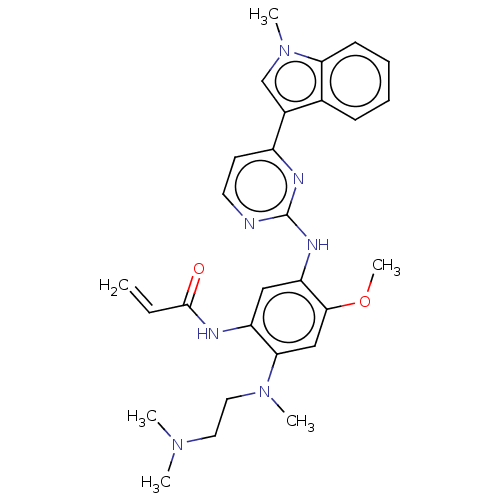
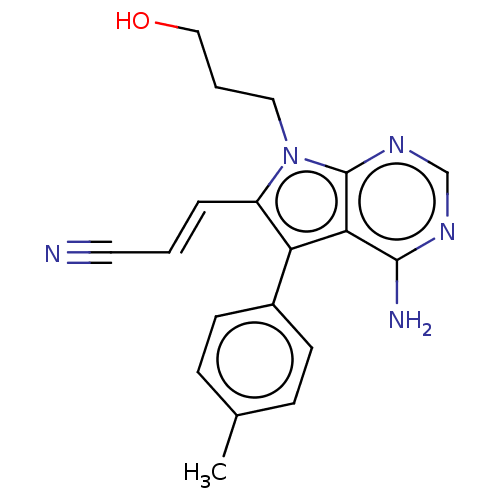
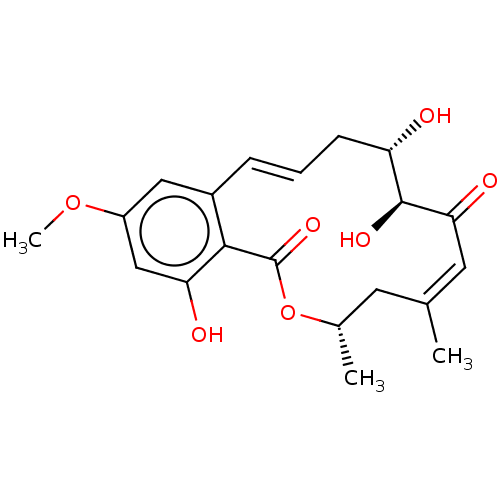
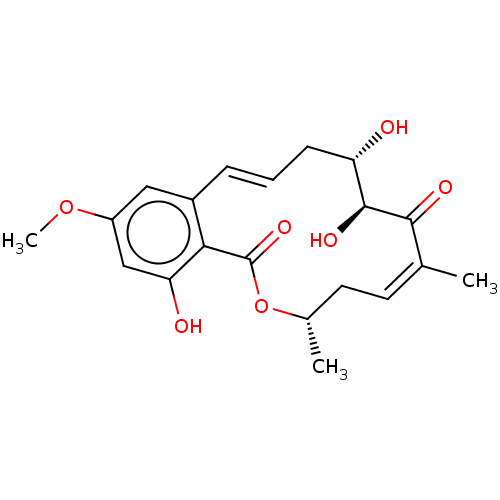
Report error Found 22 Enz. Inhib. hit(s) with all data for entry = 50000292
Affinity DataIC50: 1nMAssay Description:Inhibition of JNK3 (unknown origin) after 1 hr by FRET-based assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of JNK2 (unknown origin) after 1 hrMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of CRM1-mediated nucleocytoplasmic transport in human HeLa cells after 90 mins by immunofluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of JNK1 (unknown origin) after 1 hrMore data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of human calpain 1 protease using Ac-LLY-AFC as substrate after 1 hr by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of RSK2 in human MDA-MB-231 cells after 2 hrsMore data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:Inhibition of RSK2 in human MDA-MB-231 cells after 2 hrsMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of CRM1-mediated nucleocytoplasmic transport in human HeLa cells after 90 mins by immunofluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Inhibition of TNFalpha-PLAP (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:Inhibition of CRM1-mediated nucleocytoplasmic transport in human HeLa cells after 90 mins by immunofluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 36nMAssay Description:Inhibition of TNFalpha-PLAP (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 250nMAssay Description:Inhibition of RSK2 in human MDA-MB-231 cells after 2 hrsMore data for this Ligand-Target Pair
Affinity DataKi: 303nMAssay Description:Binding affinity to recombinant human GST-tagged EGFR (668 to 1210 residues) cytoplasmic domain expressed in baculovirus expression systemMore data for this Ligand-Target Pair
Affinity DataIC50: 480nMAssay Description:Inhibition of EGF-stimulated wild type EGFR phosphorylation in human LoVo cells incubated for 2 hrs followed by EGF stimulation for 10 mins by ELISAMore data for this Ligand-Target Pair
Affinity DataIC50: 750nMAssay Description:Inhibition of RSK2 in human MDA-MB-231 cells after 2 hrsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of TNFalpha-PLAP (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of TNFalpha-PLAP (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 4.68E+4nMAssay Description:Inhibition of [3H]dopamine uptake at DAT in rat brain striatal synaptosomes by liquid scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 5.25E+4nMAssay Description:Inhibition of [3H]dopamine uptake at DAT in rat brain striatal synaptosomes by liquid scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 3.31E+5nMAssay Description:Inhibition of [3H]dopamine uptake at DAT in rat brain striatal synaptosomes by liquid scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 4.38E+8nMAssay Description:Inhibition of [3H]dopamine uptake at DAT in rat brain striatal synaptosomes by liquid scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 4.61E+8nMAssay Description:Inhibition of [3H]dopamine uptake at DAT in rat brain striatal synaptosomes by liquid scintillation counting analysisMore data for this Ligand-Target Pair
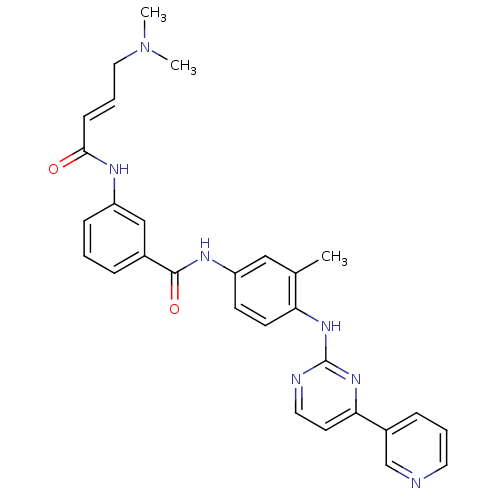
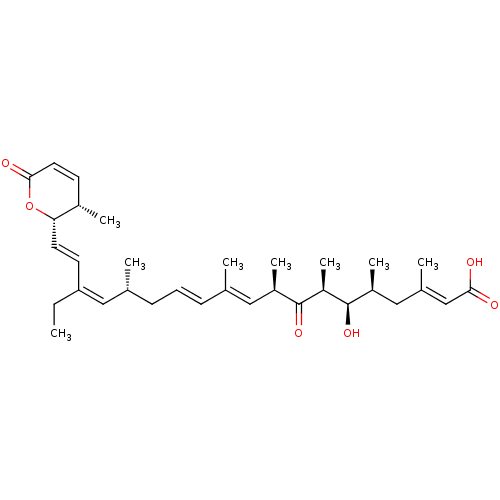
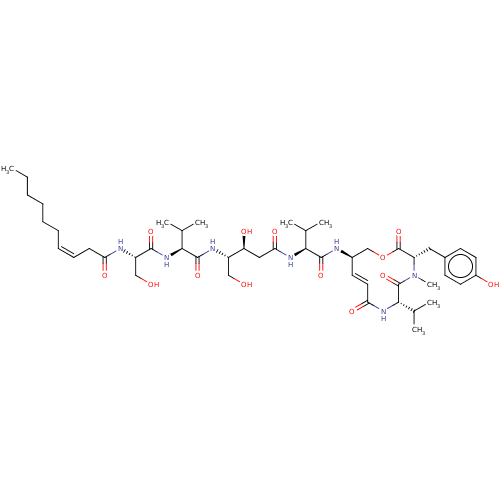
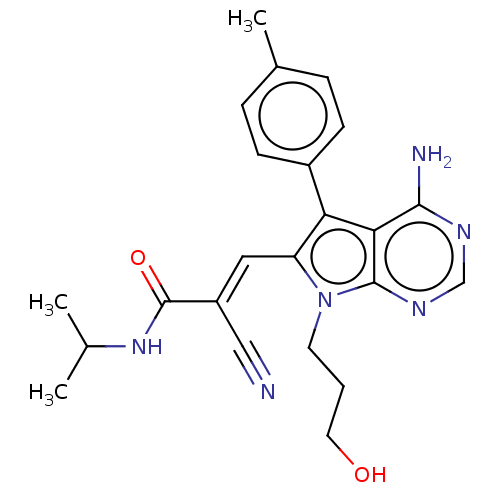


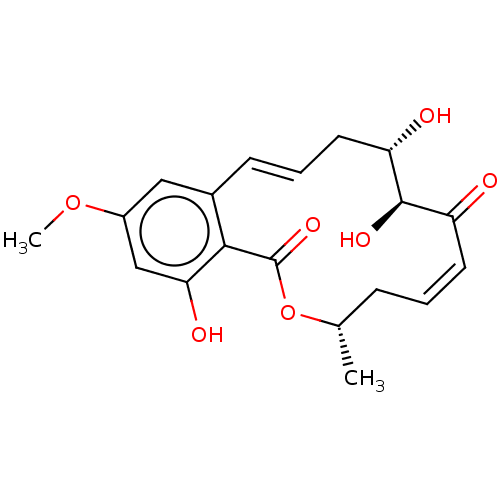

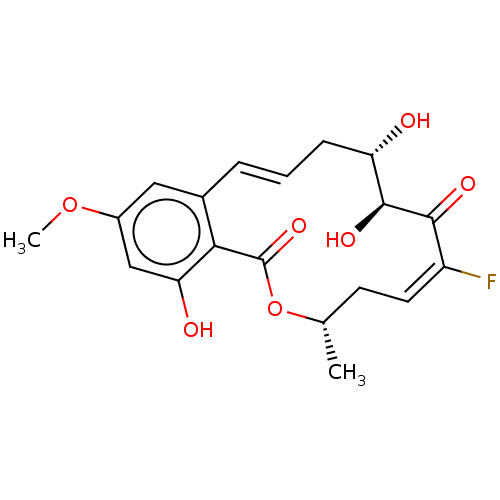
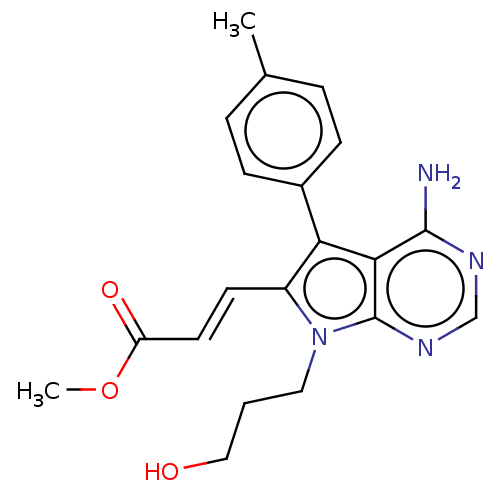
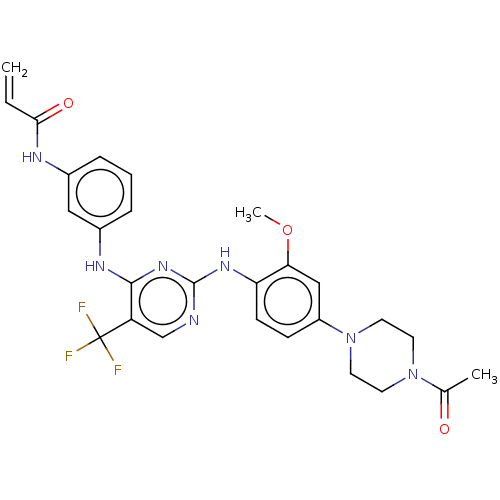
 3D Structure (crystal)
3D Structure (crystal)