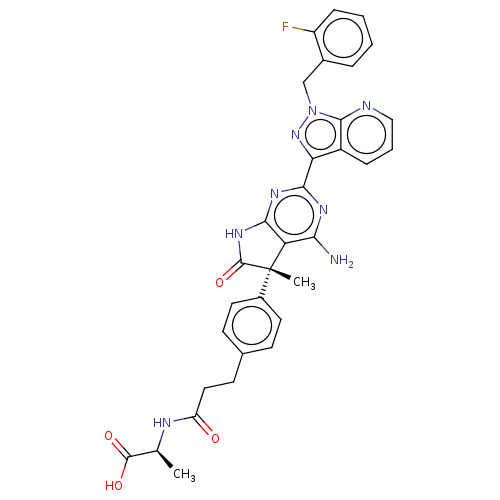
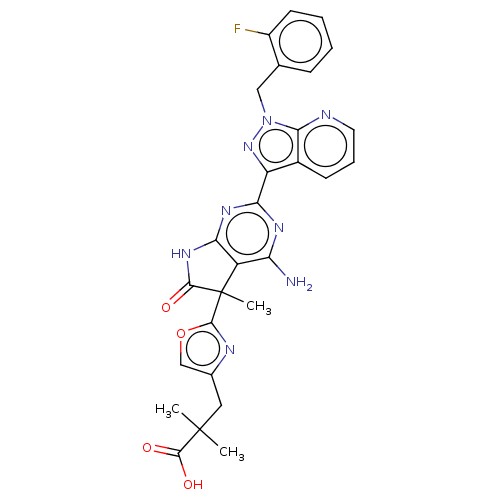
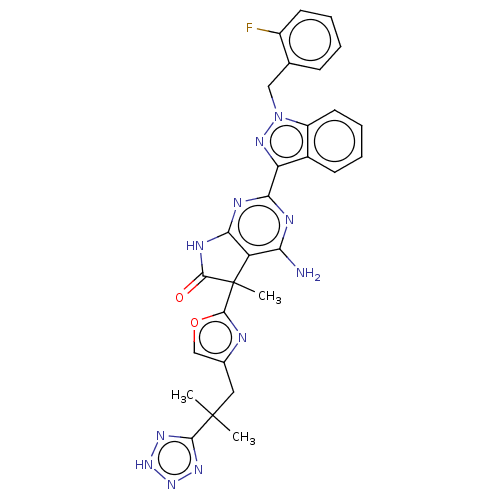
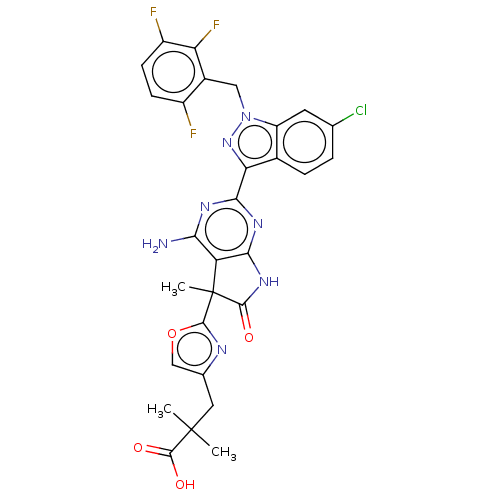
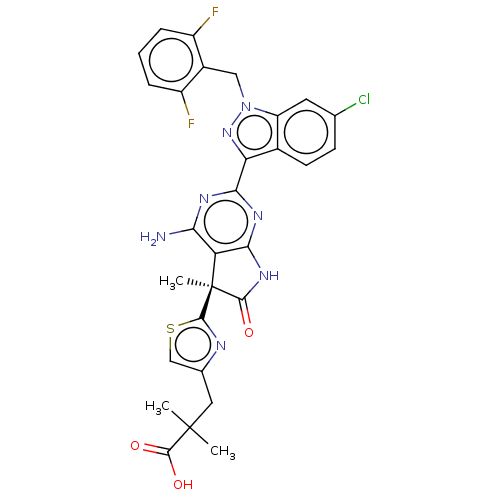
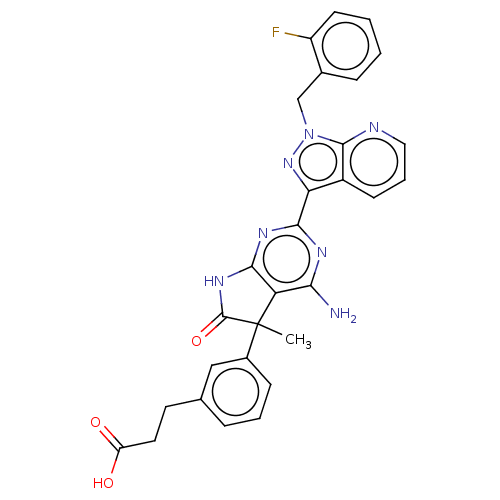
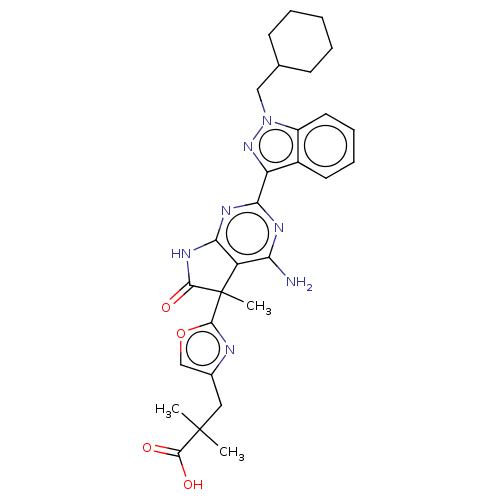
Report error Found 247 Enz. Inhib. hit(s) with all data for entry = 8736
Affinity DataKi: 0.0220nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0260nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0290nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0310nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0310nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0310nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0390nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0410nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0420nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0450nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0470nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0480nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0490nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0520nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0520nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0560nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0570nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0570nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0580nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0580nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0580nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0620nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0620nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0620nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0630nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0630nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0640nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0660nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0660nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0670nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0670nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0670nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0680nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0680nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0700nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0700nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0720nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0730nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0740nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0740nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0740nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0750nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0750nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0750nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0760nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0770nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0780nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair
Affinity DataKi: 0.0790nMAssay Description:The binding buffer was composed of 50 mM triethanolamine, pH 7.4, 3 mM MgCl2, 0.025% BSA, 2 mM dithiothreitol (DTT), 300 μM DETA/NO and 400 _...More data for this Ligand-Target Pair