Report error Found 89 Enz. Inhib. hit(s) with Target = '3-hydroxyacyl-[acyl-carrier-protein] dehydratase'
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
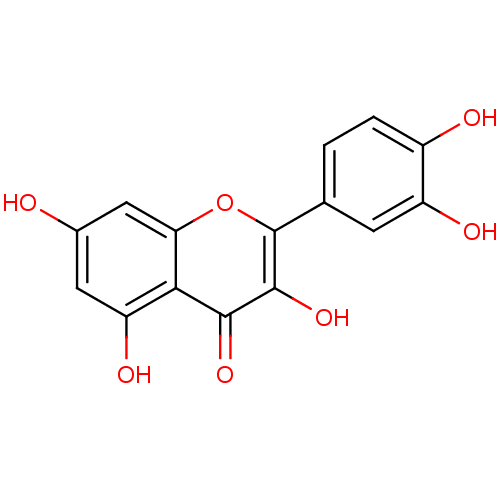
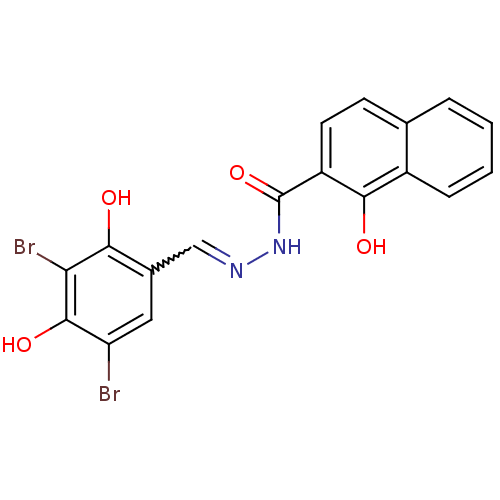
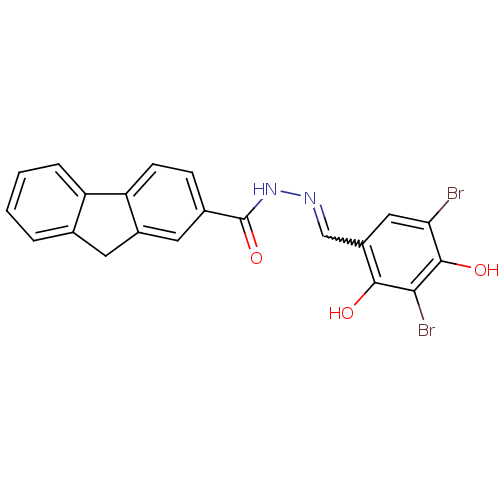
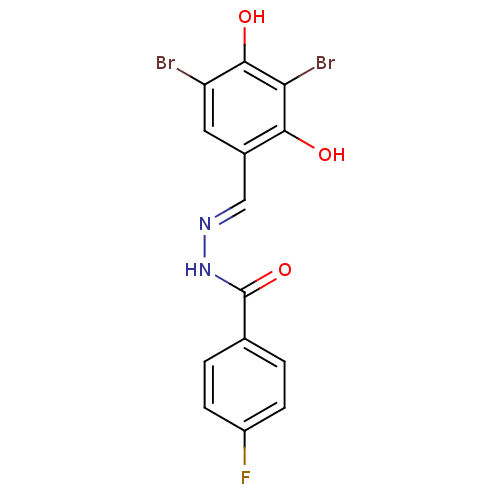
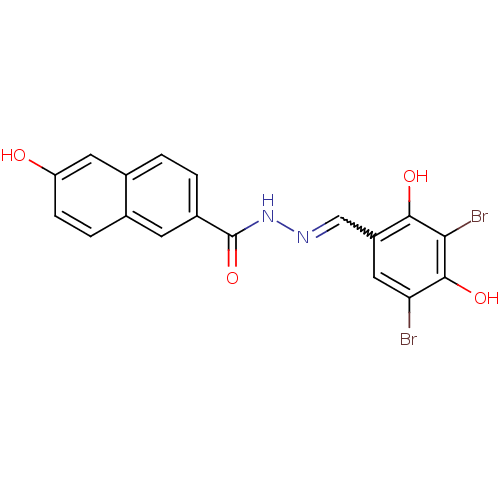
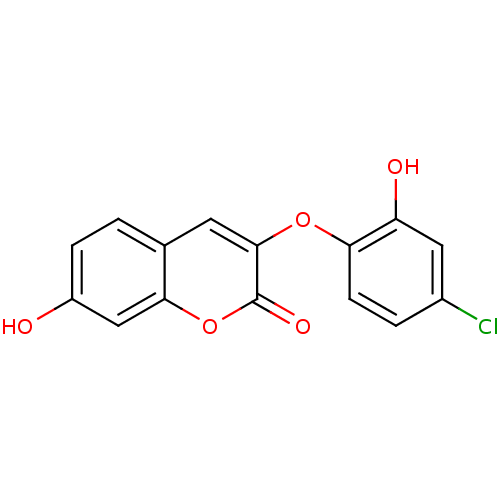
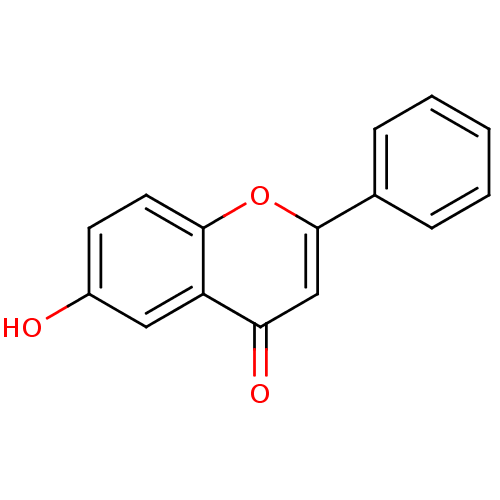
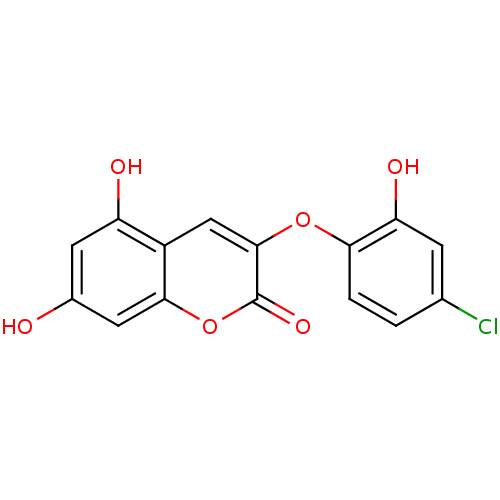
Affinity DataIC50: 30nMAssay Description:Inhibition of recombinant Plasmodium falciparum FabZ using crotonoyl-CoA as substrate after 10 minsMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Shanghai Institute of Materia Medica
Shanghai Institute of Materia Medica
Affinity DataIC50: 860nMAssay Description:The enzymatic inhibition assay of HpFabZ was monitored by the spectrophotometric method. The activity of HpFabZ was measured by detection of the decr...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Shanghai Institute of Materia Medica
Shanghai Institute of Materia Medica
Affinity DataIC50: 1.42E+3nMpH: 8.0 T: 2°CAssay Description:The enzymatic inhibition assay of HpFabZ was monitored by the spectrophotometric method. The activity of HpFabZ was measured by detection of the decr...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of FabZMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Shanghai Institute of Materia Medica
Shanghai Institute of Materia Medica
Affinity DataIC50: 1.50E+3nMpH: 8.0 T: 2°CAssay Description:The enzymatic inhibition assay of HpFabZ was monitored by the spectrophotometric method. The activity of HpFabZ was measured by detection of the decr...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Shanghai Institute of Materia Medica
Shanghai Institute of Materia Medica
Affinity DataIC50: 1.52E+3nMpH: 8.0 T: 2°CAssay Description:The enzymatic inhibition assay of HpFabZ was monitored by the spectrophotometric method. The activity of HpFabZ was measured by detection of the decr...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Shanghai Institute of Materia Medica
Shanghai Institute of Materia Medica
Affinity DataIC50: 1.52E+3nMpH: 8.0 T: 2°CAssay Description:The enzymatic inhibition assay of HpFabZ was monitored by the spectrophotometric method. The activity of HpFabZ was measured by detection of the decr...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Shanghai Institute of Materia Medica
Shanghai Institute of Materia Medica
Affinity DataIC50: 1.97E+3nMAssay Description:The enzymatic inhibition assay of HpFabZ was monitored by the spectrophotometric method. The activity of HpFabZ was measured by detection of the decr...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Shanghai Institute of Materia Medica
Shanghai Institute of Materia Medica
Affinity DataIC50: 1.98E+3nMpH: 8.0 T: 2°CAssay Description:The enzymatic inhibition assay of HpFabZ was monitored by the spectrophotometric method. The activity of HpFabZ was measured by detection of the decr...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of FabZMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of FabZMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Shanghai Institute of Materia Medica
Shanghai Institute of Materia Medica
Affinity DataIC50: 2.13E+3nMAssay Description:The enzymatic inhibition assay of HpFabZ was monitored by the spectrophotometric method. The activity of HpFabZ was measured by detection of the decr...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Shanghai Institute of Materia Medica
Shanghai Institute of Materia Medica
Affinity DataIC50: 3.55E+3nMAssay Description:The enzymatic inhibition assay of HpFabZ was monitored by the spectrophotometric method. The activity of HpFabZ was measured by detection of the decr...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Shanghai Institute of Materia Medica
Shanghai Institute of Materia Medica
Affinity DataIC50: 3.75E+3nMAssay Description:The enzymatic inhibition assay of HpFabZ was monitored by the spectrophotometric method. The activity of HpFabZ was measured by detection of the decr...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataKi: 4.80E+3nMAssay Description:Competitive inhibition of Plasmodium falciparum FabZ using crotonyl-CoA as substrate by Michaelis-Menten steady state analysisMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of FabZMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ [1-175](Yersinia pestis)
Brookhaven National Laboratory
Brookhaven National Laboratory
Affinity DataIC50: 5.30E+3nMpH: 7.0 T: 2°CAssay Description:The enzymatic activities of FtFabZ and YpFabZ were determined via the reportedspectrophotometric method using the substrate analogue crotonoyl-CoA.7 ...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Francisella tularensis)
Brookhaven National Laboratory
Brookhaven National Laboratory
Affinity DataIC50: 5.40E+3nMpH: 7.0 T: 2°CAssay Description:The enzymatic activities of FtFabZ and YpFabZ were determined via the reportedspectrophotometric method using the substrate analogue crotonoyl-CoA.7 ...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ [1-175](Yersinia pestis)
Brookhaven National Laboratory
Brookhaven National Laboratory
Affinity DataIC50: 6.10E+3nMpH: 7.0 T: 2°CAssay Description:The enzymatic activities of FtFabZ and YpFabZ were determined via the reportedspectrophotometric method using the substrate analogue crotonoyl-CoA.7 ...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Shanghai Institute of Materia Medica
Shanghai Institute of Materia Medica
Affinity DataIC50: 6.43E+3nMpH: 8.0 T: 2°CAssay Description:The enzymatic inhibition assay of HpFabZ was monitored by the spectrophotometric method. The activity of HpFabZ was measured by detection of the decr...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 7.00E+3nMAssay Description:Inhibition of FabZMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Francisella tularensis)
Brookhaven National Laboratory
Brookhaven National Laboratory
Affinity DataIC50: 7.70E+3nMpH: 7.0 T: 2°CAssay Description:The enzymatic activities of FtFabZ and YpFabZ were determined via the reportedspectrophotometric method using the substrate analogue crotonoyl-CoA.7 ...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of FabZMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of FabZMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Shanghai Institute of Materia Medica
Shanghai Institute of Materia Medica
Affinity DataIC50: 8.77E+3nMAssay Description:The enzymatic inhibition assay of HpFabZ was monitored by the spectrophotometric method. The activity of HpFabZ was measured by detection of the decr...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Shanghai Institute of Materia Medica
Shanghai Institute of Materia Medica
Affinity DataIC50: 9.92E+3nMpH: 8.0 T: 2°CAssay Description:The enzymatic inhibition assay of HpFabZ was monitored by the spectrophotometric method. The activity of HpFabZ was measured by detection of the decr...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of recombinant Plasmodium falciparum FabZ using crotonoyl-CoA as substrate after 10 minsMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Shanghai Institute of Materia Medica
Shanghai Institute of Materia Medica
Affinity DataIC50: 1.28E+4nMAssay Description:The enzymatic inhibition assay of HpFabZ was monitored by the spectrophotometric method. The activity of HpFabZ was measured by detection of the decr...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ [1-175](Yersinia pestis)
Brookhaven National Laboratory
Brookhaven National Laboratory
Affinity DataIC50: 1.30E+4nMpH: 7.0 T: 2°CAssay Description:The enzymatic activities of FtFabZ and YpFabZ were determined via the reportedspectrophotometric method using the substrate analogue crotonoyl-CoA.7 ...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 1.30E+4nMAssay Description:Inhibition of recombinant Plasmodium falciparum FabZ using crotonoyl-CoA as substrate after 10 minsMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 1.63E+4nMAssay Description:Inhibition of recombinant Plasmodium falciparum FabZ using crotonoyl-CoA as substrate after 10 minsMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of FabZMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Francisella tularensis)
Brookhaven National Laboratory
Brookhaven National Laboratory
Affinity DataIC50: 2.78E+4nMpH: 7.0 T: 2°CAssay Description:The enzymatic activities of FtFabZ and YpFabZ were determined via the reportedspectrophotometric method using the substrate analogue crotonoyl-CoA.7 ...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 2.80E+4nMAssay Description:Inhibition of recombinant Plasmodium falciparum FabZ using crotonoyl-CoA as substrate after 10 minsMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ [1-175](Yersinia pestis)
Brookhaven National Laboratory
Brookhaven National Laboratory
Affinity DataIC50: 2.97E+4nMpH: 7.0 T: 2°CAssay Description:The enzymatic activities of FtFabZ and YpFabZ were determined via the reportedspectrophotometric method using the substrate analogue crotonoyl-CoA.7 ...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of FabZMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of FabZMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Shanghai Institute of Materia Medica
Shanghai Institute of Materia Medica
Affinity DataIC50: 3.98E+4nMpH: 8.0 T: 2°CAssay Description:The enzymatic inhibition assay of HpFabZ was monitored by the spectrophotometric method. The activity of HpFabZ was measured by detection of the decr...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibition of FabZMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Shanghai Institute of Materia Medica
Shanghai Institute of Materia Medica
Affinity DataIC50: 4.27E+4nMAssay Description:The enzymatic inhibition assay of HpFabZ was monitored by the spectrophotometric method. The activity of HpFabZ was measured by detection of the decr...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase FabZ(Francisella tularensis)
Brookhaven National Laboratory
Brookhaven National Laboratory
Affinity DataIC50: 4.31E+4nMpH: 7.0 T: 2°CAssay Description:The enzymatic activities of FtFabZ and YpFabZ were determined via the reportedspectrophotometric method using the substrate analogue crotonoyl-CoA.7 ...More data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 9.00E+4nMAssay Description:Inhibition of recombinant Plasmodium falciparum FabZ using crotonoyl-CoA as substrate after 10 minsMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant Plasmodium falciparum FabZ using crotonoyl-CoA as substrate after 10 minsMore data for this Ligand-Target Pair
Target3-hydroxyacyl-[acyl-carrier-protein] dehydratase(malaria parasite P. falciparum)
University of Bologna
Curated by ChEMBL
University of Bologna
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant Plasmodium falciparum FabZ using crotonoyl-CoA as substrate after 10 minsMore data for this Ligand-Target Pair
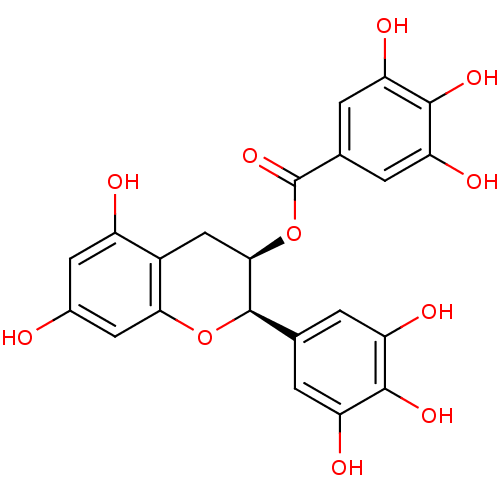
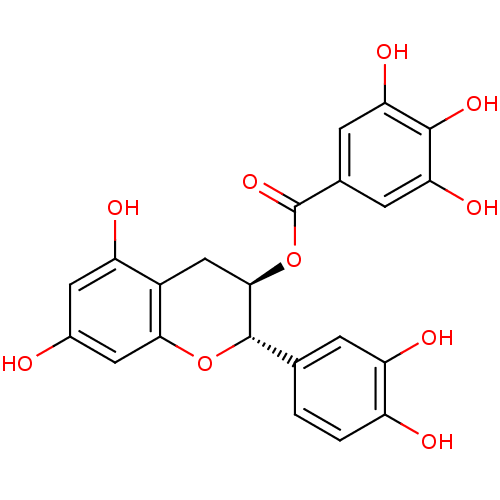
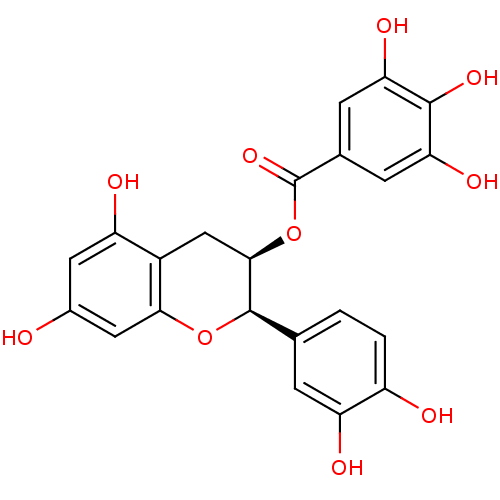
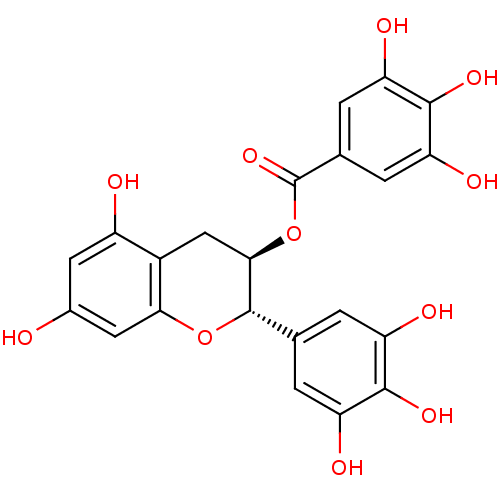
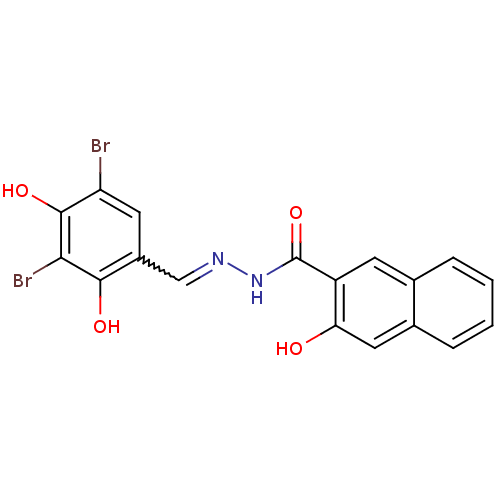
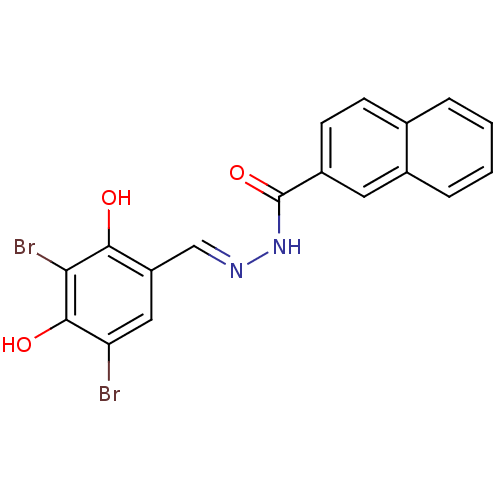
 3D Structure (crystal)
3D Structure (crystal)