Report error Found 269 Enz. Inhib. hit(s) with Target = 'Mitochondrial import inner membrane translocase subunit TIM23'
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
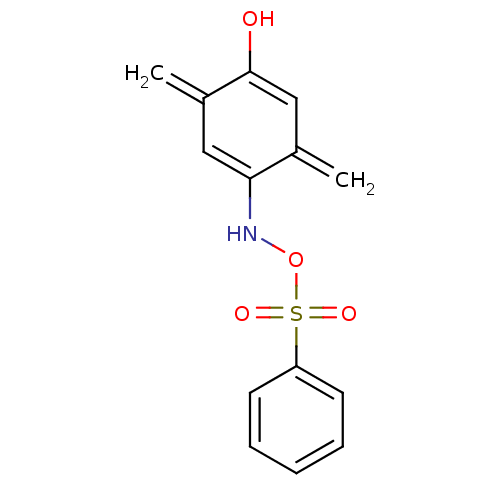
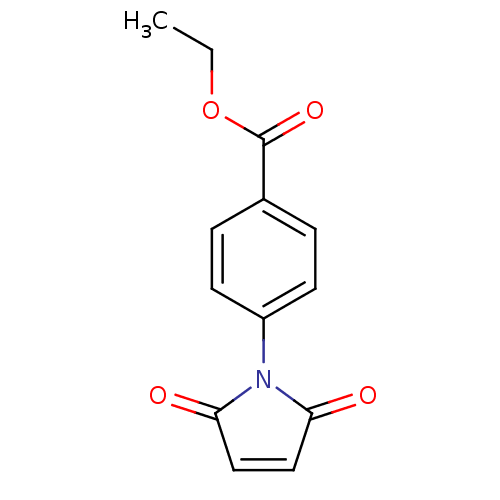
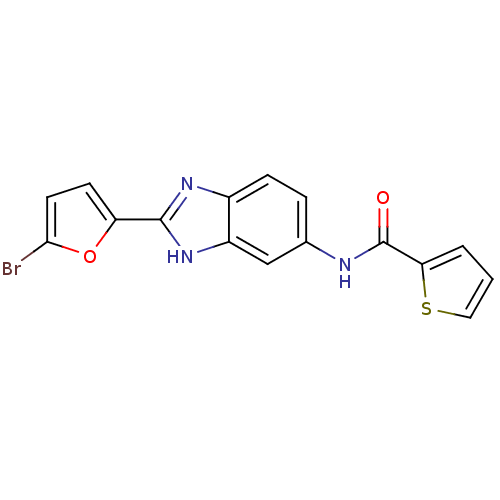
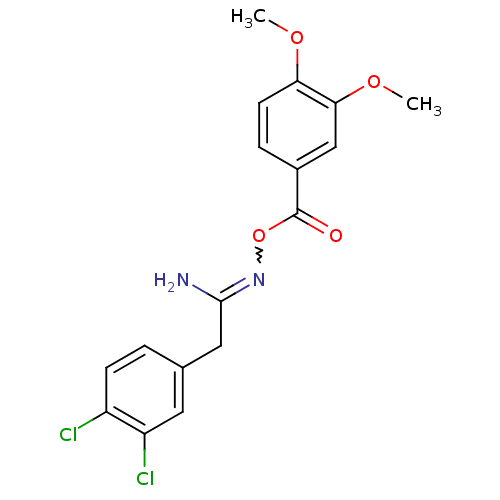
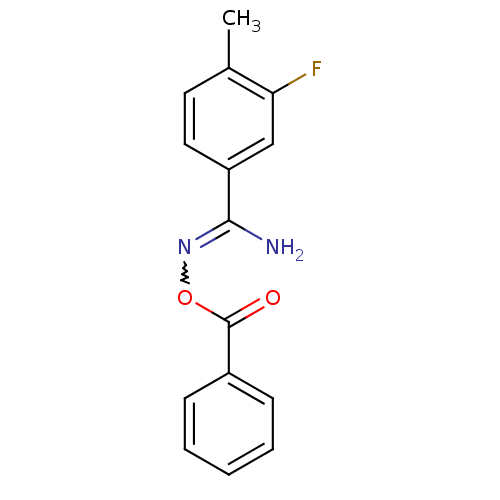
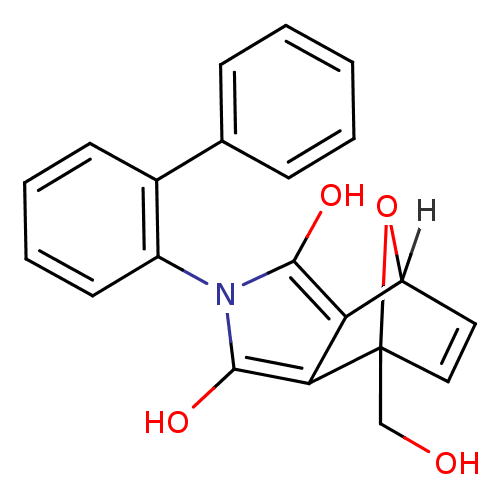
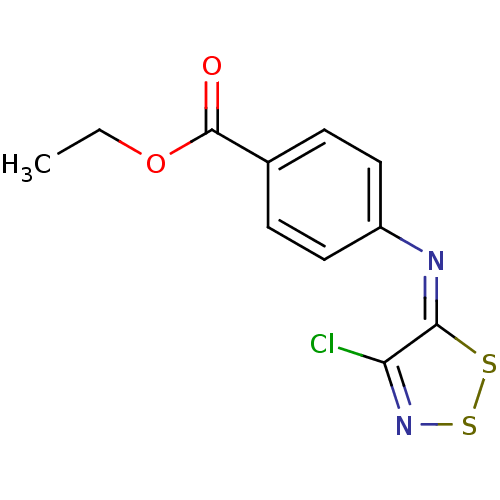
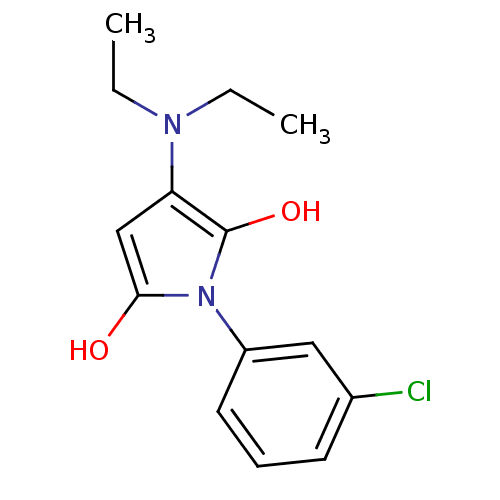
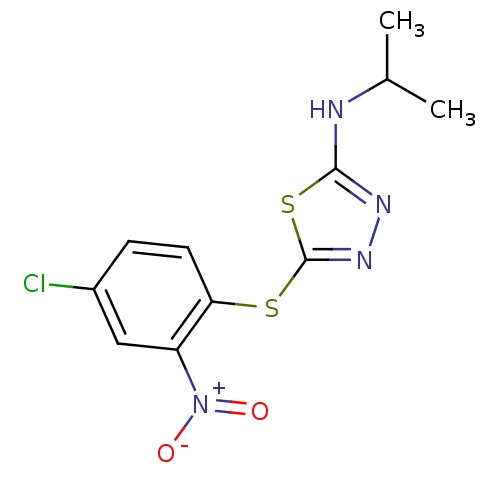
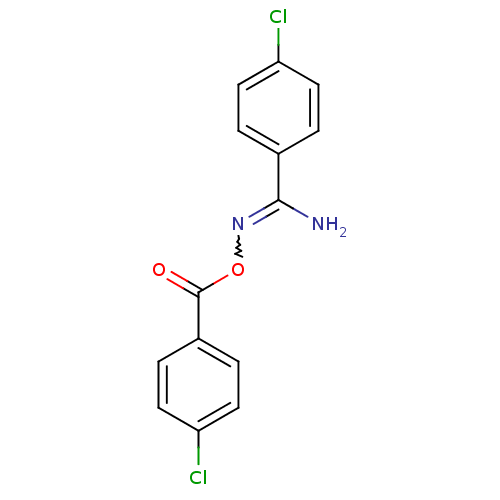
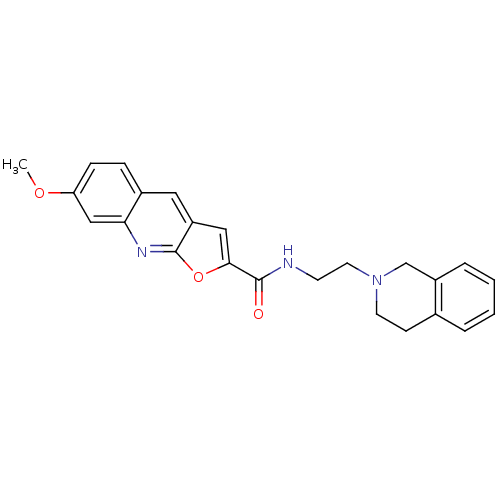
Affinity DataIC50: 200nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
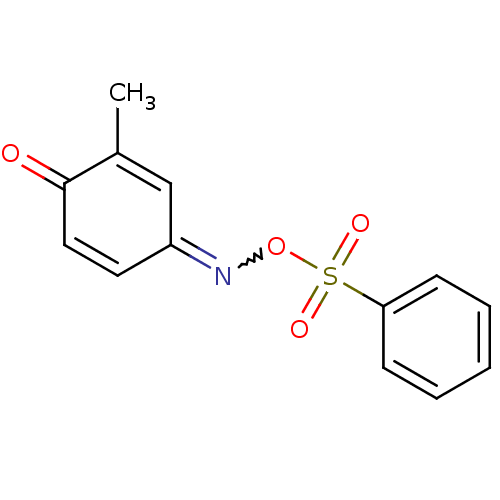
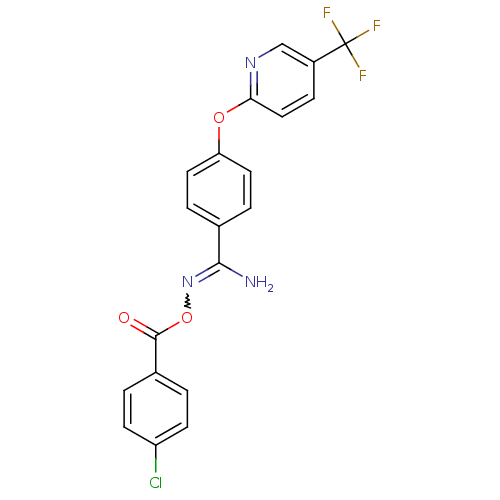
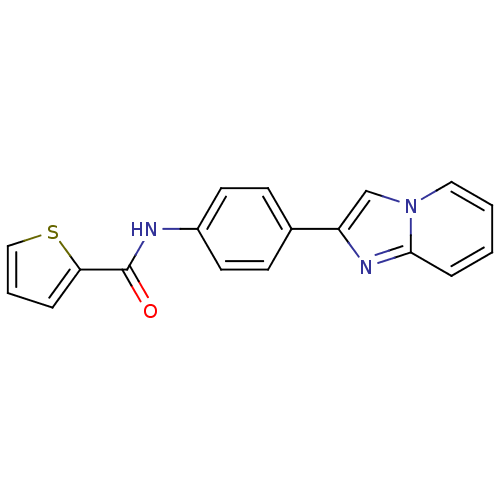
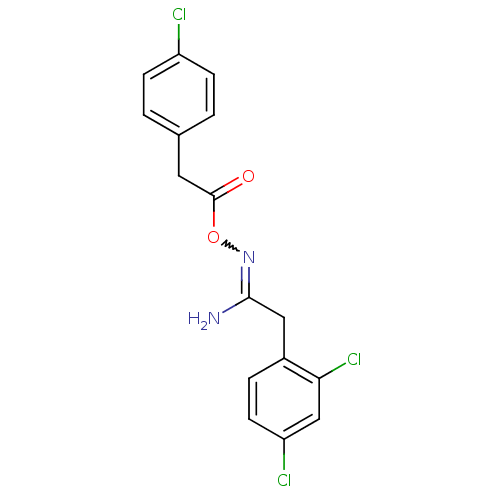
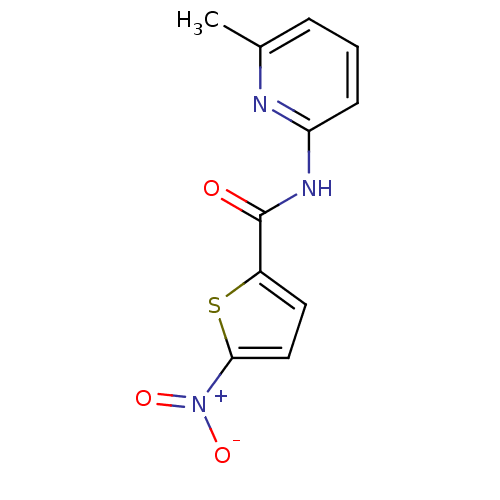
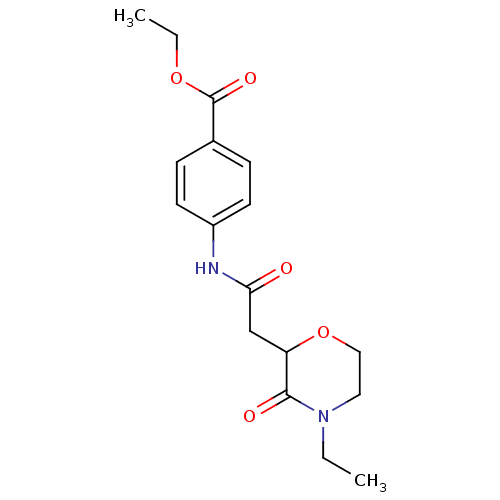
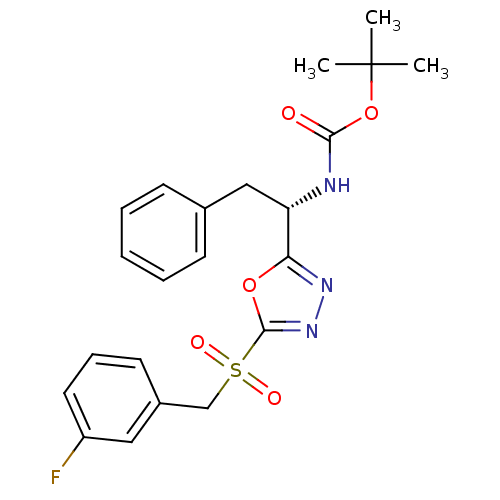
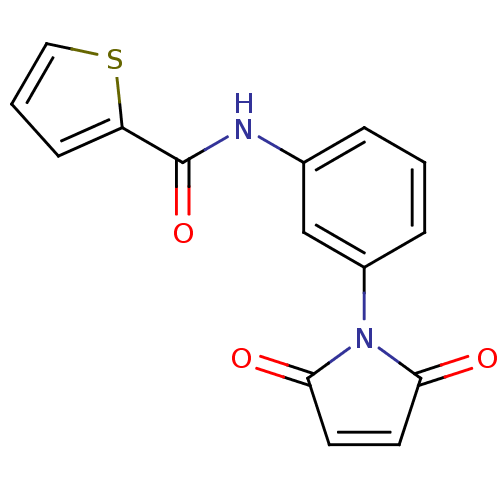
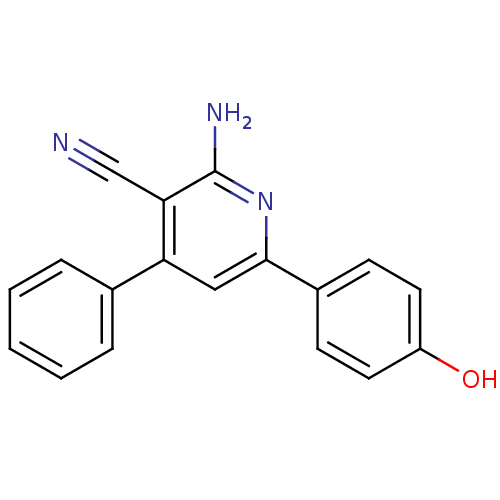
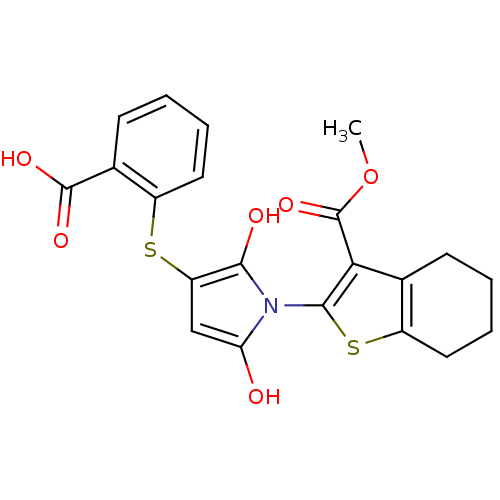
Affinity DataIC50: 270nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
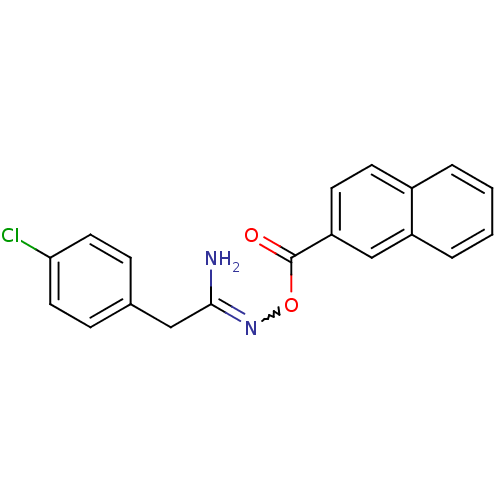
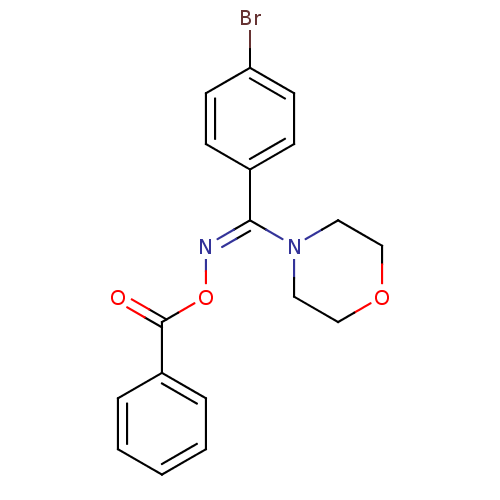
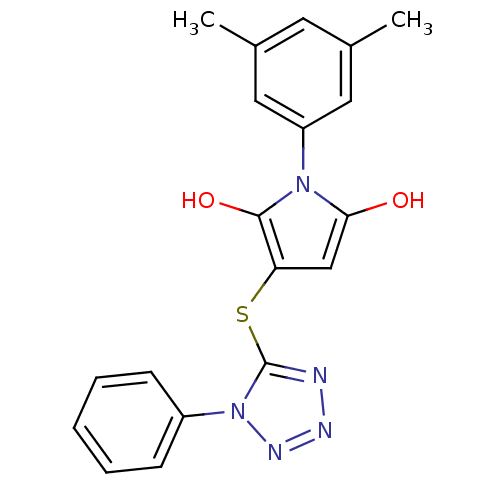
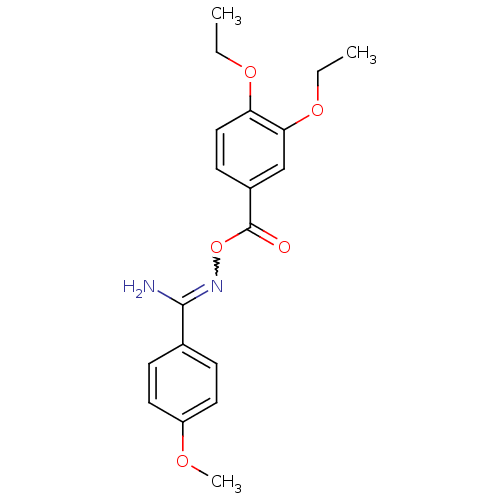
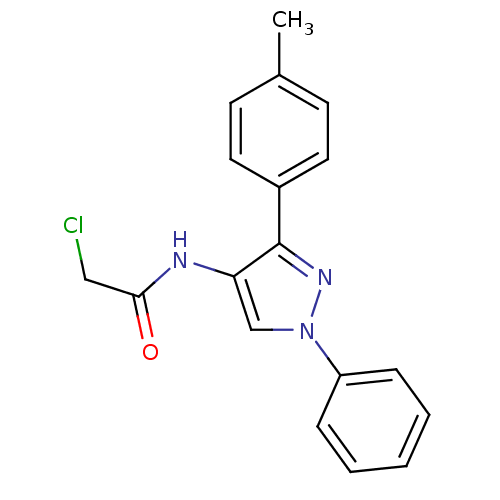
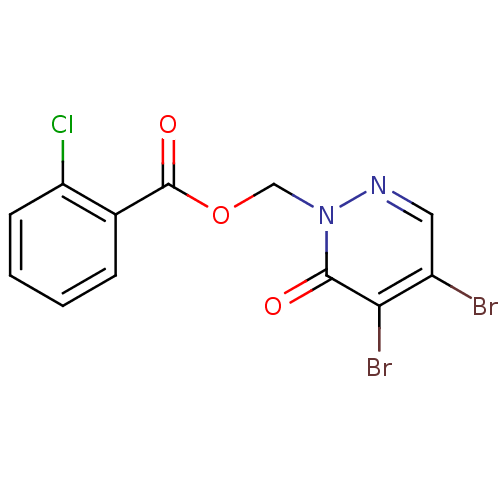
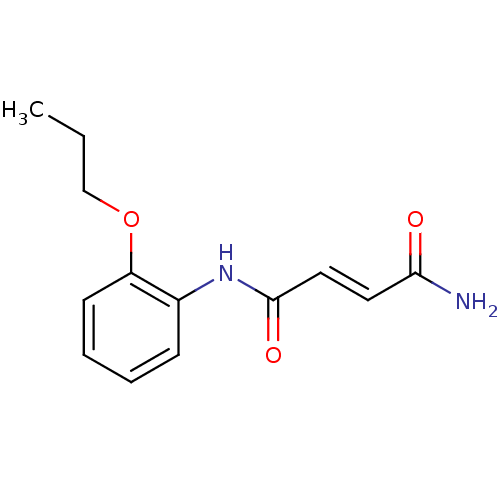
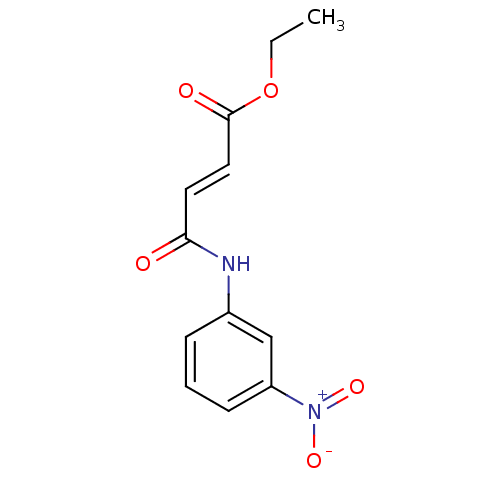
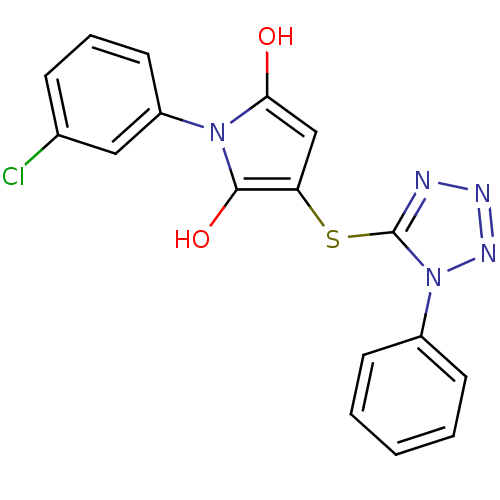
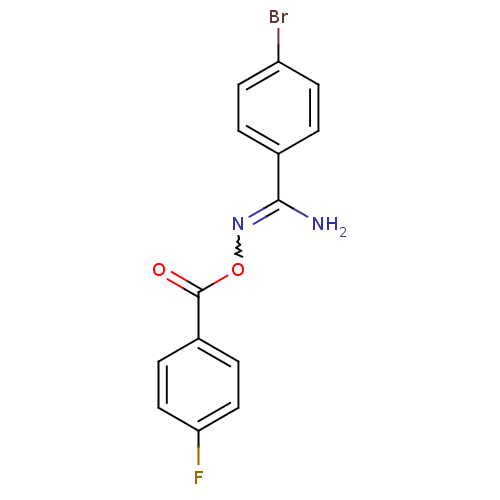
Affinity DataIC50: 380nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
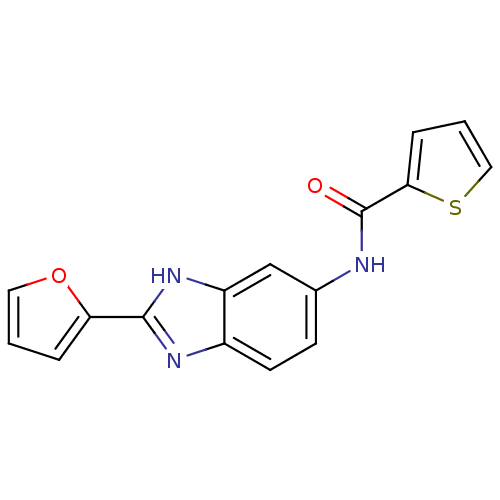
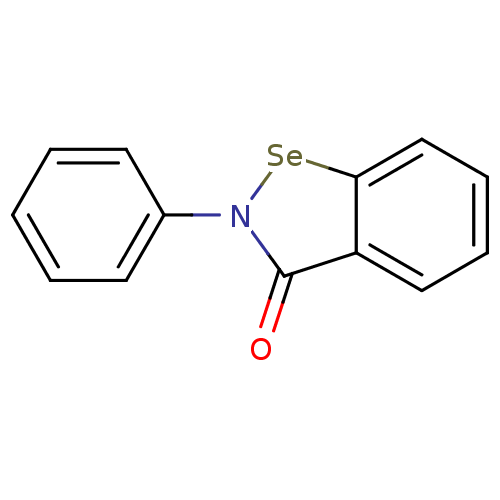
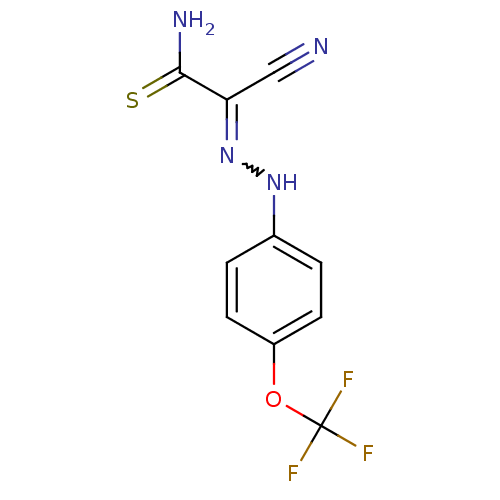
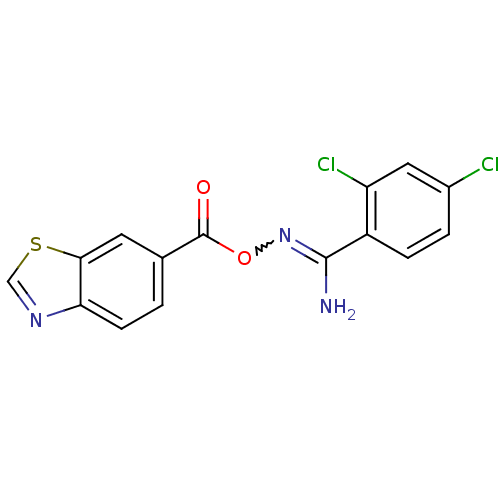
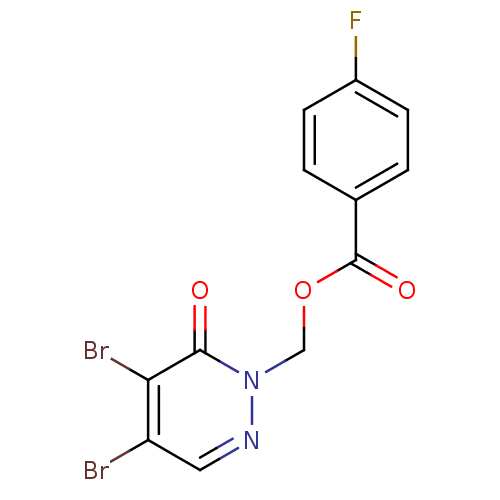
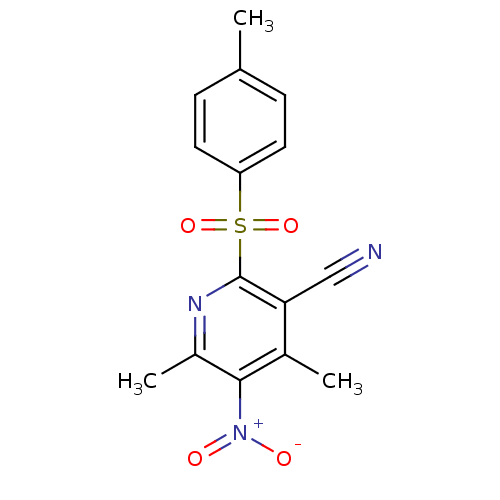
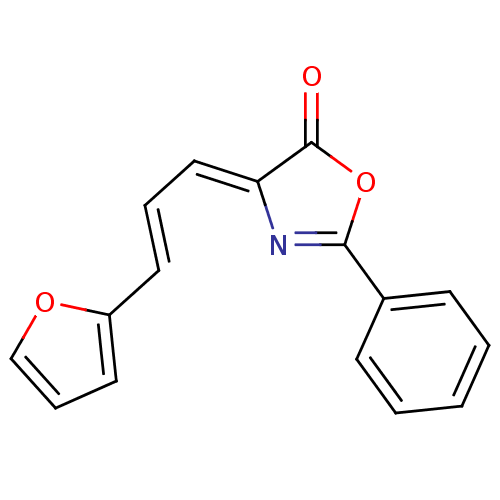
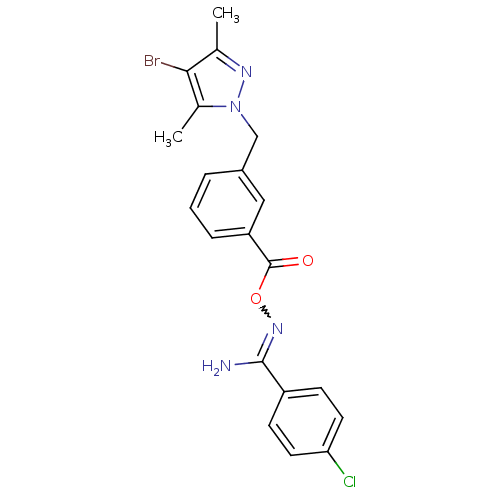
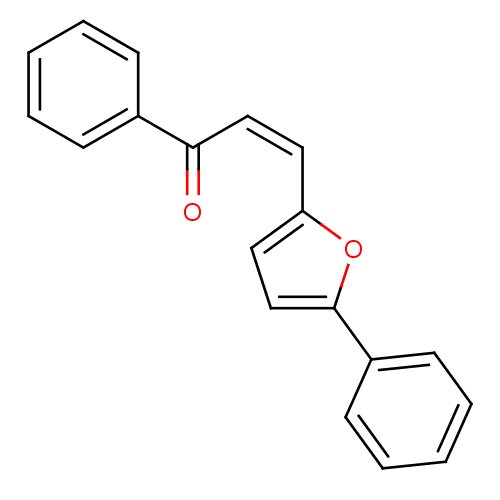
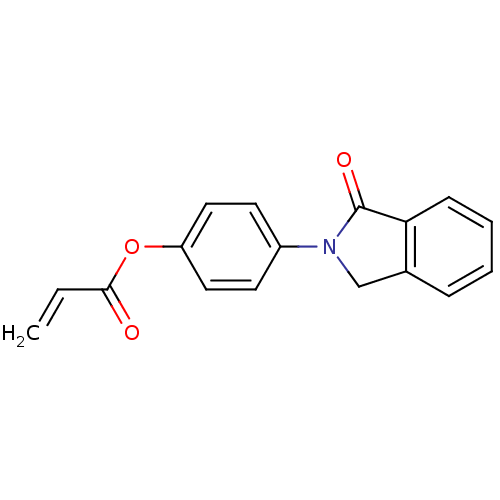
Affinity DataIC50: 459nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 480nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 570nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 617nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 787nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 880nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 882nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 897nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 987nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 1.06E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 1.26E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 1.31E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 1.38E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 1.39E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 1.42E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 1.48E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 1.48E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 1.50E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 1.52E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 1.55E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 1.57E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 1.72E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 1.75E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 1.78E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 1.83E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 2.02E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 2.07E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 2.08E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 2.22E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 2.28E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 2.30E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 2.34E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 2.37E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 2.44E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 2.46E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 2.52E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 2.61E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 2.65E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 2.66E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 2.90E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 2.92E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 3.01E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 3.02E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 3.13E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 3.14E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 3.33E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 3.34E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair