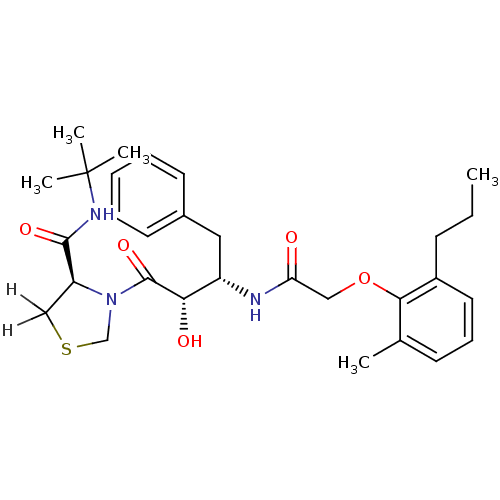
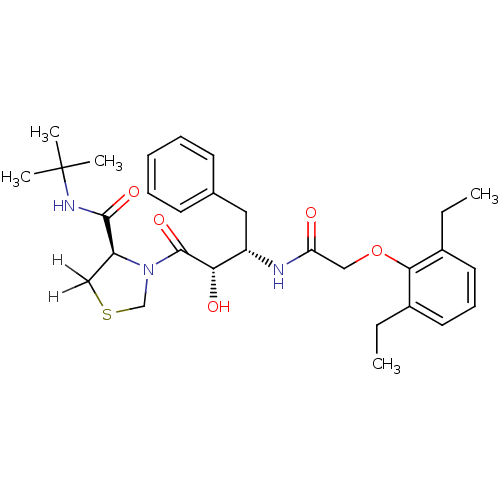
Report error Found 18 Enz. Inhib. hit(s) with all data for entry = 333
Affinity DataKi: 0.0880nM ΔG°: -59.7kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 0.160nM ΔG°: -58.2kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 0.290nM ΔG°: -56.6kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 0.330nM ΔG°: -56.3kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 0.740nM ΔG°: -54.2kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 1.40nM ΔG°: -52.6kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 2.24nM ΔG°: -51.4kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 5.14nM ΔG°: -49.2kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 8.91nM ΔG°: -47.8kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 21.7nM ΔG°: -45.5kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 24.9nM ΔG°: -45.1kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DatapH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DatapH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DatapH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DatapH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DatapH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DatapH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DatapH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair