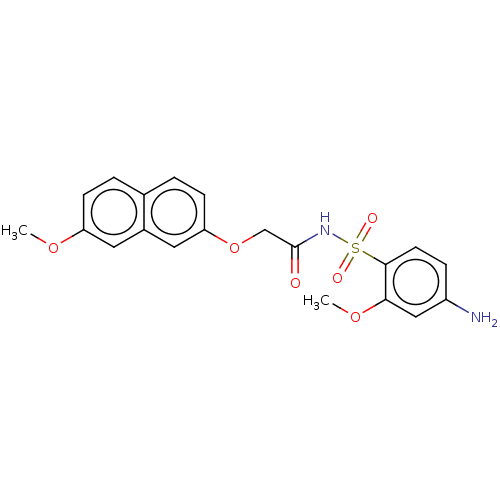
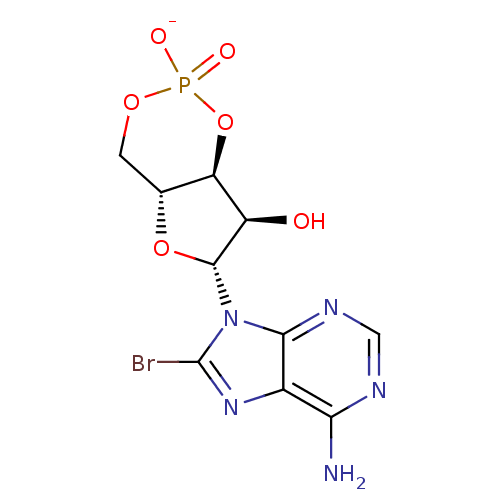
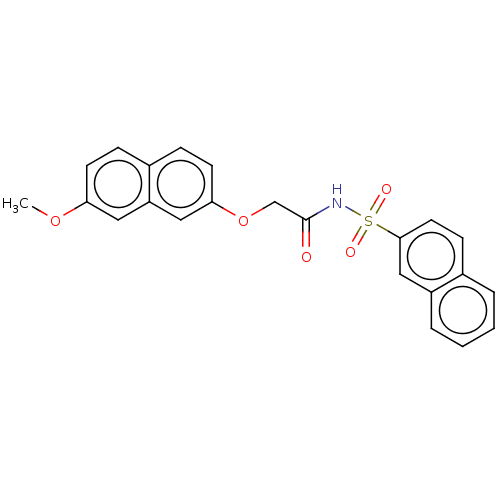
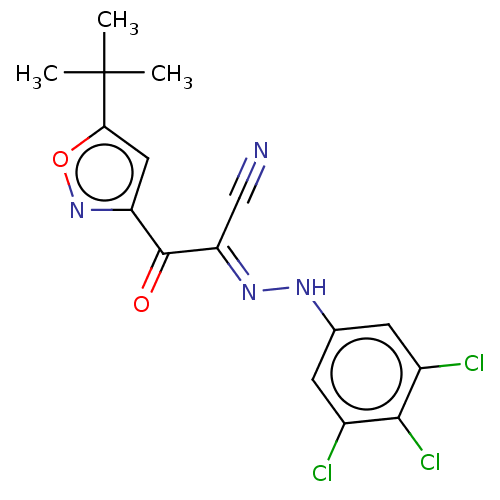
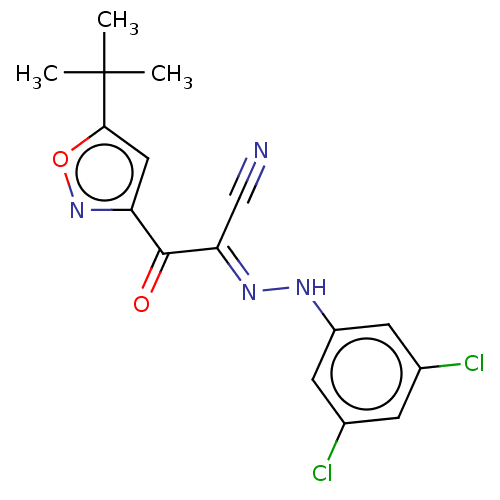
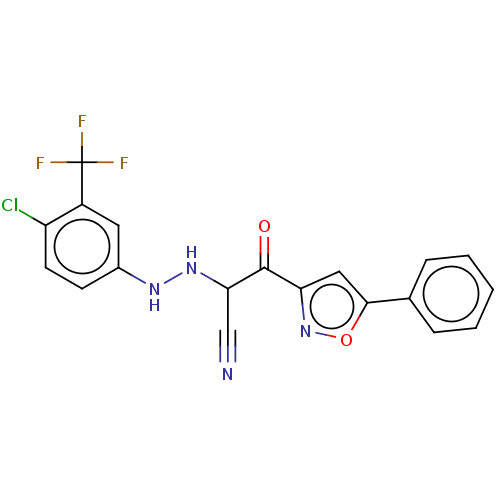
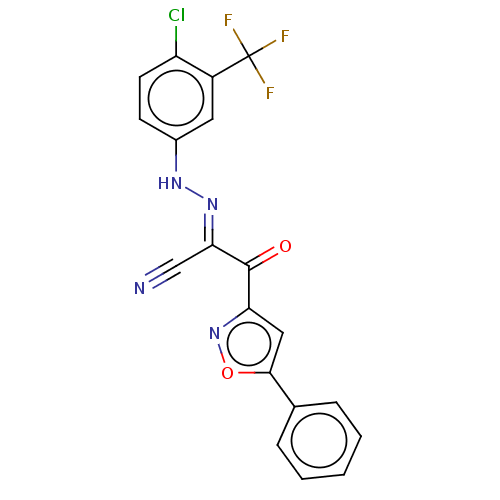
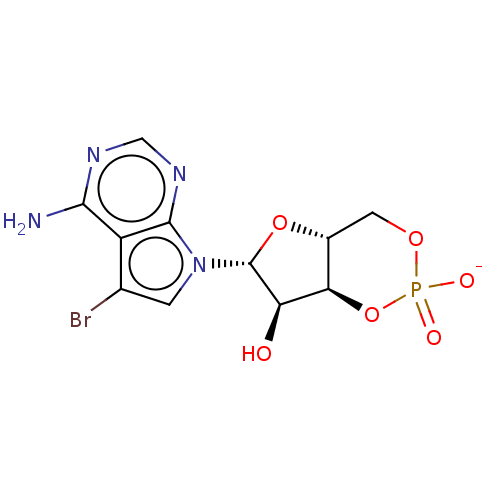
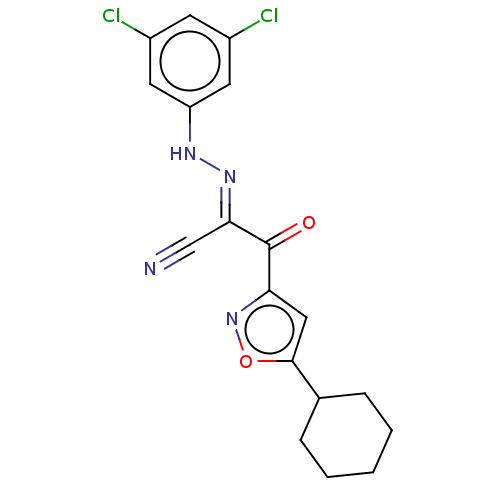
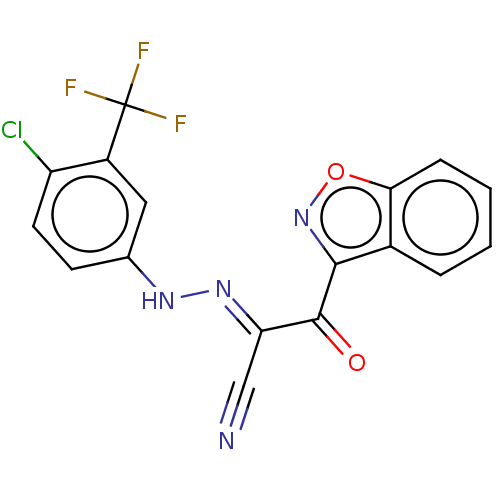
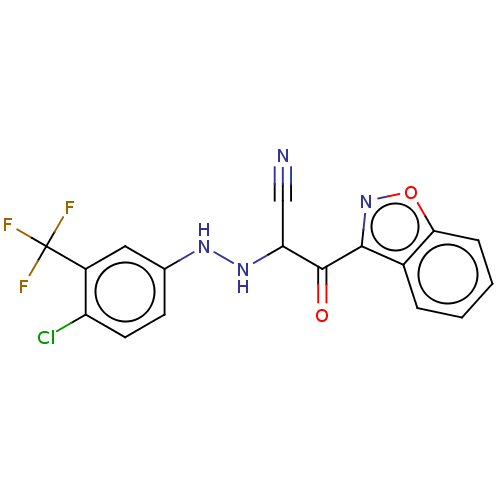
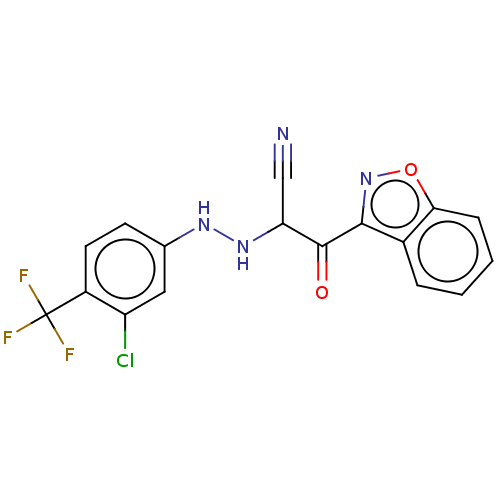
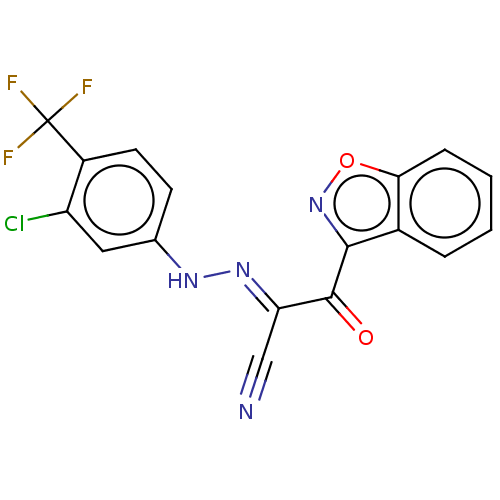
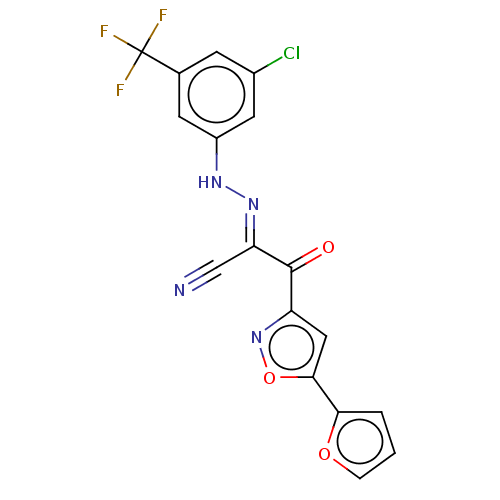
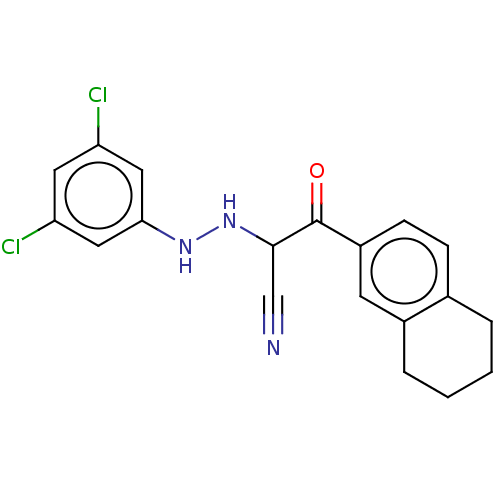
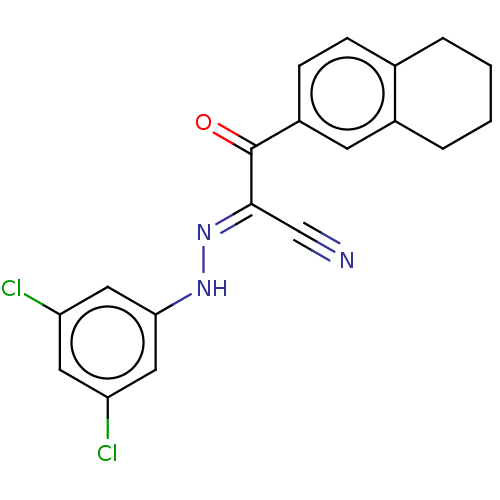
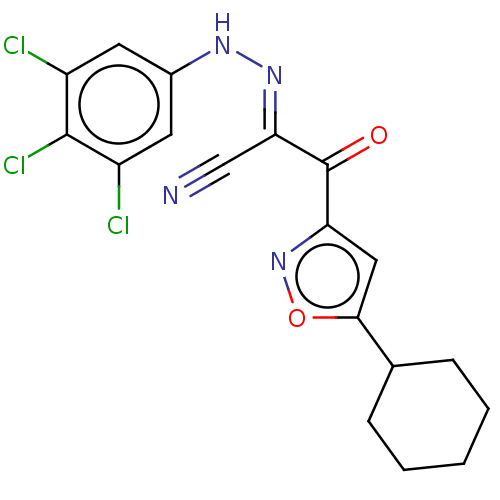
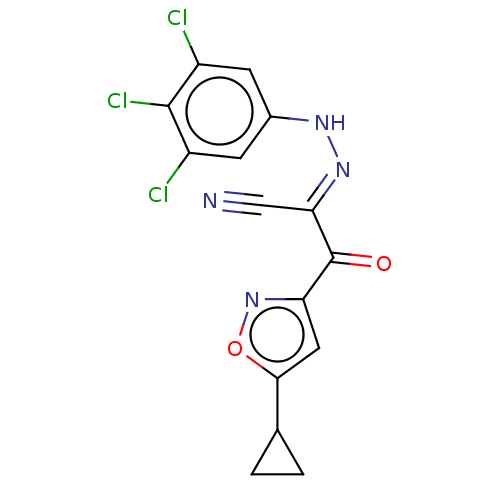
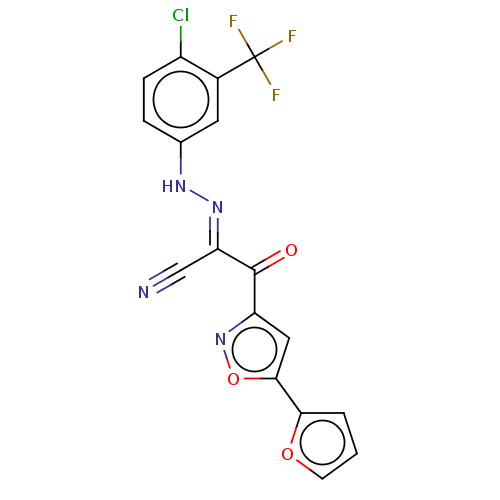
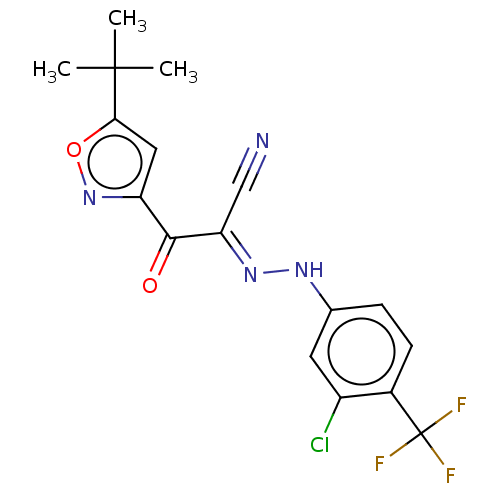
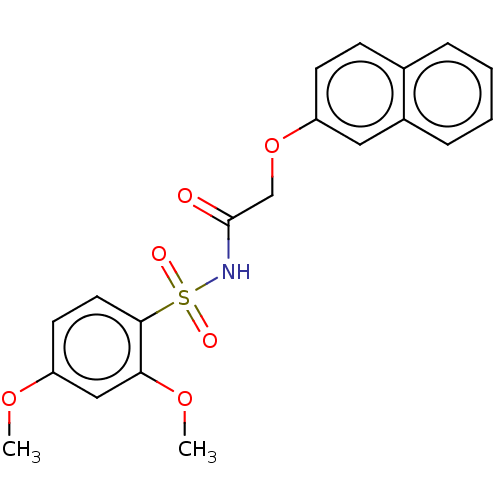
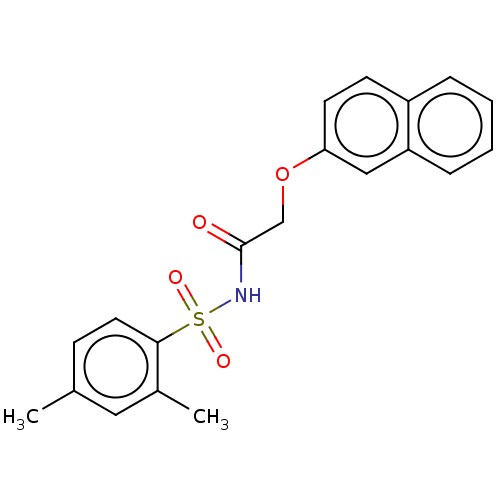
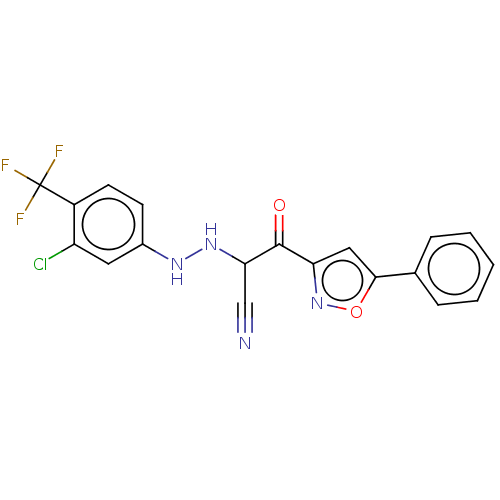
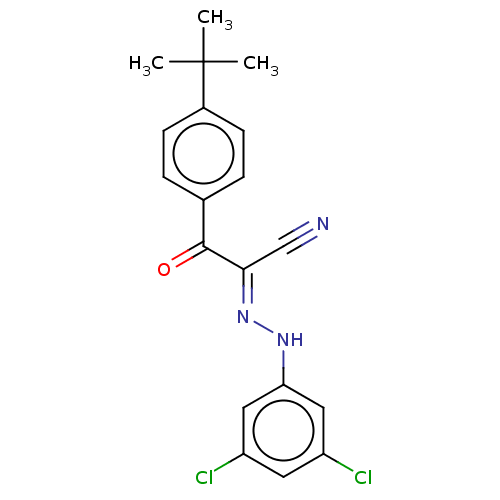
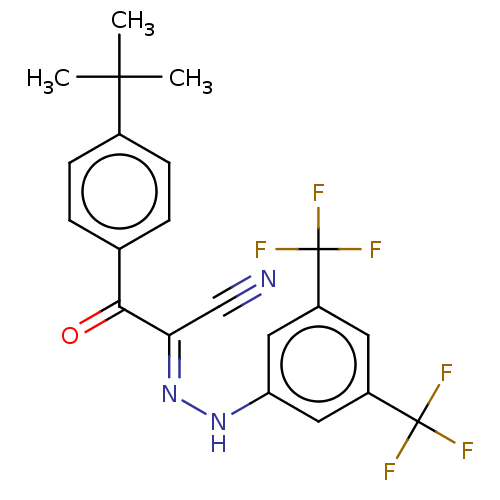
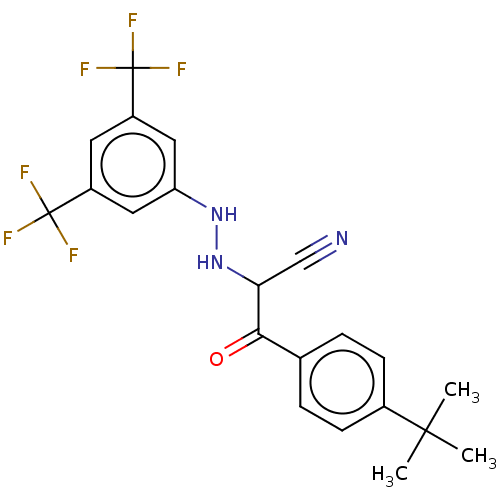
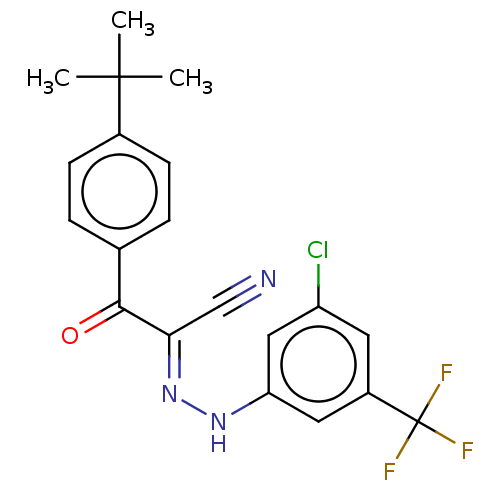
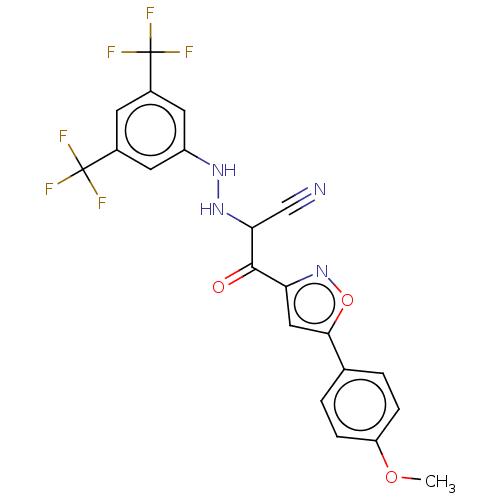
Affinity DataEC50: 530nMAssay Description:Inhibition of 8Fluo-cAMP binding to recombinant human His-tagged EPAC1 (157 to 881 residues) expressed in Escherichia coli measured after 5 mins by f...More data for this Ligand-Target Pair
Affinity DataIC50: 900nMAssay Description:Binding affinity to GST-tagged recombinant EPAC1 CNBD (169 to 318 residues) (unknown origin) expressed in Escherichia coli BL21 Star (DE3) incubated ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.09E+3nMAssay Description:Inhibition of 8Fluo-cAMP binding to recombinant human His-tagged EPAC1 (157 to 881 residues) expressed in Escherichia coli measured after 5 mins by f...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Binding affinity to GST-tagged recombinant EPAC1 CNBD (169 to 318 residues) (unknown origin) expressed in Escherichia coli BL21 Star (DE3) incubated ...More data for this Ligand-Target Pair
Affinity DataEC50: 2.20E+3nMAssay Description:Activation of EPAC1 DEP1 deletion mutant (unknown origin) expressed in mouse NIH-3T3-A14 cells assessed as increase in Rap1b- fluorescent nucleotide ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMAssay Description:Antagonist activity at human recombinant EPAC1 assessed as inhibition of Rap1b-bGDP exchange activityMore data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMAssay Description:Displacement of 8-NBD-cAMP from recombinant human EPAC1 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMAssay Description:Antagonist activity at human recombinant EPAC1 assessed as inhibition of Rap1b-bGDP exchange activityMore data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMAssay Description:Displacement of 8-NBD-cAMP from recombinant human EPAC1 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMAssay Description:To explore the SARs and examine how the modifications on the isoxazole ring affect biological activities of newly synthesized analogues, we first eva...More data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMAssay Description:Antagonist activity at recombinant human full length EPAC1 assessed as inhibition of EPAC1-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide e...More data for this Ligand-Target Pair
Affinity DataEC50: 2.54E+3nMAssay Description:Inhibition of 8Fluo-cAMP binding to recombinant human His-tagged EPAC1 (157 to 881 residues) expressed in Escherichia coli measured after 5 mins by f...More data for this Ligand-Target Pair
Affinity DataIC50: 2.60E+3nMAssay Description:Antagonist activity at human recombinant EPAC1 assessed as inhibition of Rap1b-bGDP exchange activityMore data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+3nMAssay Description:Antagonist activity at recombinant human full length EPAC1 assessed as inhibition of EPAC1-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide e...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+3nMAssay Description:Antagonist activity at recombinant human full length EPAC1 assessed as inhibition of EPAC1-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide e...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+3nMAssay Description:To explore the SARs and examine how the modifications on the isoxazole ring affect biological activities of newly synthesized analogues, we first eva...More data for this Ligand-Target Pair
Affinity DataEC50: 2.75E+3nMAssay Description:Inhibition of 8Fluo-cAMP binding to recombinant human His-tagged EPAC1 (157 to 881 residues) expressed in Escherichia coli measured after 5 mins by f...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:To explore the SARs and examine how the modifications on the isoxazole ring affect biological activities of newly synthesized analogues, we first eva...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Antagonist activity at recombinant human full length EPAC1 assessed as inhibition of EPAC1-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide e...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+3nMAssay Description:Inhibition of EPAC1 (unknown origin) assessed as cAMP-mediated GEF activityMore data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+3nMAssay Description:Inhibition of EPAC1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+3nMAssay Description:Inhibition of recombinant human GST-tagged EPAC1 (149 to 881 residues) expressed in Escherichia coli Rosetta 2(DE3) assessed as reduction in guanine ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+3nMAssay Description:To explore the SARs and examine how the modifications on the isoxazole ring affect biological activities of newly synthesized analogues, we first eva...More data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+3nMAssay Description:Antagonist activity at recombinant human full length EPAC1 assessed as inhibition of EPAC1-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide e...More data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+3nMAssay Description:To explore the SARs and examine how the modifications on the isoxazole ring affect biological activities of newly synthesized analogues, we first eva...More data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+3nMAssay Description:Antagonist activity at recombinant full length EPAC1 (unknown origin) assessed as inhibition of cAMP-mediated Rap1b-BODIPY GDP nucleotide exchange ac...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+3nMAssay Description:Antagonist activity at human recombinant EPAC1 assessed as inhibition of Rap1b-bGDP exchange activityMore data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+3nMAssay Description:Antagonist activity at human recombinant EPAC1 assessed as inhibition of Rap1b-bGDP exchange activityMore data for this Ligand-Target Pair
Affinity DataIC50: 4.20E+3nMAssay Description:To explore the SARs and examine how the modifications on the isoxazole ring affect biological activities of newly synthesized analogues, we first eva...More data for this Ligand-Target Pair
Affinity DataIC50: 4.20E+3nMAssay Description:Antagonist activity at recombinant human full length EPAC1 assessed as inhibition of EPAC1-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide e...More data for this Ligand-Target Pair
Affinity DataIC50: 4.20E+3nMAssay Description:Antagonist activity at recombinant human GST-fused EPAC1 (149 to 881 residues) expressed in Escherichia coli Rosetta (DE3) assessed as inhibition of ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+3nMAssay Description:Antagonist activity at human recombinant EPAC1 assessed as inhibition of Rap1b-bGDP exchange activityMore data for this Ligand-Target Pair
Affinity DataKd: 4.50E+3nMAssay Description:Binding affinity to GST-tagged human EPAC1 CNBD (149 to 318 residues) expressed in Escherichia coli BL21 (DE3) incubated for 30 mins by 8-NBD-cAMP ba...More data for this Ligand-Target Pair
Affinity DataIC50: 4.60E+3nMAssay Description:Antagonist activity at recombinant human full length EPAC1 assessed as inhibition of EPAC1-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide e...More data for this Ligand-Target Pair
Affinity DataIC50: 4.60E+3nMAssay Description:To explore the SARs and examine how the modifications on the isoxazole ring affect biological activities of newly synthesized analogues, we first eva...More data for this Ligand-Target Pair
Affinity DataIC50: 4.60E+3nMAssay Description:Binding affinity to GST-tagged recombinant EPAC1 CNBD (169 to 318 residues) (unknown origin) expressed in Escherichia coli BL21 Star (DE3) incubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.80E+3nMAssay Description:Binding affinity to GST-tagged recombinant EPAC1 CNBD (169 to 318 residues) (unknown origin) expressed in Escherichia coli BL21 Star (DE3) incubated ...More data for this Ligand-Target Pair
Affinity DataKd: 4.90E+3nMAssay Description:Binding affinity to GST-tagged human EPAC1 L273W mutant expressed in Escherichia coli BL21 (DE3) incubated for 30 mins by 8-NBD-cAMP based competitiv...More data for this Ligand-Target Pair
Affinity DataEC50: 5.00E+3nMAssay Description:Activation of recombinant EPAC1 (149 to 881 residues) (unknown origin) assessed as stimulation of dissociation of fluorescent MANT-GDP from recombina...More data for this Ligand-Target Pair
Affinity DataEC50: 5.00E+3nMAssay Description:Activation of recombinant EPAC1 (149 to 881 residues) (unknown origin) assessed as stimulation of dissociation of fluorescent MANT-GDP from recombina...More data for this Ligand-Target Pair
Affinity DataEC50: 5.00E+3nMAssay Description:Activation of recombinant EPAC1 (149 to 881 residues) (unknown origin) assessed as stimulation of dissociation of fluorescent MANT-GDP from recombina...More data for this Ligand-Target Pair
Affinity DataIC50: 5.10E+3nMAssay Description:To explore the SARs and examine how the modifications on the isoxazole ring affect biological activities of newly synthesized analogues, we first eva...More data for this Ligand-Target Pair
Affinity DataIC50: 5.40E+3nMAssay Description:To explore the SARs and examine how the modifications on the isoxazole ring affect biological activities of newly synthesized analogues, we first eva...More data for this Ligand-Target Pair
Affinity DataIC50: 5.40E+3nMAssay Description:Antagonist activity at recombinant full length EPAC1 (unknown origin) assessed as inhibition of cAMP-mediated Rap1b-BODIPY GDP nucleotide exchange ac...More data for this Ligand-Target Pair
Affinity DataIC50: 5.50E+3nMAssay Description:Antagonist activity at recombinant full length EPAC1 (unknown origin) assessed as inhibition of cAMP-mediated Rap1b-BODIPY GDP nucleotide exchange ac...More data for this Ligand-Target Pair
Affinity DataIC50: 5.50E+3nMAssay Description:To explore the SARs and examine how the modifications on the isoxazole ring affect biological activities of newly synthesized analogues, we first eva...More data for this Ligand-Target Pair
Affinity DataIC50: 5.50E+3nMAssay Description:Antagonist activity at recombinant full length EPAC1 (unknown origin) assessed as inhibition of cAMP-mediated Rap1b-BODIPY GDP nucleotide exchange ac...More data for this Ligand-Target Pair
Affinity DataIC50: 5.60E+3nMAssay Description:To explore the SARs and examine how the modifications on the isoxazole ring affect biological activities of newly synthesized analogues, we first eva...More data for this Ligand-Target Pair
Affinity DataIC50: 5.60E+3nMAssay Description:Antagonist activity at recombinant full length EPAC1 (unknown origin) assessed as inhibition of cAMP-mediated Rap1b-BODIPY GDP nucleotide exchange ac...More data for this Ligand-Target Pair
Affinity DataIC50: 5.60E+3nMAssay Description:To explore the SARs and examine how the modifications on the isoxazole ring affect biological activities of newly synthesized analogues, we first eva...More data for this Ligand-Target Pair