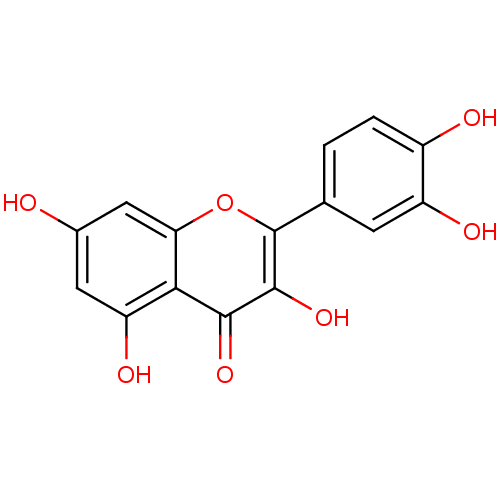
BDBM7460 2-(3,4-dihydroxyphenyl)-3,5,7-trihydroxy-4H-chromen-4-one::2-(3,4-dihydroxyphenyl)-3,5,7-trihydroxy-chromen-4-one::2-(3,4-dihydroxyphenyl)-3,5,7-trihydroxy-chromone;hydrate::CHEMBL50::Quercetin::Quercetin (10)::Quercetin (21)::Quercetin (Qur)::US11021454, Compound Quercetin::US9180183, Quercetin::med.21724, Compound 4
SMILES Oc1cc(O)c2c(c1)oc(-c1ccc(O)c(O)c1)c(O)c2=O
InChI Key InChIKey=REFJWTPEDVJJIY-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 420 hits for monomerid = 7460
Found 420 hits for monomerid = 7460
TargetEnoyl-acyl-carrier protein reductase(malaria parasite P. falciparum)
National Institute of Immunology
Curated by ChEMBL
National Institute of Immunology
Curated by ChEMBL
Affinity DataKi: 22nMAssay Description:Inhibition of Plasmodium falciparum ENR in presence of triclosanMore data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Inhibition of CYP1B1 EROD activity assessed as inhibition of deethylation of 7-ethoxyresorufin to resorufinMore data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Inhibition of CYP1B1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 45.3nMAssay Description:Inhibition of rat ecto-5'-nucleotidase expressed in Sf9 cells by capillary electrophoresis methodMore data for this Ligand-Target Pair
Affinity DataKi: 280nMAssay Description:Inhibition of xanthine oxidase (unknown origin) at 37 degC at ph 7..8More data for this Ligand-Target Pair
Affinity DataKi: 472nMAssay Description:Inhibition of YES (unknown origin)More data for this Ligand-Target Pair
TargetEnoyl-acyl-carrier protein reductase(malaria parasite P. falciparum)
National Institute of Immunology
Curated by ChEMBL
National Institute of Immunology
Curated by ChEMBL
Affinity DataKi: 473nMAssay Description:Inhibition of Plasmodium falciparum ENR using crotonyl-CoA substrateMore data for this Ligand-Target Pair
Affinity DataKi: 660nMAssay Description:Inhibition of CYP1A1 EROD activity assessed as inhibition of deethylation of 7-ethoxyresorufin to resorufinMore data for this Ligand-Target Pair
TargetEnoyl-acyl-carrier protein reductase(malaria parasite P. falciparum)
National Institute of Immunology
Curated by ChEMBL
National Institute of Immunology
Curated by ChEMBL
Affinity DataKi: 1.09E+3nMAssay Description:Inhibition of Plasmodium falciparum ENR using NADH substrateMore data for this Ligand-Target Pair
Affinity DataKi: 1.18E+3nM ΔG°: -8.00kcal/molepH: 7.5 T: 22°CAssay Description:In vitro kinase assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of 100 uM ATP/ [gam...More data for this Ligand-Target Pair
Affinity DataKi: 1.18E+3nMAssay Description:Inhibition of CK2 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 1.18E+3nMAssay Description:Inhibition of CK2alpha (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+3nMAssay Description:Inhibition of bovine xanthine oxidaseMore data for this Ligand-Target Pair
TargetMultidrug resistance-associated protein 1(Human)
The Hong Kong Polytechnic University
Curated by ChEMBL
The Hong Kong Polytechnic University
Curated by ChEMBL
Affinity DataKi: 2.40E+3nMAssay Description:Inhibition of MRP1 transfected in human HeLa cells assessed as inhibition of [3H]LTC4 transport by rapid filtration assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.40E+3nMpH: 7.4Assay Description:Carbonic anhydrase activity was assayed by following the change in absorbance at 348 nm of 4-nitrophenylacetate (NPA) to 4-nitrophenylate ion over a ...More data for this Ligand-Target Pair
Affinity DataKi: 2.46E+3nMAssay Description:Ability to displace [3H]N6-phenylisopropyladenosine binding from adenosine A1 receptor.Checked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 2.47E+3nMAssay Description:Displacement of specific [3H]-PIA binding from adenosine A1 receptor in rat brain membranes.More data for this Ligand-Target Pair
Affinity DataKi: 2.54E+3nMAssay Description:Inhibition of human CA2 by stopped-flow CO2 hydration assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.68E+3nMAssay Description:Inhibition of human CA1 by stopped-flow CO2 hydration assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.60E+3nMpH: 7.4Assay Description:Carbonic anhydrase activity was assayed by following the change in absorbance at 348 nm of 4-nitrophenylacetate (NPA) to 4-nitrophenylate ion over a ...More data for this Ligand-Target Pair
Affinity DataKi: 3.73E+3nMpH: 7.4Assay Description:CA activity was assayed by following the change in absorbance at 348 nm of 4-NPA to 4-nitrophenylate ion over a period of 3 min at 25�C using a spect...More data for this Ligand-Target Pair
Affinity DataKi: 4.30E+3nMAssay Description:Competitive inhibition of Helicobacter pylori Ddl using ATP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: 4.84E+3nMAssay Description:Inhibition of human CA7 by stopped-flow CO2 hydration assayMore data for this Ligand-Target Pair
Affinity DataKi: 5.41E+3nMAssay Description:Inhibition of human CA14 by stopped-flow CO2 hydration assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.17E+3nMAssay Description:Inhibition of human CA6 by stopped-flow CO2 hydration assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.81E+3nMAssay Description:Inhibition of human CA5A by stopped-flow CO2 hydration assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.92E+3nMAssay Description:Ability to displace [3H]-CGS- 21680 binding from adenosine A2A receptor.Checked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 6.99E+3nMAssay Description:Affinity at Adenosine A2A receptor in rat striatal membranes by [3H]- CGS 21680 displacement.More data for this Ligand-Target Pair
Affinity DataKi: 7.00E+3nMAssay Description:Inhibition of human CA9 by stopped-flow CO2 hydration assayMore data for this Ligand-Target Pair
Affinity DataKi: 7.40E+3nMAssay Description:This is a review article. Please point to the original journal.More data for this Ligand-Target Pair
Affinity DataKi: 7.85E+3nMAssay Description:Binding affinity to dopamine D4 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 7.89E+3nMAssay Description:Inhibition of human CA4 by stopped-flow CO2 hydration assayMore data for this Ligand-Target Pair
Affinity DataKi: 8.10E+3nMAssay Description:Inhibition of human CA3 by stopped-flow CO2 hydration assayMore data for this Ligand-Target Pair
TargetMultidrug resistance-associated protein 1(Human)
The Hong Kong Polytechnic University
Curated by ChEMBL
The Hong Kong Polytechnic University
Curated by ChEMBL
Affinity DataKi: 8.10E+3nMAssay Description:TP_TRANSPORTER: inhibition of LTC4 uptake in membrane vesicle from MRP1-expressing HeLa cellsMore data for this Ligand-Target Pair
Affinity DataKi: 9.03E+3nMAssay Description:Inhibition of mouse CA13 by stopped-flow CO2 hydration assayMore data for this Ligand-Target Pair
Affinity DataKi: 9.10E+3nMpH: 7.4Assay Description:Carbonic anhydrase activity was assayed by following the change in absorbance at 348 nm of 4-nitrophenylacetate (NPA) to 4-nitrophenylate ion over a ...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Mazandaran University of Medical Sciences
Curated by ChEMBL
Mazandaran University of Medical Sciences
Curated by ChEMBL
Affinity DataKi: 9.34E+3nMAssay Description:Inhibition of Helicobacter pylori urease assessed as measuring ammonia production incubated for 1.5 hr by indophenol methodMore data for this Ligand-Target Pair
Affinity DataKi: 9.39E+3nMAssay Description:Inhibition of human CA12 by stopped-flow CO2 hydration assayMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Bovine)
WestfÄLische Wilhelms-UniversitÄ
Curated by PDSP Ki Database
WestfÄLische Wilhelms-UniversitÄ
Curated by PDSP Ki Database
TargetGamma-aminobutyric acid receptor subunit alpha-1(Rat)
WestfÄLische Wilhelms-UniversitÄ
Curated by PDSP Ki Database
WestfÄLische Wilhelms-UniversitÄ
Curated by PDSP Ki Database
TargetGamma-aminobutyric acid receptor subunit alpha-1(Rat)
WestfÄLische Wilhelms-UniversitÄ
Curated by PDSP Ki Database
WestfÄLische Wilhelms-UniversitÄ
Curated by PDSP Ki Database
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
WestfÄLische Wilhelms-UniversitÄ
Curated by PDSP Ki Database
WestfÄLische Wilhelms-UniversitÄ
Curated by PDSP Ki Database
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
WestfÄLische Wilhelms-UniversitÄ
Curated by PDSP Ki Database
WestfÄLische Wilhelms-UniversitÄ
Curated by PDSP Ki Database
TargetSodium-dependent noradrenaline transporter(Human)
WestfÄLische Wilhelms-UniversitÄ
Curated by PDSP Ki Database
WestfÄLische Wilhelms-UniversitÄ
Curated by PDSP Ki Database
Activity Spreadsheet -- ITC Data from BindingDB
 Found 4 hits for monomerid = 7460
Found 4 hits for monomerid = 7460
ITC DataΔG°: -9.78kcal/mole −TΔS°: -0.257kcal/mole ΔH°: -9.58kcal/mole logk: 3.93E+7
pH: 7.5 T: 10.00°C
pH: 7.5 T: 10.00°C
ITC DataΔG°: -8.08kcal/mole −TΔS°: 0.840kcal/mole ΔH°: -8.92kcal/mole logk: 6.82E+5
pH: 7.0 T: 30.00°C
pH: 7.0 T: 30.00°C
ITC DataΔG°: -7.40kcal/mole −TΔS°: -2.66kcal/mole ΔH°: -4.74kcal/mole logk: 2.20E+5
pH: 7.0 T: 30.00°C
pH: 7.0 T: 30.00°C
ITC DataΔG°: -7.94kcal/mole −TΔS°: -2.82kcal/mole ΔH°: -5.12kcal/mole logk: 5.42E+5
pH: 7.0 T: 30.00°C
pH: 7.0 T: 30.00°C
 3D Structure (crystal)
3D Structure (crystal)