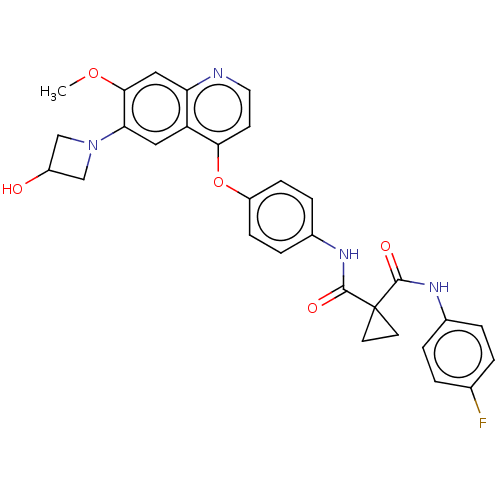
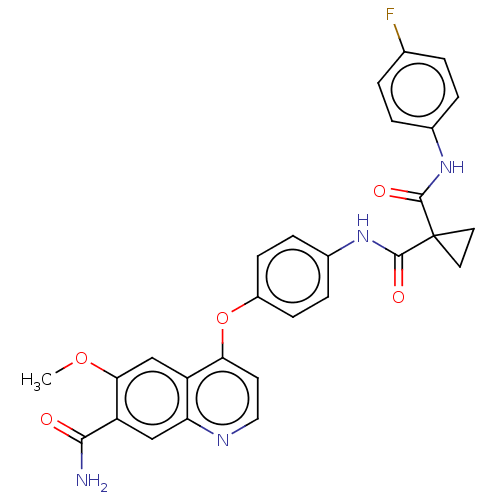
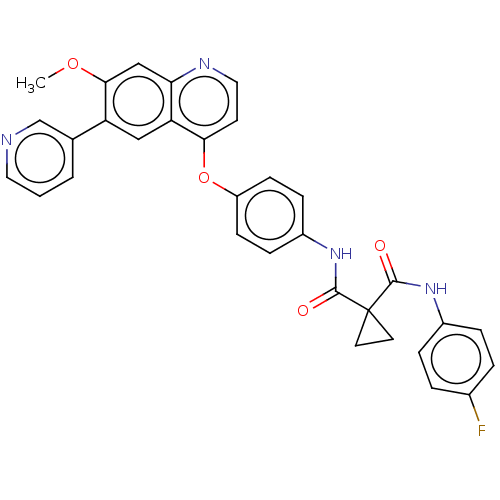
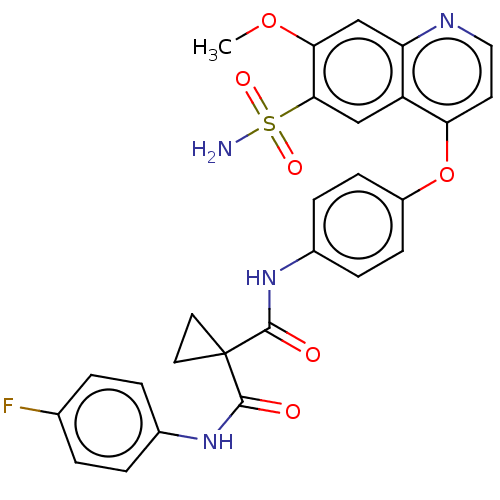
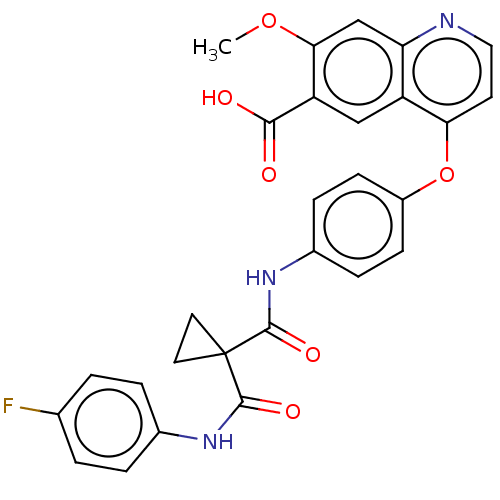
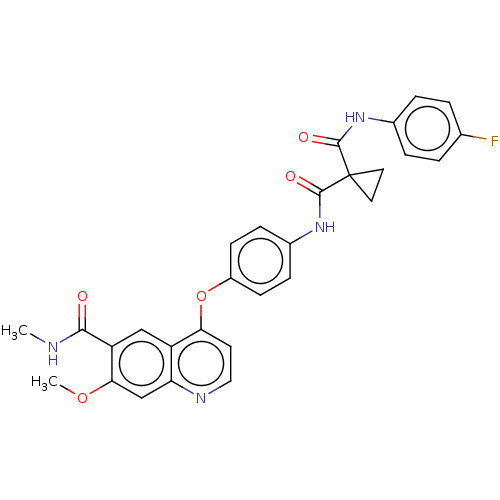
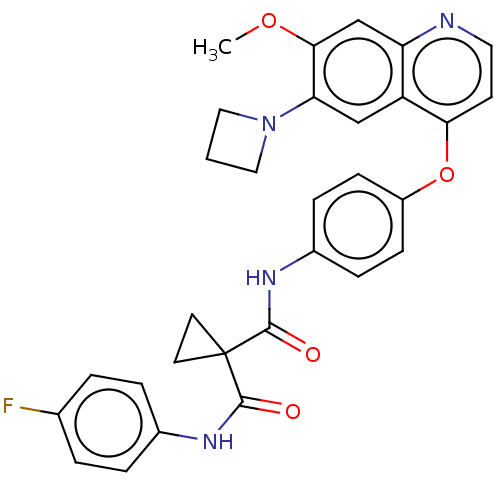
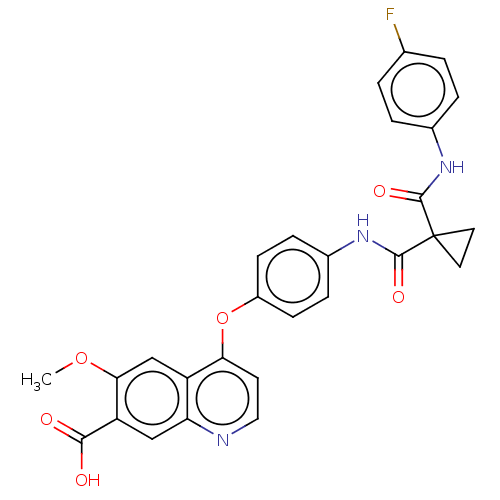
Report error Found 65 Enz. Inhib. hit(s) with Target = 'Hepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L]'
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
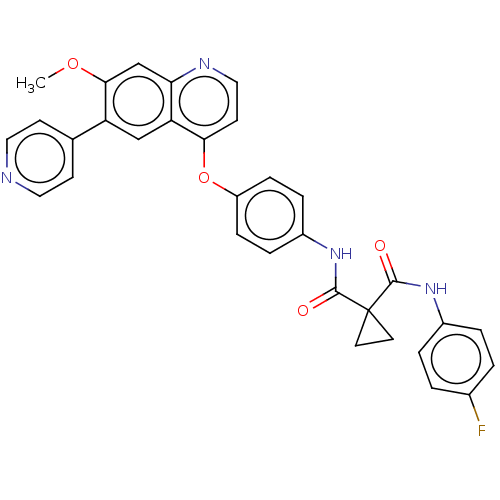
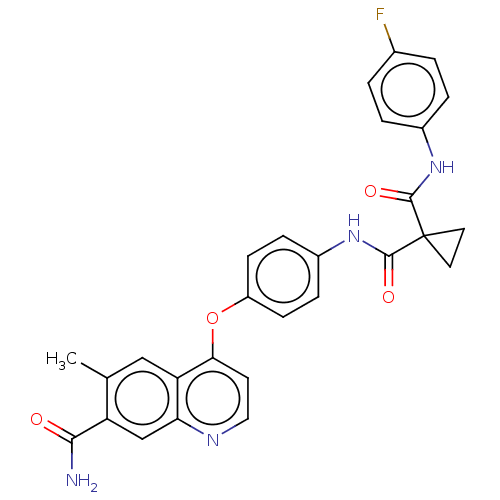
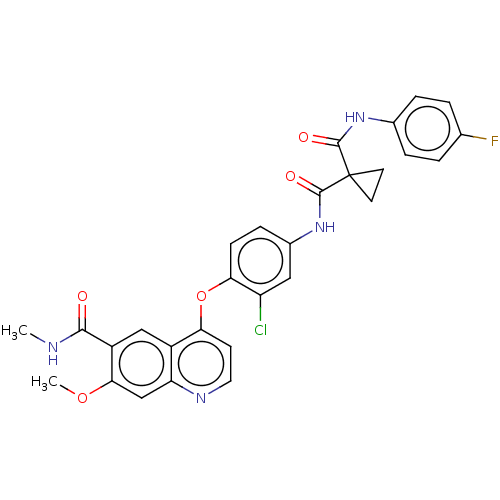
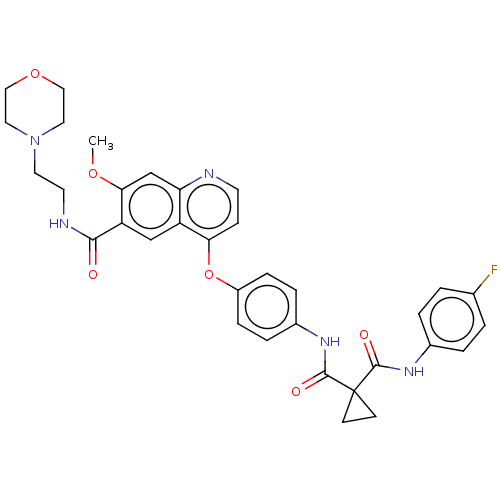
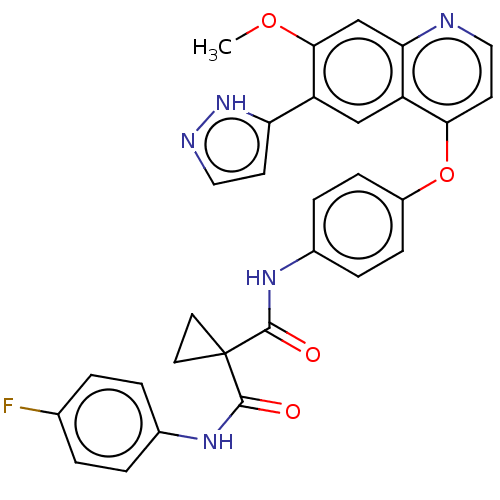
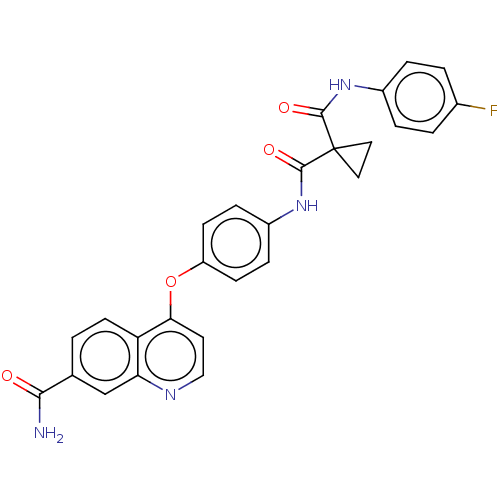
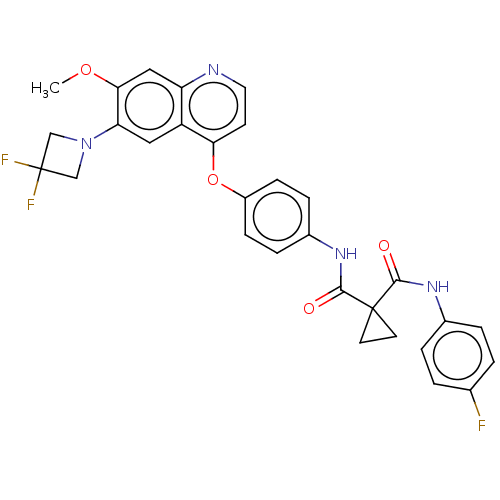
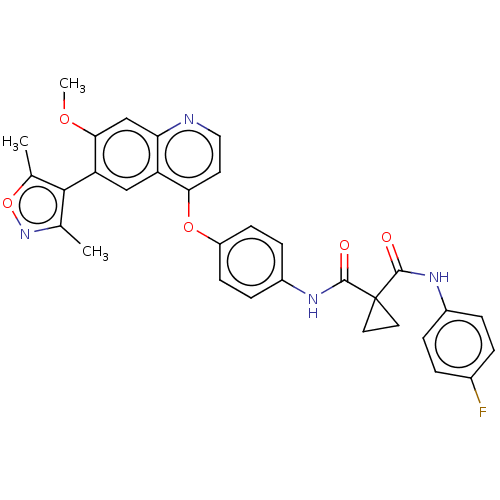
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
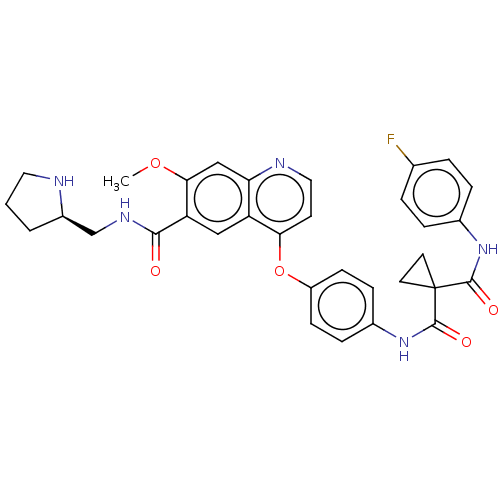
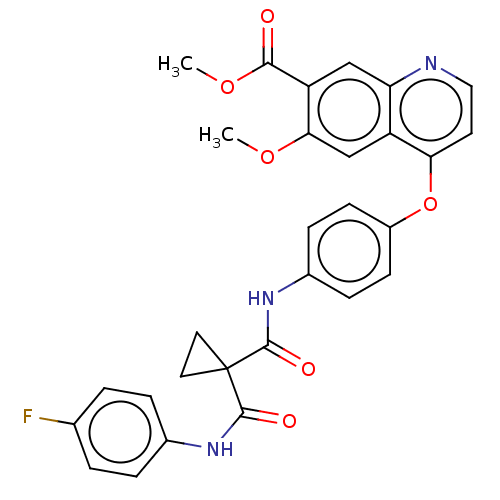
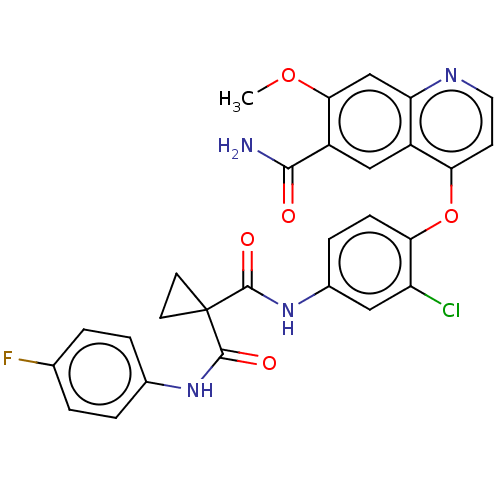
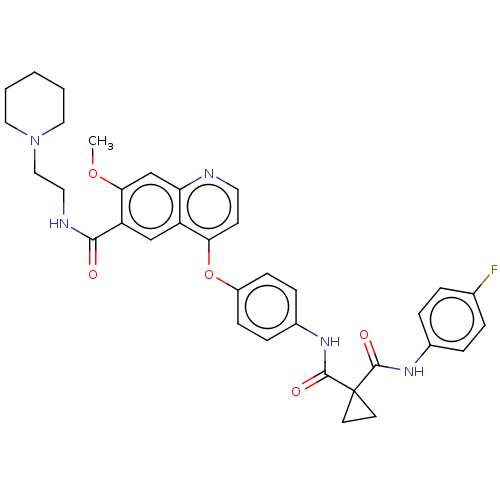
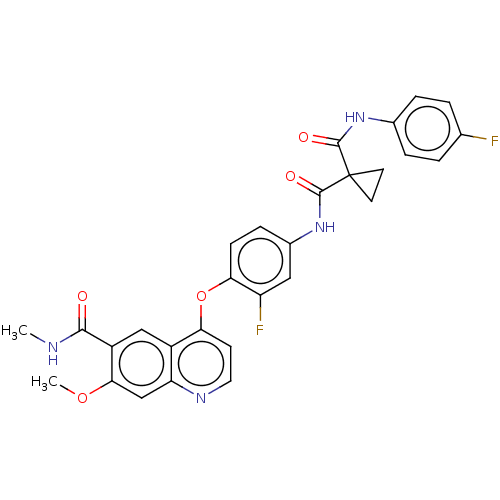
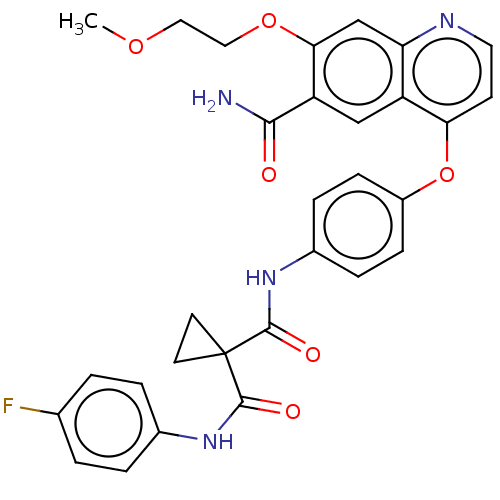
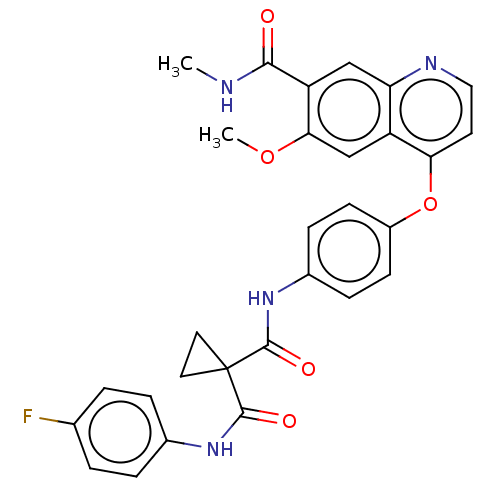
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
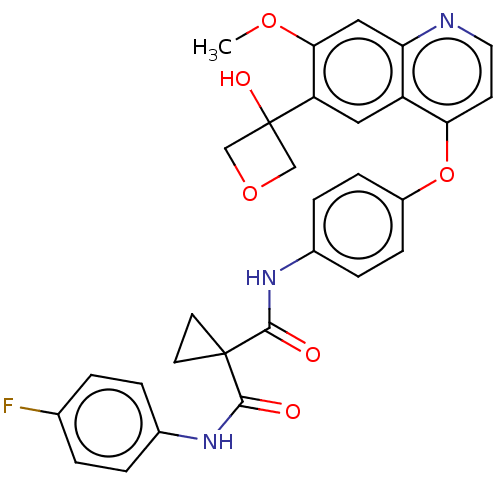
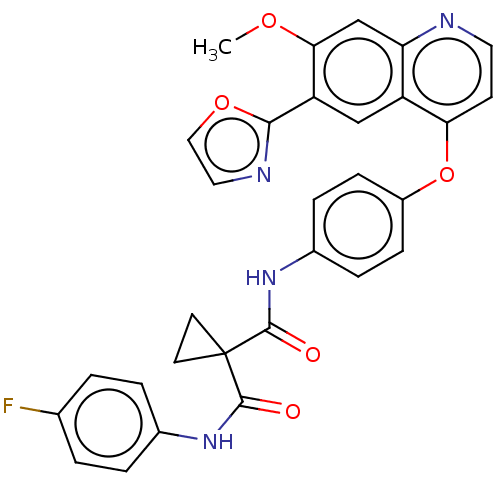
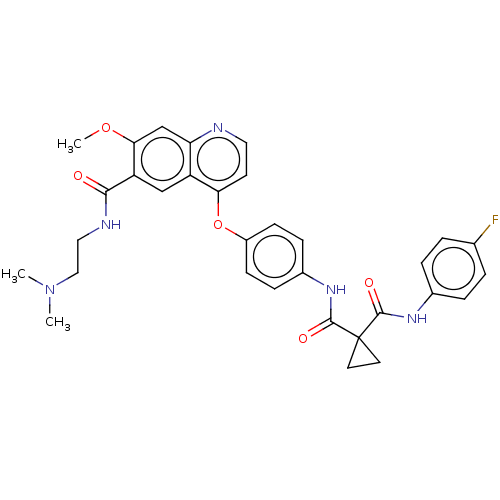
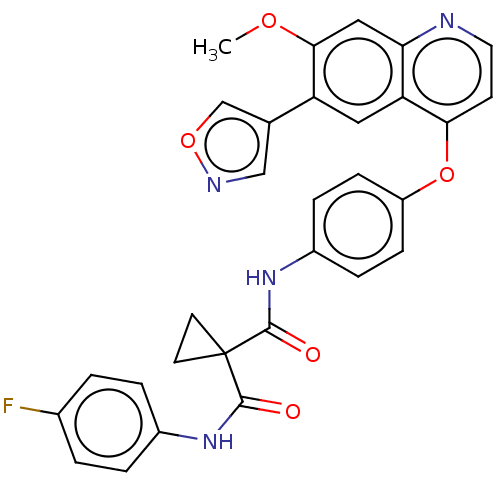
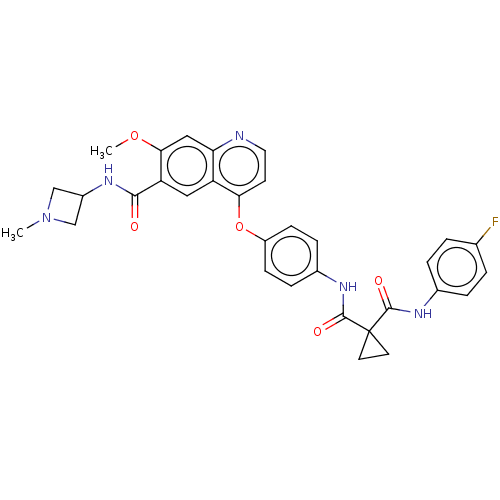
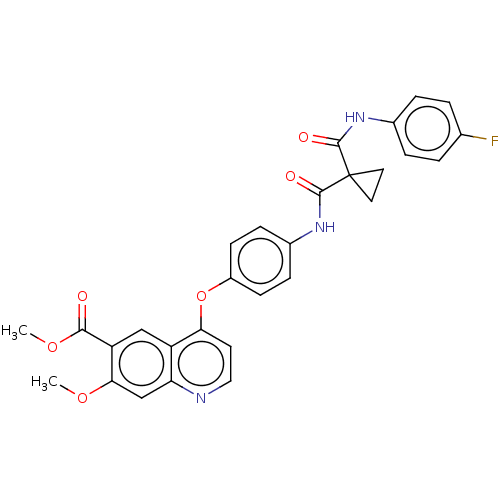
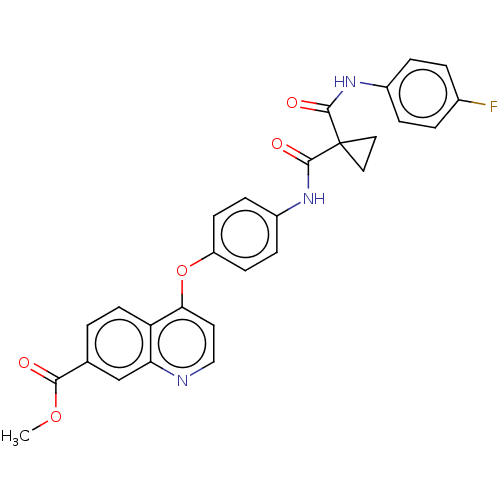
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
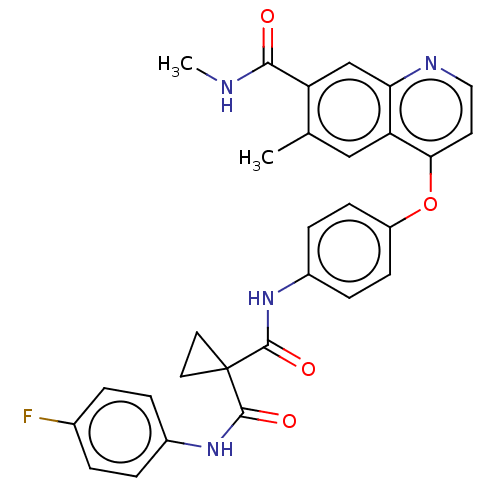
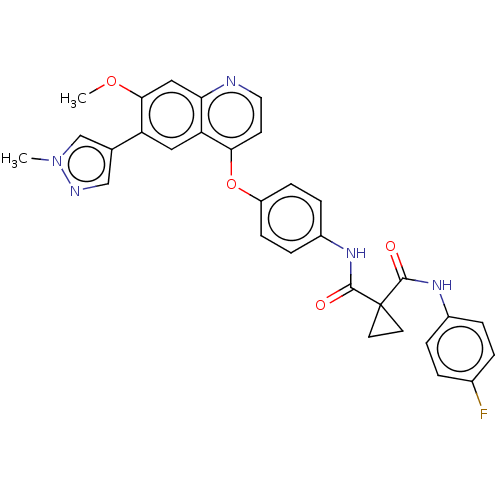
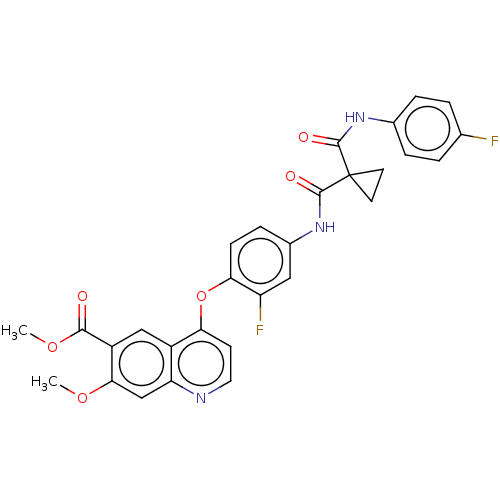
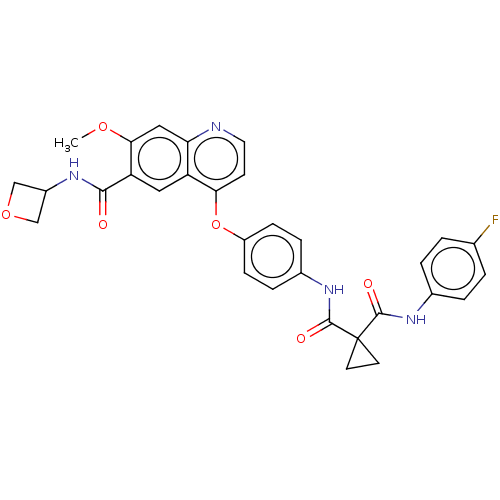
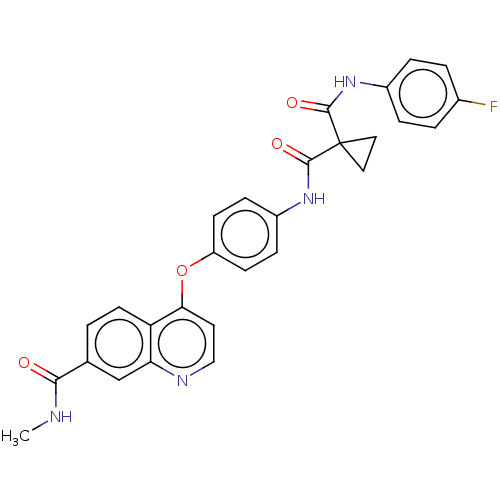
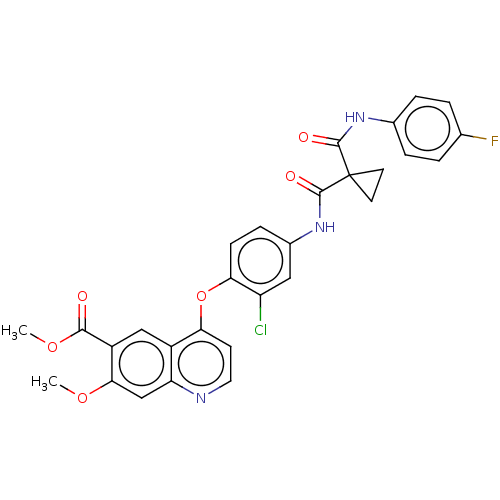
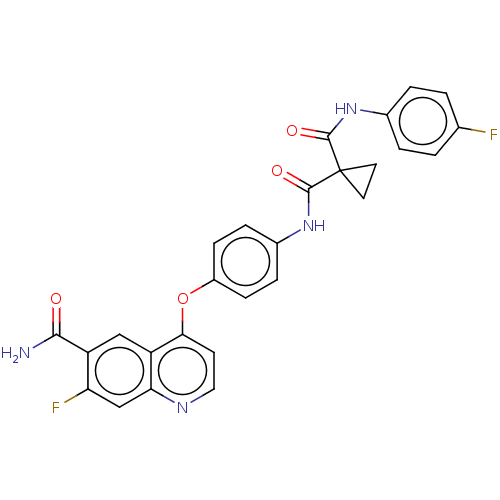
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 10nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 55nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 55nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 55nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 55nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 55nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 55nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 55nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 55nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetHepatocyte growth factor receptor MET Isoform 2 [974-1390,A1209G,V1290L](Human)
Exelixis
US Patent
Exelixis
US Patent
Affinity DataIC50: 55nMAssay Description:Human Met (residues R974-S1390 with A1209G and V1290L, 3.4 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 250 μM KKKGQEEEYVFIE (SEO ID NO...More data for this Ligand-Target Pair
Ligand InfoSimilars