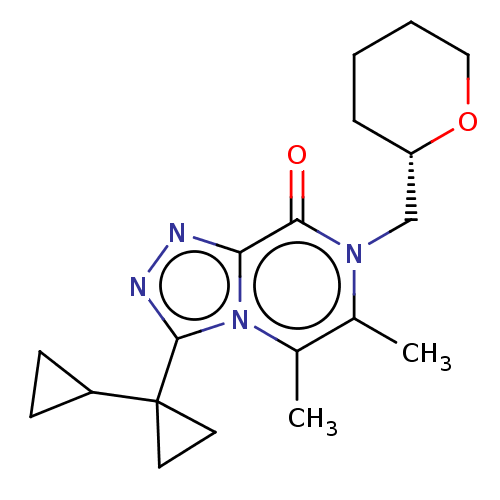
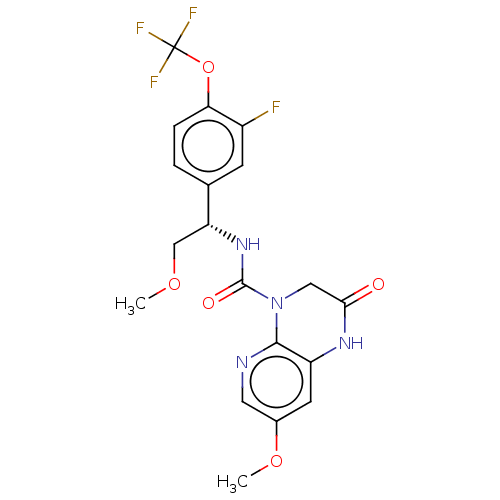
Report error Found 43 Enz. Inhib. hit(s) with Target = 'Rod cGMP-specific 3',5'-cyclic phosphodiesterase subunit alpha/beta'
Affinity DataIC50: 611nMAssay Description:The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of human PDE6...More data for this Ligand-Target Pair
Affinity DataIC50: 982nMAssay Description:The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of human PDE6...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of human PDE6...More data for this Ligand-Target Pair
Affinity DataIC50: 2.66E+3nMAssay Description:Table 7: The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of h...More data for this Ligand-Target Pair
Affinity DataIC50: 2.99E+3nMAssay Description:The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of human PDE6...More data for this Ligand-Target Pair
Affinity DataIC50: 3.06E+3nMAssay Description:The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of human PDE6...More data for this Ligand-Target Pair
Affinity DataIC50: 3.29E+3nMAssay Description:The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of human PDE6...More data for this Ligand-Target Pair
Affinity DataIC50: 3.45E+3nMAssay Description:The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of human PDE6...More data for this Ligand-Target Pair
Affinity DataIC50: 3.85E+3nMAssay Description:The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of human PDE6...More data for this Ligand-Target Pair
Affinity DataIC50: 7.62E+3nMAssay Description:Table 7: The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of h...More data for this Ligand-Target Pair
Affinity DataIC50: 9.59E+3nMAssay Description:The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of human PDE6...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of recombinant human PDE6A/PDE6B expressed in insect cells using [3H]cGMP as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human PDE6ABMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of recombinant human PDE6A/PDE6B expressed in baculovirus infected Sf9 cell expression system using [3H]cGMP as substrate preincubated for...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Table 7: The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of h...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Table 7: The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of h...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Table 7: The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of h...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Table 7: The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of h...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:All 3′, 5′ cyclic nucleotide phosphodiesterase (PDE) enzyme activities are measured with a radiometric enzyme assay based on SPA detectio...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:All 3′, 5′ cyclic nucleotide phosphodiesterase (PDE) enzyme activities are measured with a radiometric enzyme assay based on SPA detectio...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:All 3′, 5′ cyclic nucleotide phosphodiesterase (PDE) enzyme activities are measured with a radiometric enzyme assay based on SPA detectio...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:All 3′, 5′ cyclic nucleotide phosphodiesterase (PDE) enzyme activities are measured with a radiometric enzyme assay based on SPA detectio...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:All 3′, 5′ cyclic nucleotide phosphodiesterase (PDE) enzyme activities are measured with a radiometric enzyme assay based on SPA detectio...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:All 3′, 5′ cyclic nucleotide phosphodiesterase (PDE) enzyme activities are measured with a radiometric enzyme assay based on SPA detectio...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:All 3′, 5′ cyclic nucleotide phosphodiesterase (PDE) enzyme activities are measured with a radiometric enzyme assay based on SPA detectio...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:All 3′, 5′ cyclic nucleotide phosphodiesterase (PDE) enzyme activities are measured with a radiometric enzyme assay based on SPA detectio...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Table 7: The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of h...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Table 7: The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of h...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Table 7: The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of h...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Table 7: The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of h...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Table 7: The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of h...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of human PDE6...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Table 7: The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of h...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of human PDE6...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Table 7: The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of h...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of human PDE6...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Table 7: The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of h...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of human PDE6...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of human PDE6...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Table 7: The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of h...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of human PDE6...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Table 7: The following phosphodiesterase activities are measured using [3H]cGMP as reaction substrate: human PDE6A/6B. The catalytic active form of h...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of recombinant human PDE6A/PDE6B expressed in insect cells using [3H]cGMP as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair