Report error Found 158 Enz. Inhib. hit(s) with all data for entry = 10461
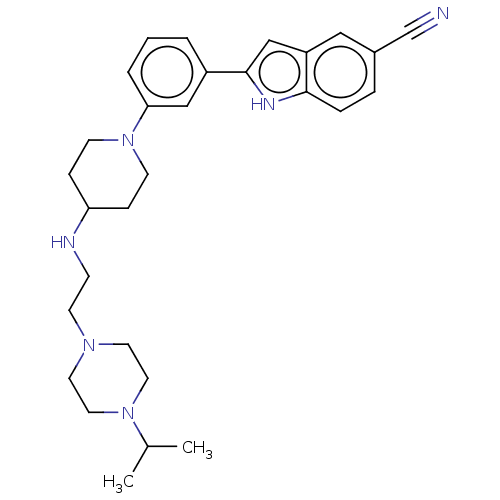
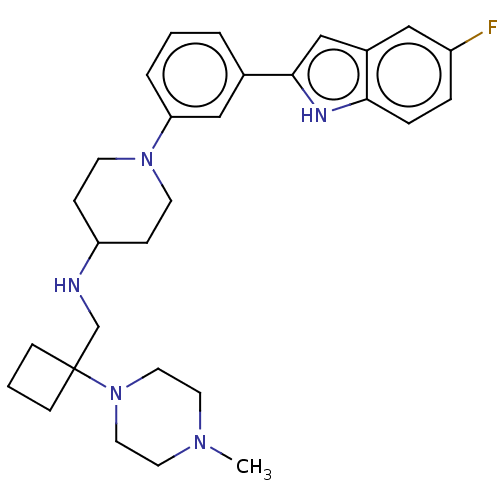
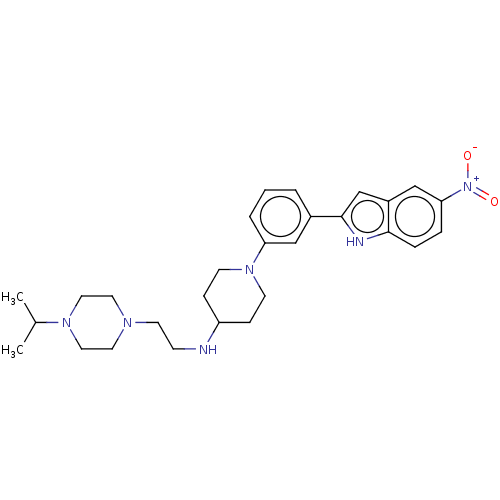
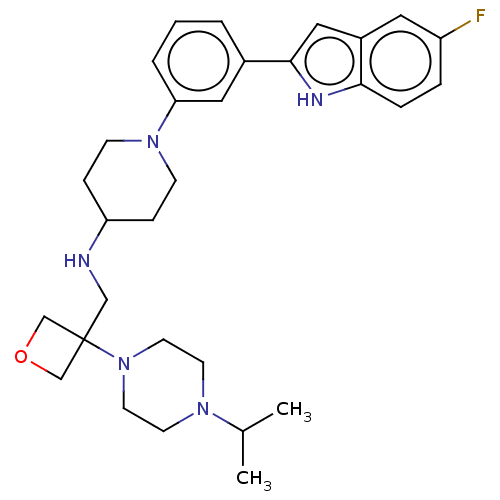
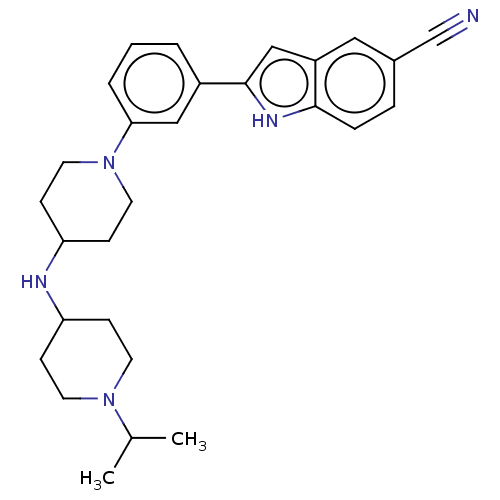
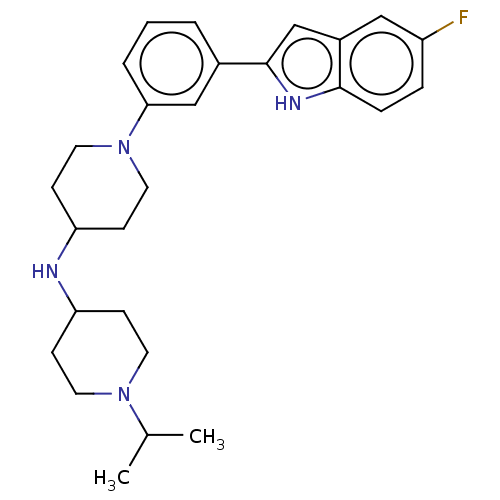
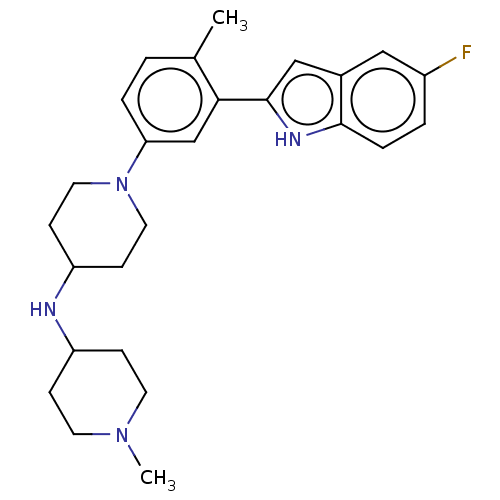
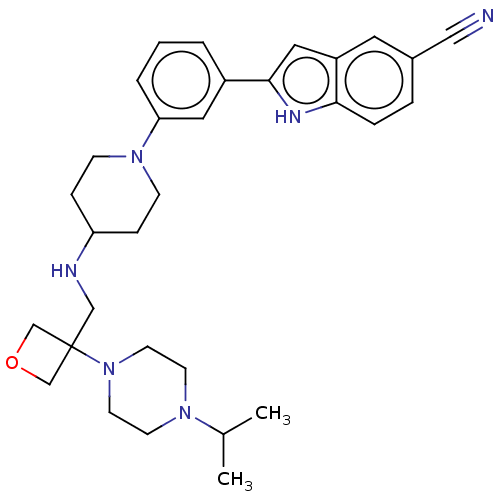
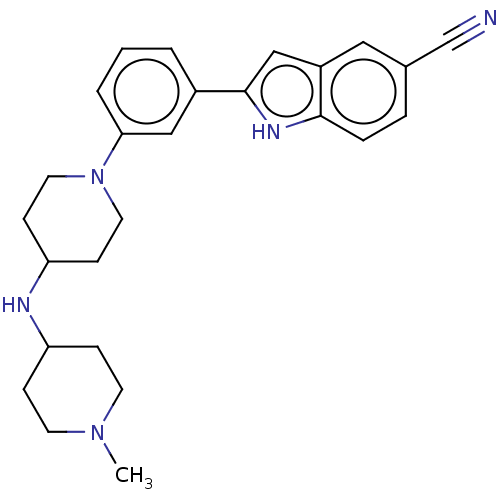
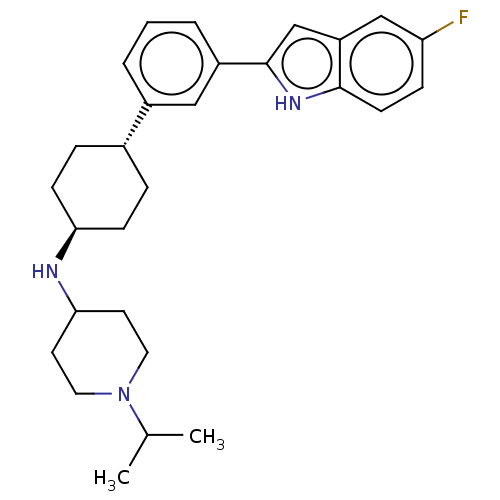
Affinity DataIC50: 15nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
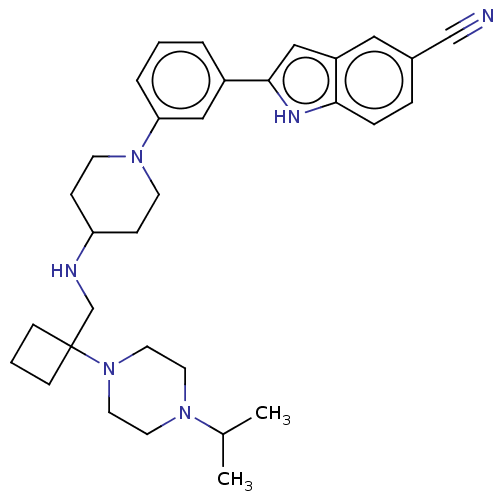
Affinity DataIC50: 25nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
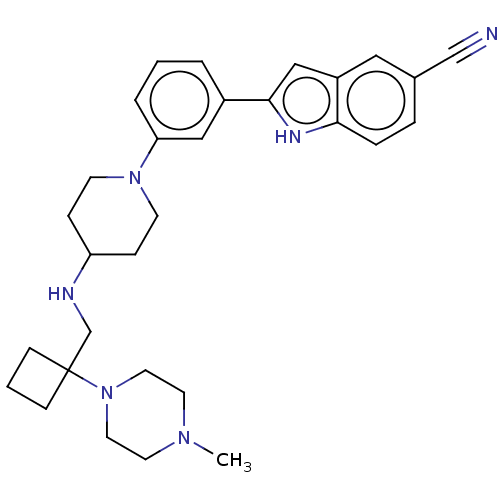
Affinity DataIC50: 26nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
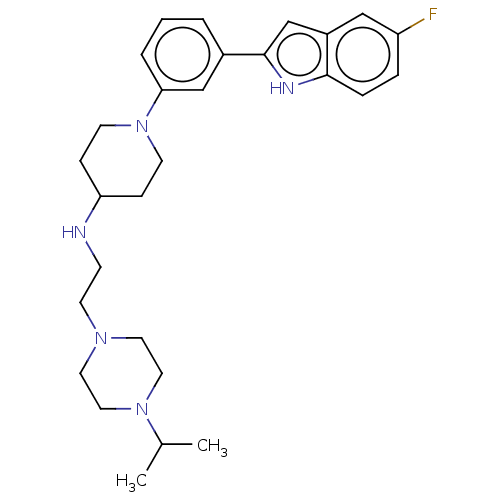
Affinity DataIC50: 27nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 36nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 41nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 47nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 49nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 82nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 86nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 87nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 92nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 140nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 140nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 140nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 140nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 175nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 180nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 190nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 210nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 260nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 290nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 290nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 310nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair
Affinity DataIC50: 320nMAssay Description:To optimize p97 inhibitors, the C-5 trifluoromethylated trifluoromethylated indole 12 was generated as a promising lead structure. In the ADP-Glo ass...More data for this Ligand-Target Pair