Report error Found 28 Enz. Inhib. hit(s) with Target = 'Bifunctional ligase/repressor BirA'
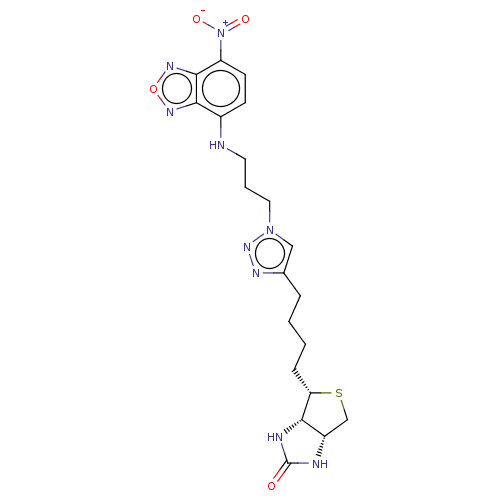
TargetBifunctional ligase/repressor BirA(Escherichia coli (strain K12))
University of Nebraska-Lincoln
Curated by ChEMBL
University of Nebraska-Lincoln
Curated by ChEMBL
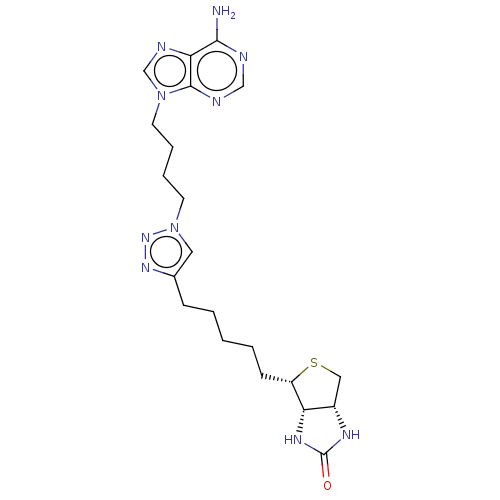
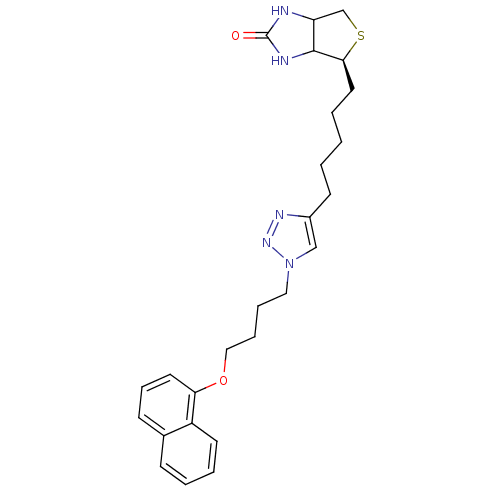
Affinity DataKd: 1.5nMAssay Description:Binding affinity to Escherichia coli biotin protein ligaseMore data for this Ligand-Target Pair
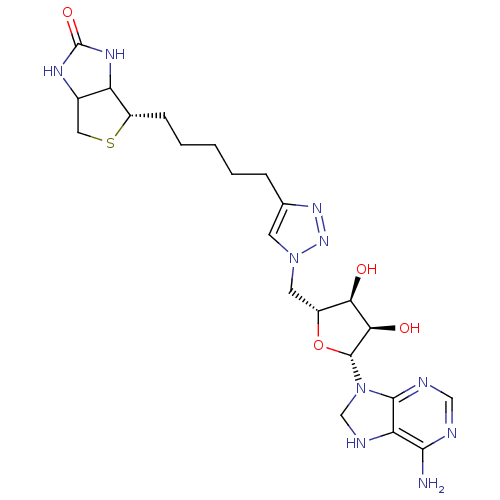
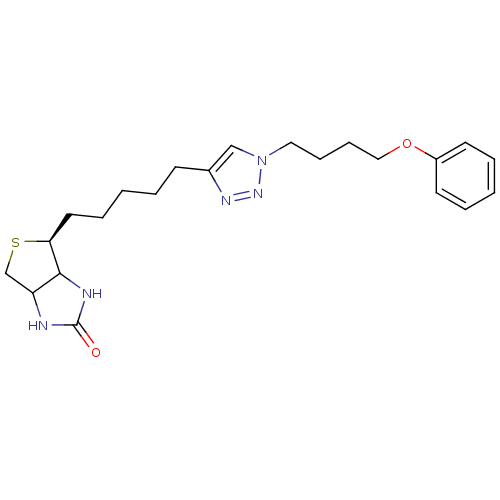
Affinity DataKi: 30nM ΔG°: -44.7kJ/mole IC50: 203nMpH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
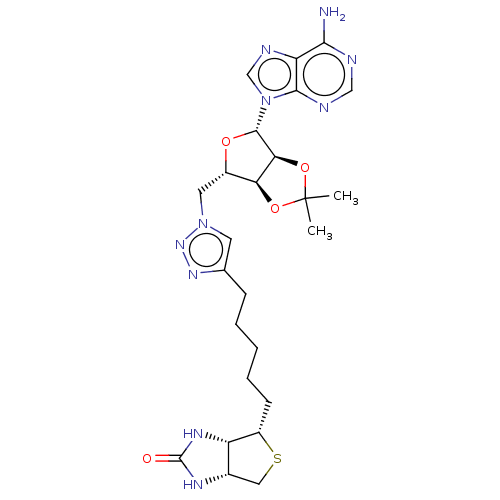
TargetBifunctional ligase/repressor BirA(Escherichia coli (strain K12))
University of Nebraska-Lincoln
Curated by ChEMBL
University of Nebraska-Lincoln
Curated by ChEMBL
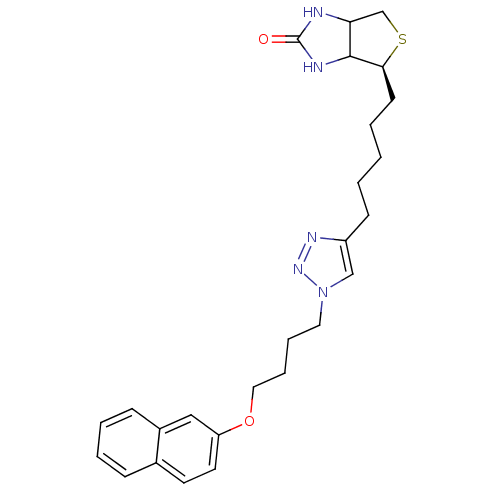
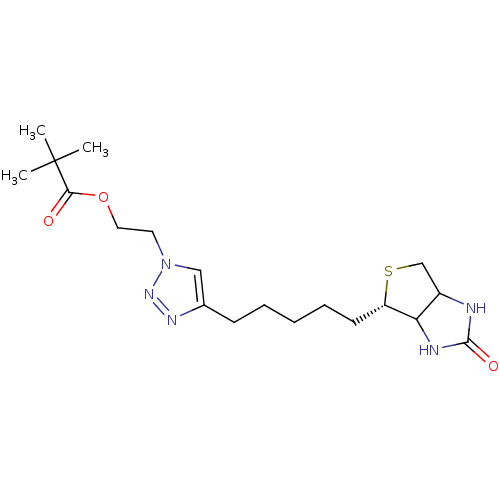
Affinity DataKi: 87nMAssay Description:Inhibition of Escherichia coli recombinant BPL using 3H-biotin as substrate after 10 mins by liquid scintillation counting analysisMore data for this Ligand-Target Pair
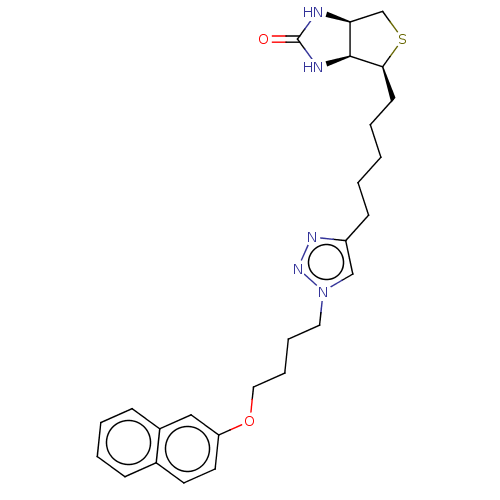
Affinity DataKi: 90nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: 90nM ΔG°: -41.8kJ/mole IC50: 530nMpH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
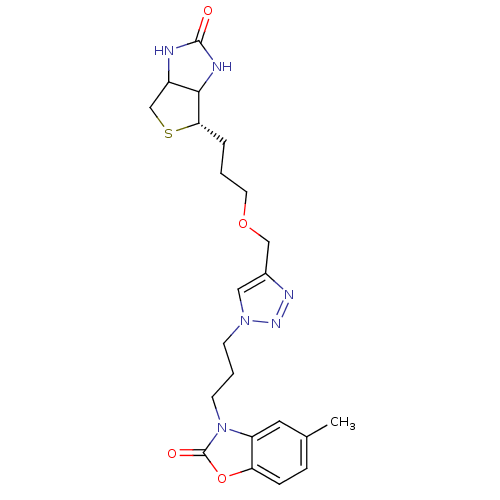
TargetBifunctional ligase/repressor BirA(Escherichia coli (strain K12))
University of Nebraska-Lincoln
Curated by ChEMBL
University of Nebraska-Lincoln
Curated by ChEMBL
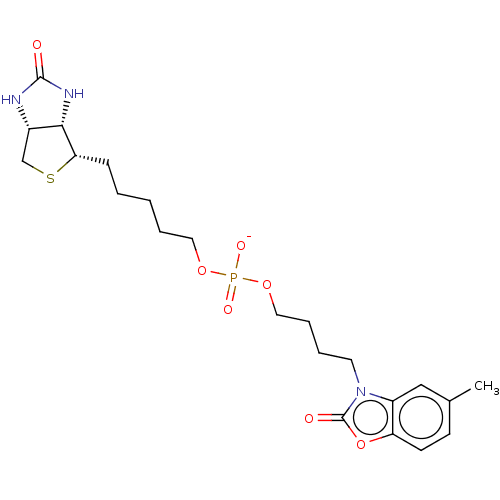
Affinity DataKi: 225nM ΔG°: -39.5kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
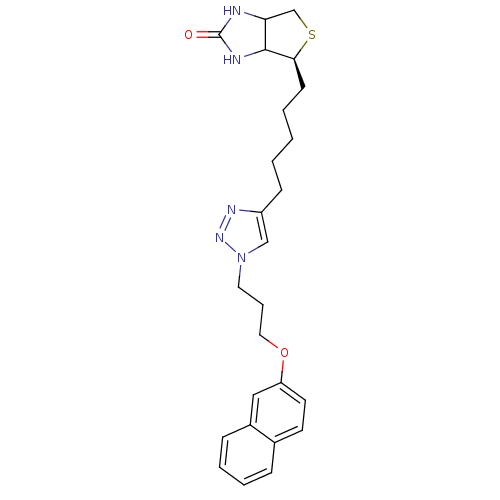
TargetBifunctional ligase/repressor BirA(Escherichia coli (strain K12))
University of Nebraska-Lincoln
Curated by ChEMBL
University of Nebraska-Lincoln
Curated by ChEMBL
Affinity DataKi: 420nM ΔG°: -37.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
TargetBifunctional ligase/repressor BirA(Staphylococcus aureus)
University of Groningen
Curated by ChEMBL
University of Groningen
Curated by ChEMBL
Affinity DataKi: 660nMAssay Description:Inhibition of Staphylococcus aureus biotin protein ligase by LC/MS-SIM analysisMore data for this Ligand-Target Pair
Affinity DataKi: 660nM ΔG°: -36.7kJ/mole IC50: 4.00E+3nMpH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
Affinity DataKi: 1.17E+3nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: 1.17E+3nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: 1.17E+3nM ΔG°: -35.2kJ/mole IC50: 7.00E+3nMpH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
TargetBifunctional ligase/repressor BirA(Staphylococcus aureus)
University of Groningen
Curated by ChEMBL
University of Groningen
Curated by ChEMBL
Affinity DataKi: 1.40E+3nMAssay Description:Binding affinity to Staphylococcus aureus biotin protein ligaseMore data for this Ligand-Target Pair
Affinity DataKi: 1.83E+3nM ΔG°: -34.1kJ/mole IC50: 1.16E+4nMpH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nM ΔG°: >-29.7kJ/mole IC50: 5.00E+4nMpH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
TargetBifunctional ligase/repressor BirA(Escherichia coli (strain K12))
University of Nebraska-Lincoln
Curated by ChEMBL
University of Nebraska-Lincoln
Curated by ChEMBL
Affinity DataKi: >3.00E+4nM ΔG°: >-26.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
TargetBifunctional ligase/repressor BirA(Escherichia coli (strain K12))
University of Nebraska-Lincoln
Curated by ChEMBL
University of Nebraska-Lincoln
Curated by ChEMBL
Affinity DataKi: >3.00E+4nM ΔG°: >-26.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
TargetBifunctional ligase/repressor BirA(Escherichia coli (strain K12))
University of Nebraska-Lincoln
Curated by ChEMBL
University of Nebraska-Lincoln
Curated by ChEMBL
Affinity DataKi: >3.00E+4nM ΔG°: >-26.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
TargetBifunctional ligase/repressor BirA(Escherichia coli (strain K12))
University of Nebraska-Lincoln
Curated by ChEMBL
University of Nebraska-Lincoln
Curated by ChEMBL
Affinity DataKi: >3.00E+4nM ΔG°: >-26.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
TargetBifunctional ligase/repressor BirA(Escherichia coli (strain K12))
University of Nebraska-Lincoln
Curated by ChEMBL
University of Nebraska-Lincoln
Curated by ChEMBL
Affinity DataKi: >3.00E+4nM ΔG°: >-26.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
TargetBifunctional ligase/repressor BirA(Escherichia coli (strain K12))
University of Nebraska-Lincoln
Curated by ChEMBL
University of Nebraska-Lincoln
Curated by ChEMBL
Affinity DataKi: >3.00E+4nM ΔG°: >-26.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
TargetBifunctional ligase/repressor BirA(Escherichia coli (strain K12))
University of Nebraska-Lincoln
Curated by ChEMBL
University of Nebraska-Lincoln
Curated by ChEMBL
Affinity DataKi: >3.00E+4nM ΔG°: >-26.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
TargetBifunctional ligase/repressor BirA(Escherichia coli (strain K12))
University of Nebraska-Lincoln
Curated by ChEMBL
University of Nebraska-Lincoln
Curated by ChEMBL
Affinity DataKi: >3.00E+4nM ΔG°: >-26.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
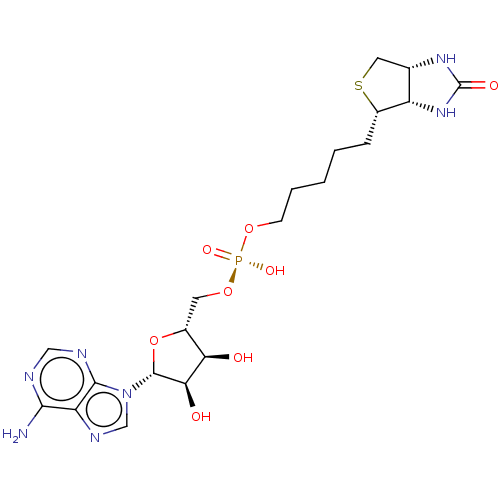
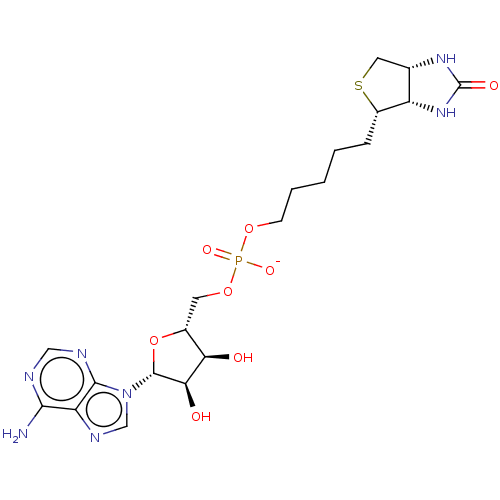

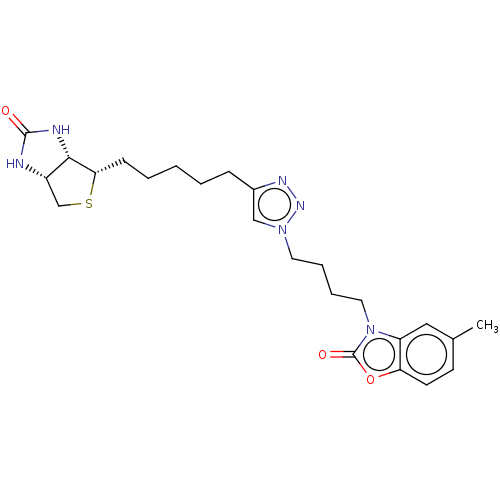
 3D Structure (crystal)
3D Structure (crystal)