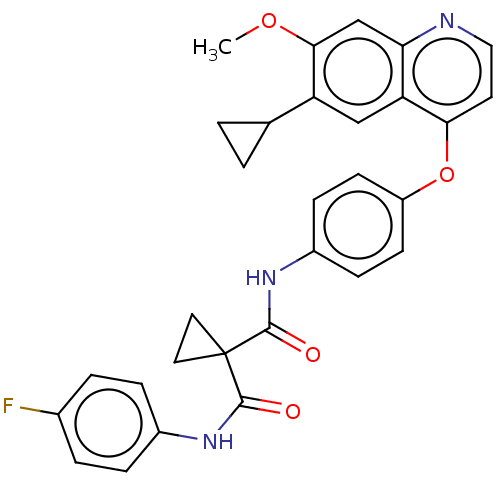
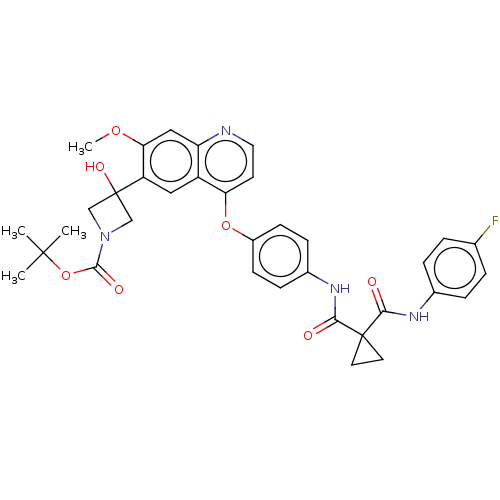
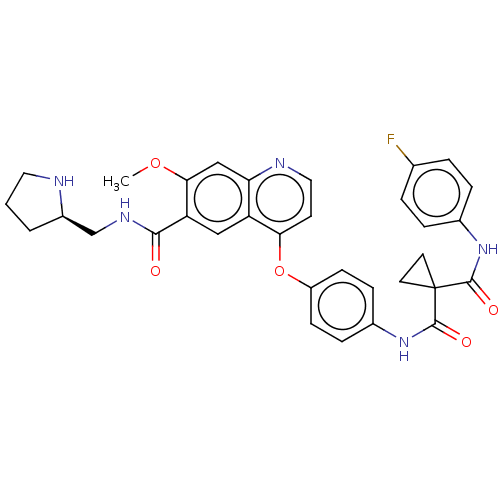
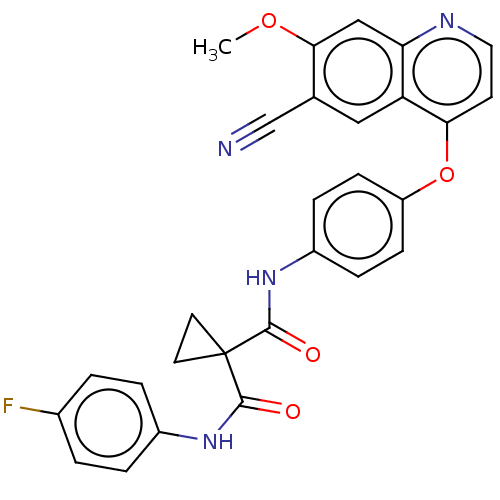
Report error Found 76 Enz. Inhib. hit(s) with Target = 'Tyrosine-protein kinase Mer [557-882,H628Q,R794A]'
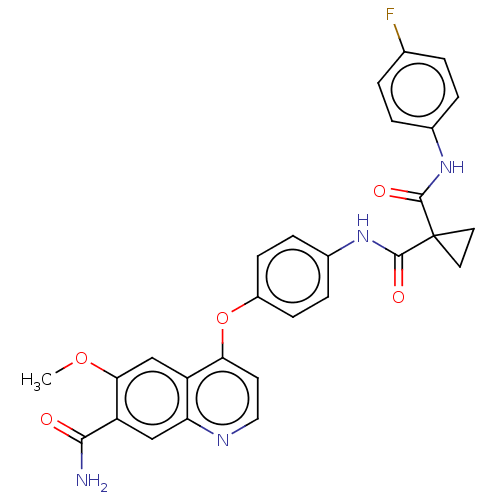
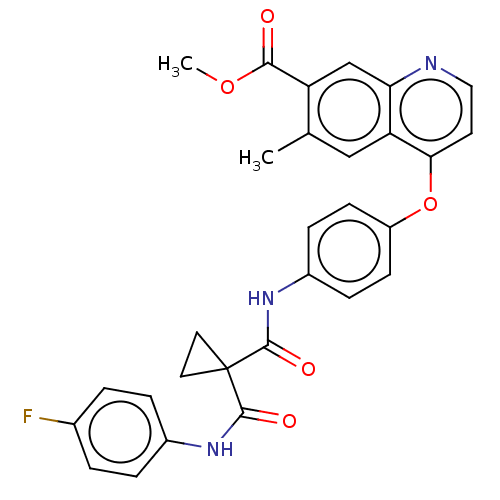
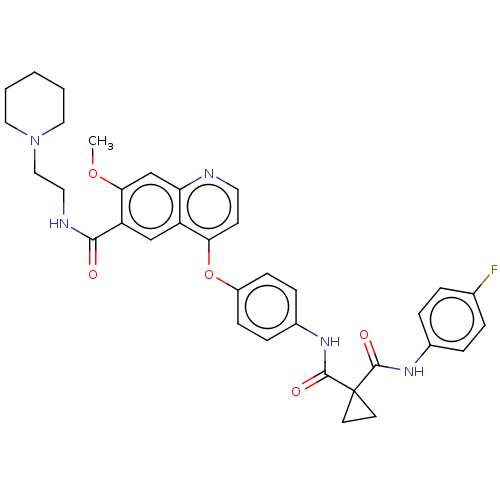
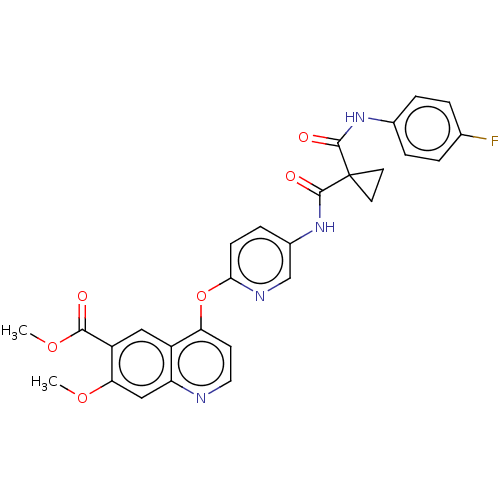
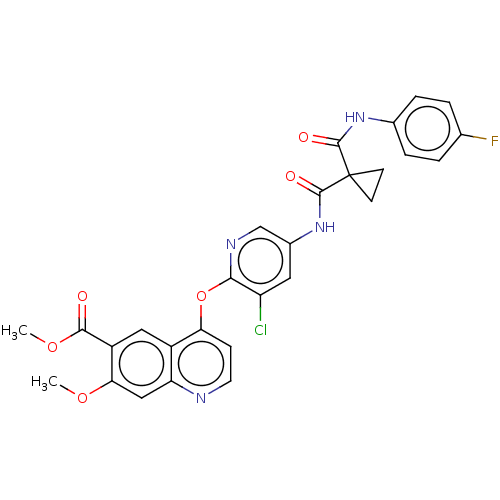
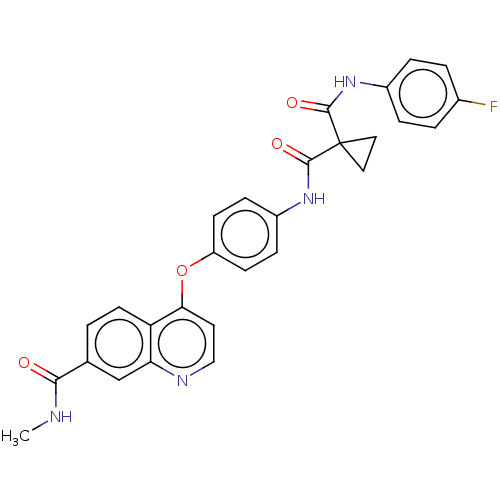
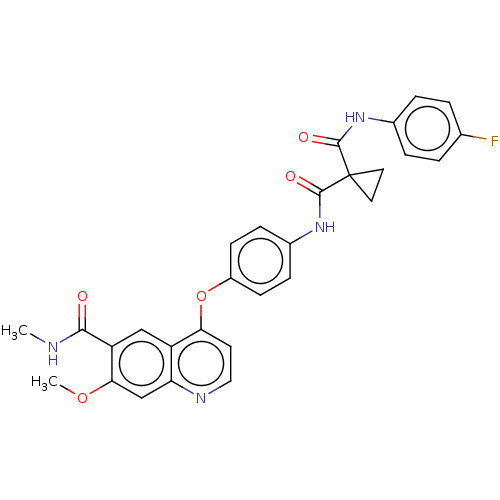
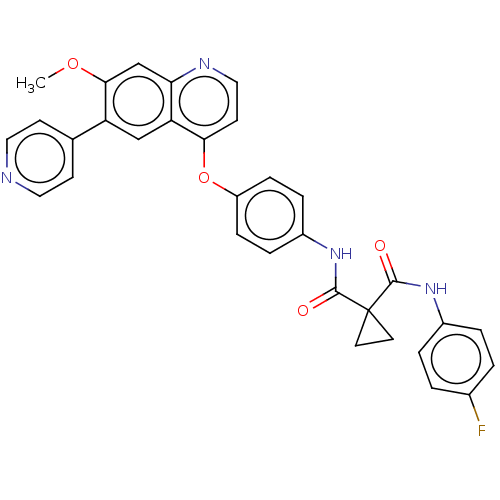
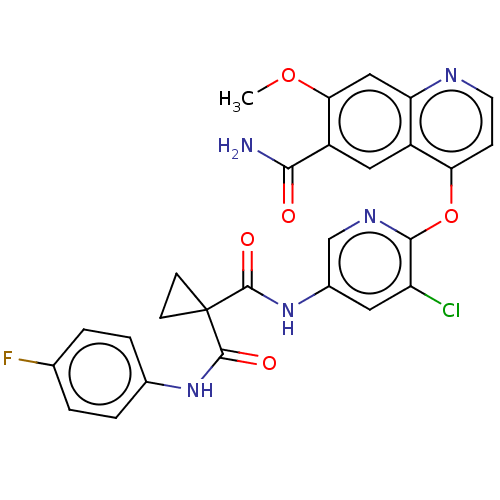
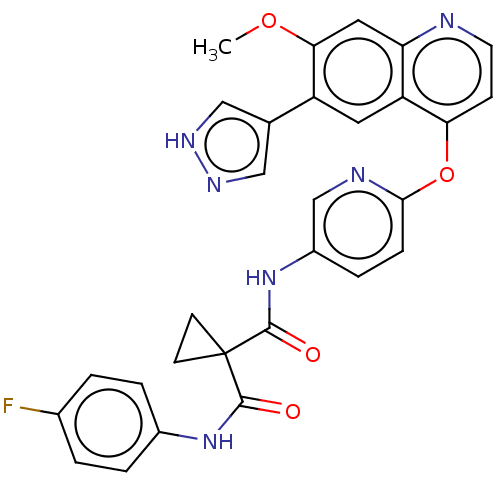
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
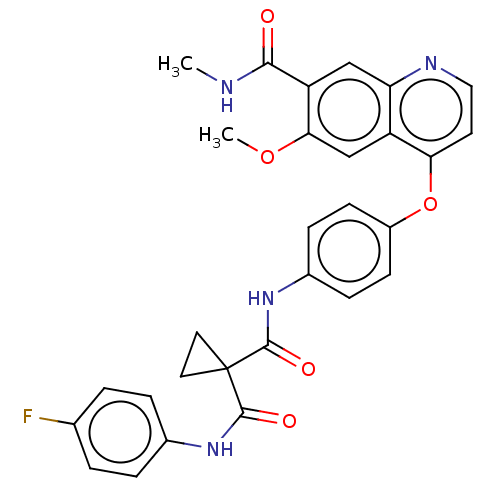
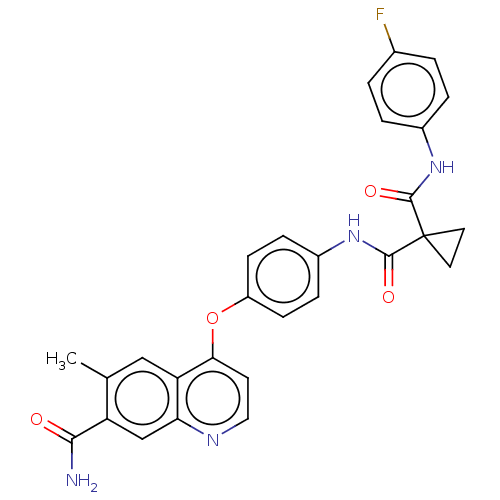
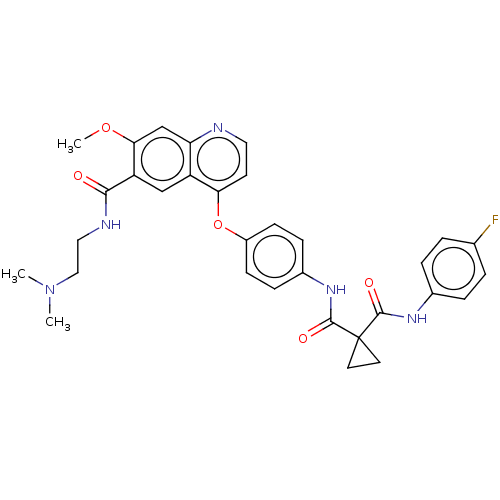
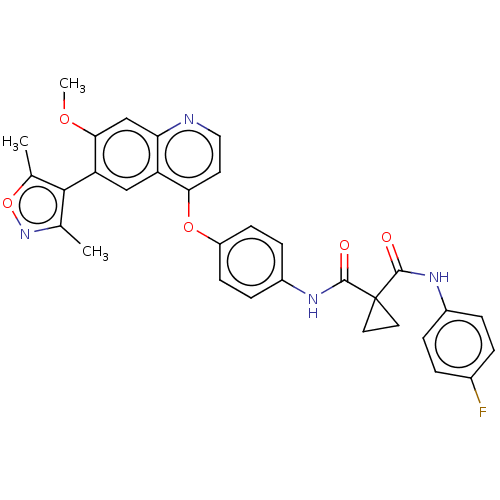
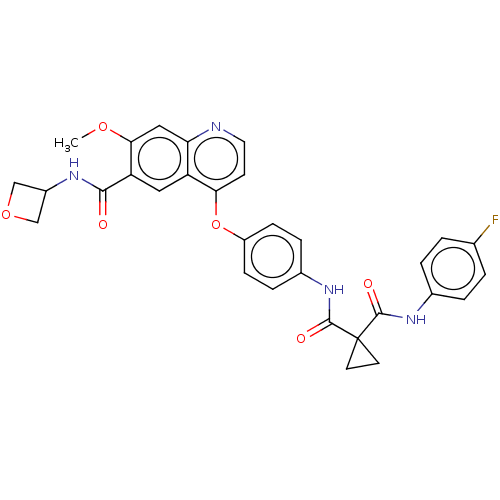
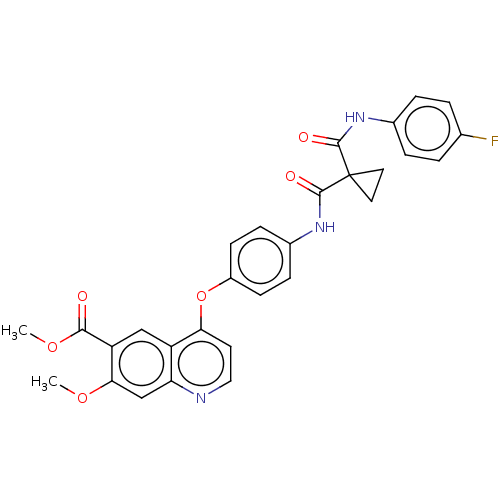
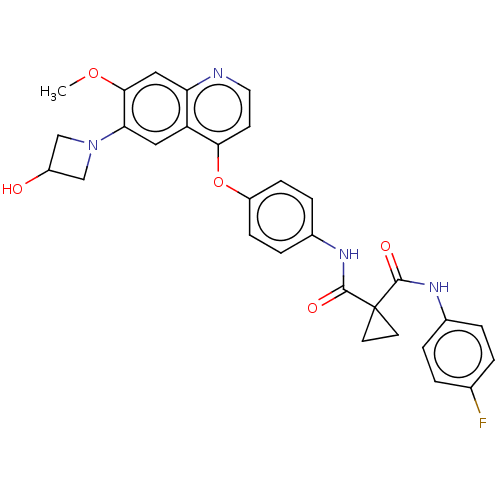
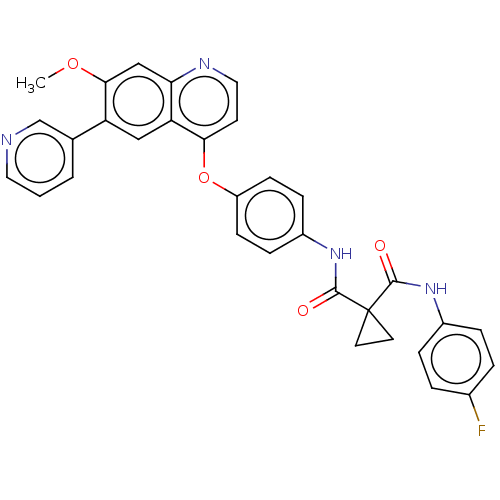
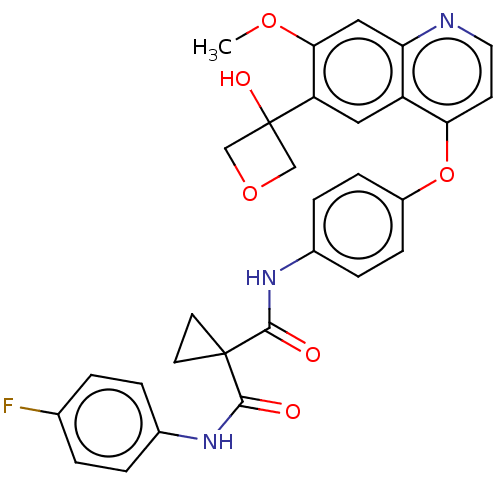
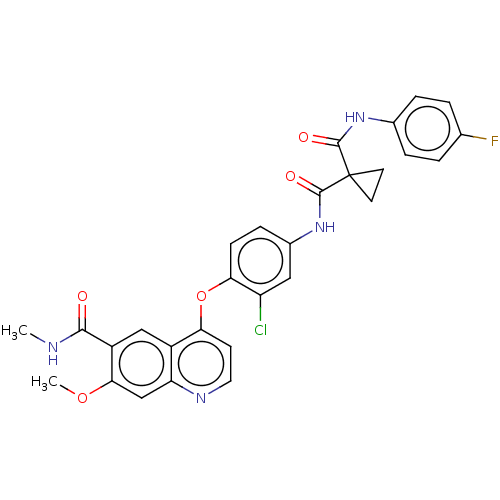
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
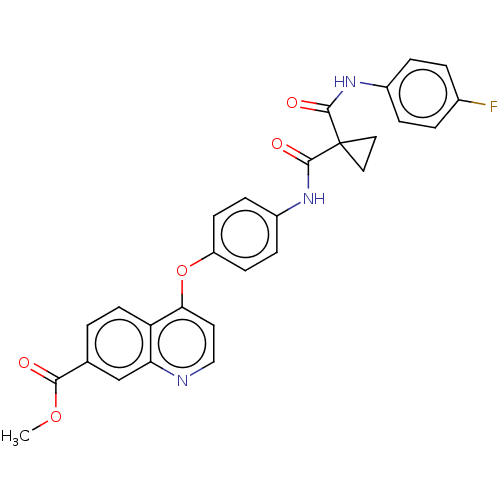
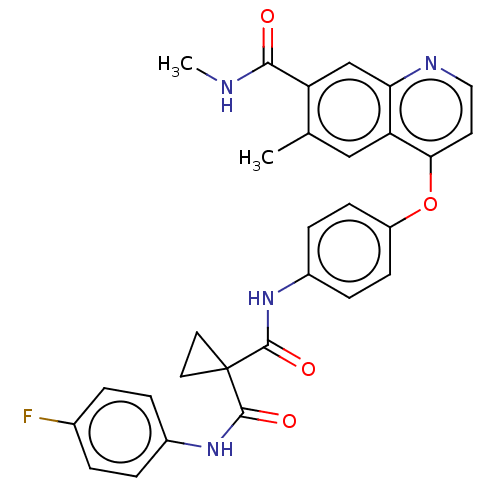
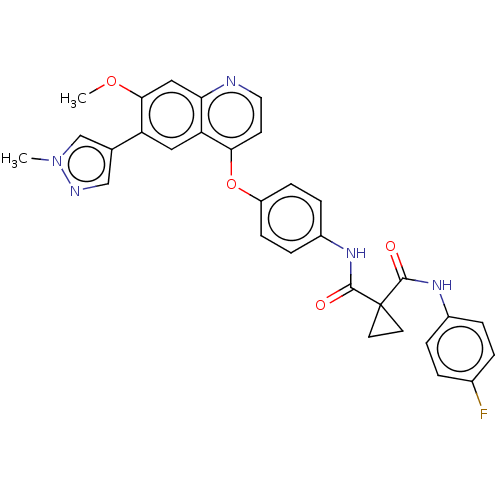
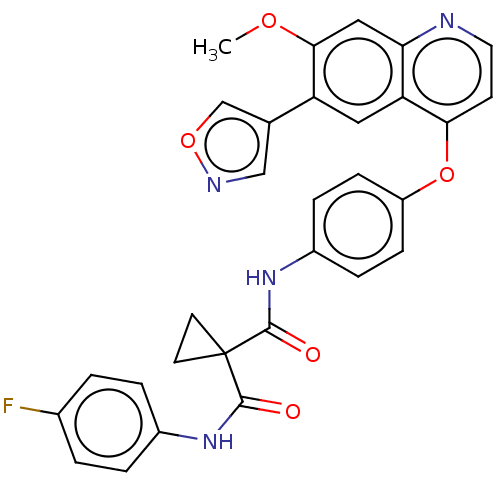
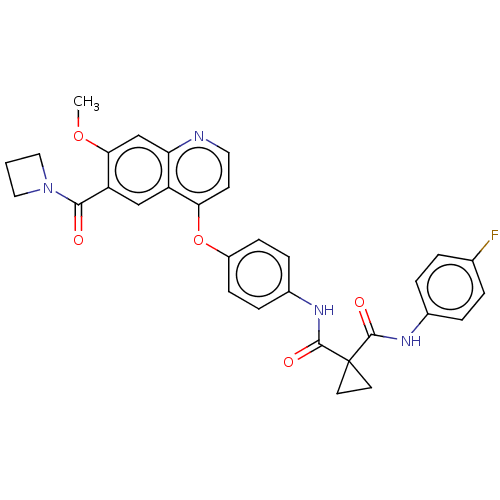
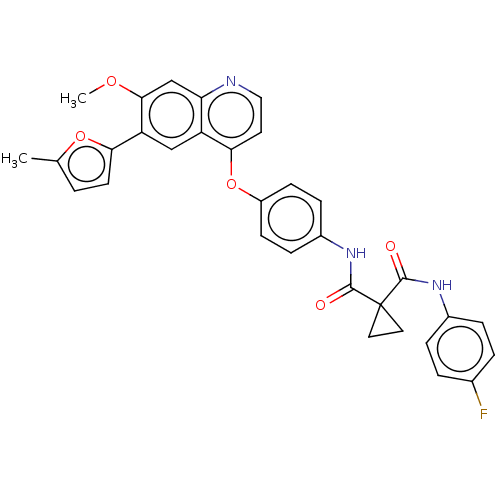
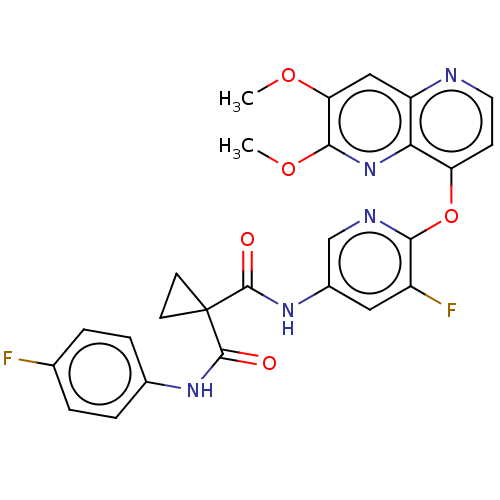
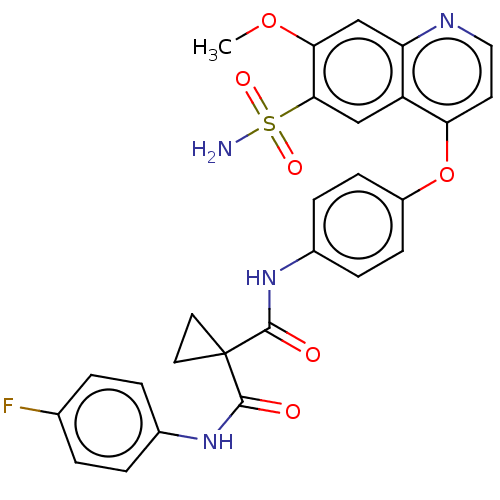
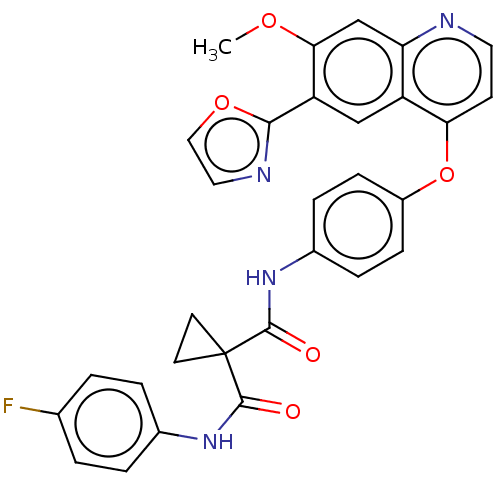
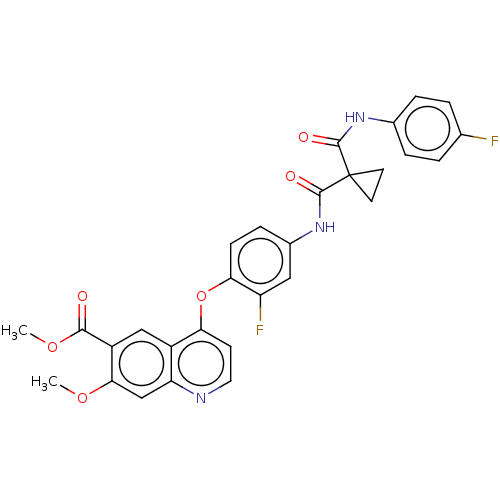
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
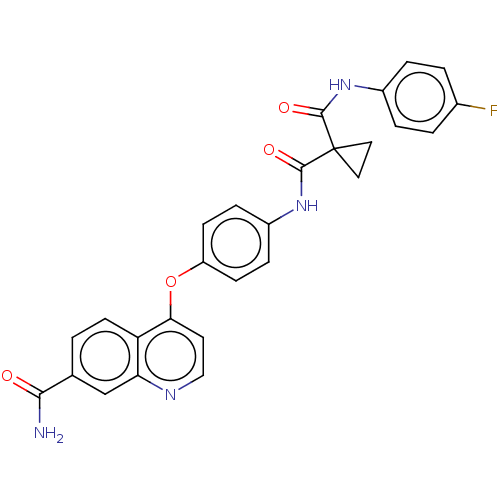
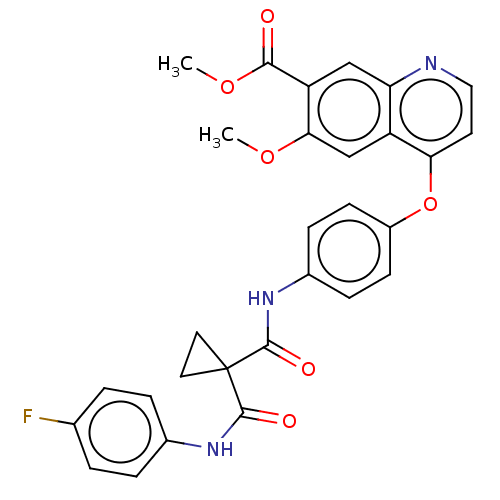
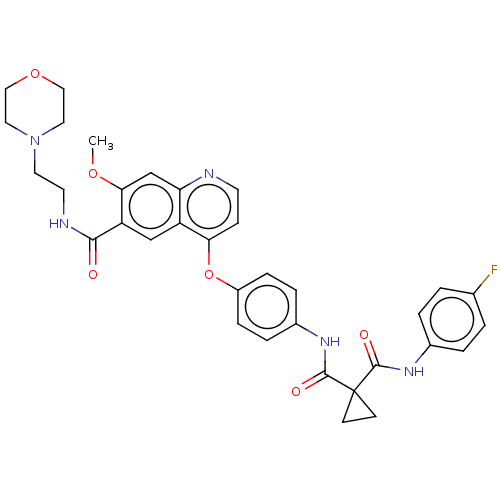
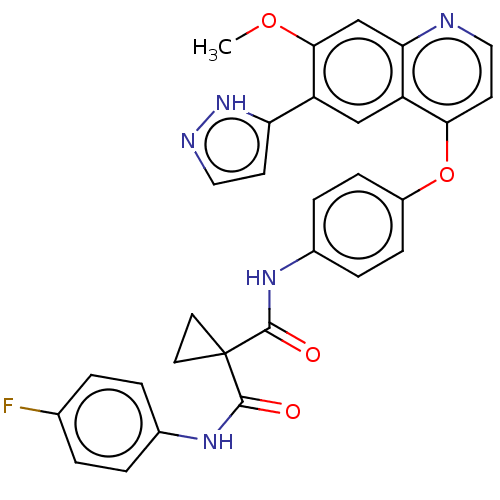
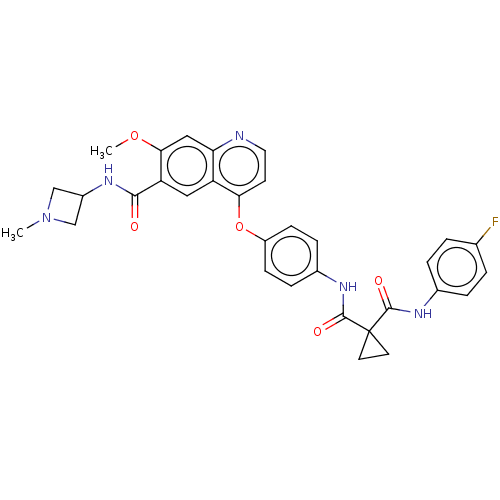
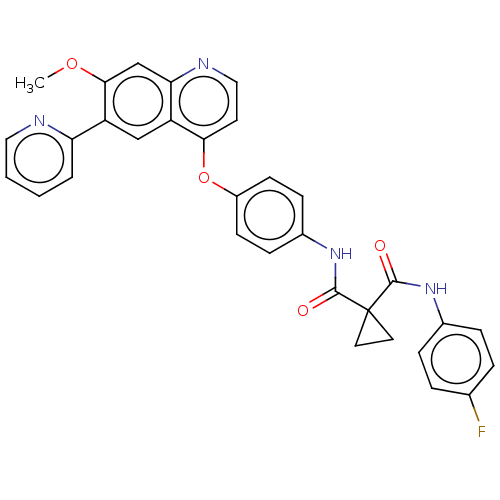
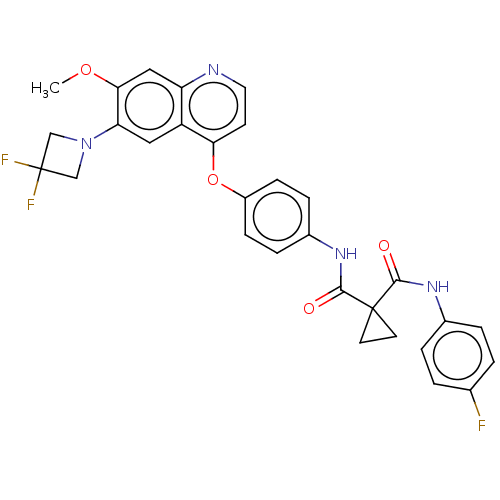
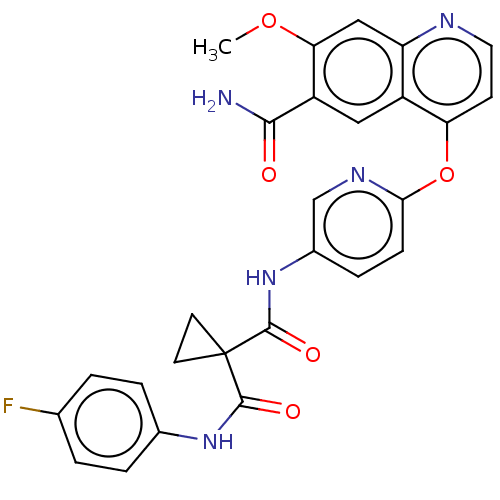
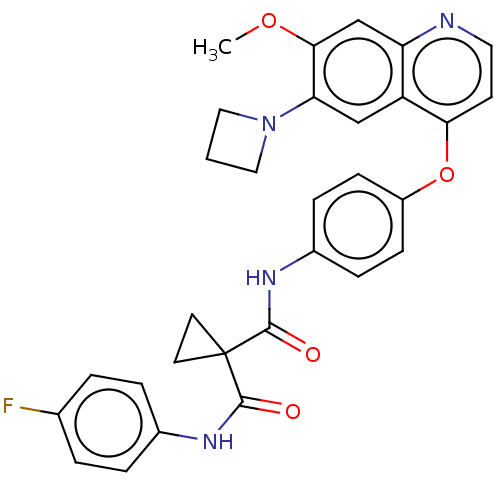
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 10nMAssay Description:Human Mer (residues R557-E882 with H628Q and R794A, 0.7 nM) was incubated with 8 mM MOPS pH 7.0, 0.2 mM EDTA, 30 mM NaCl, 250 μM GGMEDIYFEFMGGKK...More data for this Ligand-Target Pair
Ligand InfoSimilars